8.2 Genome projects & gene technologies
1/128
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
129 Terms
Genome
The complete set of genes / DNA in a cell
Proteome
The full range of proteins that a cell can produce (coded for by the cell's DNA / genome)
Cellular proteome
The proteins produced by a given type of cell under a certain set of conditions.
What is genome sequencing and why is it important?
● Identifying the DNA base sequence of an organism's genome
● So amino acid sequences of proteins that derive from an organism's genetic code can be determined (proteome)
Sequencing projects have read the genomes of a range of organisms, including humans
What can the sequencing of a simple organism's genome allow?
Can predict the proteome
As they lack introns and regulatory genes
So the relationship between the genome and proteome is straight forward
Explain how determining the genome of a pathogen could allow vaccines to be developed
● Could identify the pathogen's proteome
● So could identify potential antigens to use in the vaccine
Suggest some other potential applications of genome sequencing projects
● Identify genes/alleles associated with genetic diseases / cancers
○ New targeted drugs / gene therapy can be developed
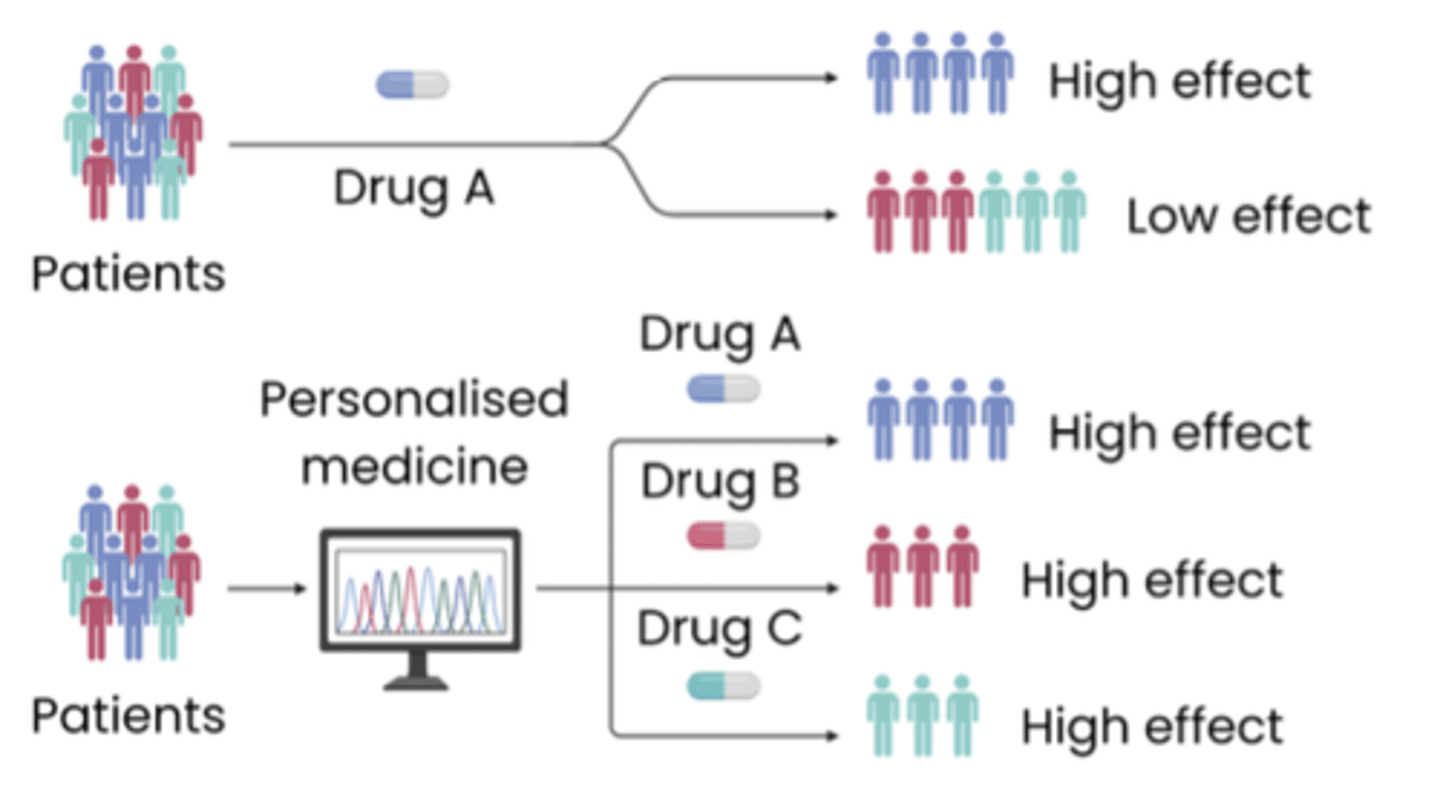
○ Can screen patients, allowing early prevention / personalised medicine
● Identification of species and evolutionary relationships
Why is it more difficult to determine the proteome of humans compared to bacteria ?
The human genome contains:
● Introns
● Regulatory genes
Explain why the genome cannot be directly translated into the proteome in complex organisms
● Presence of non-coding DNA (e.g. introns within genes do not code for polypeptides)
● Presence of regulatory genes (code for proteins which regulate expression of other genes, e.g. by coding for miRNA)
Describe how sequencing methods are changing
● They've become automated (faster, more cost-effective, larger scale)
● Continuously updated
8.4 GENE TECHNOLOGIES
Describe how whole-genome shotgun sequencing is used to determine the DNA sequence of an organism.
The entire genome (DNA) is cut into small, easily sequenced sections and computer algorithms are used to (reassemble) align the overlapping segments.
What is recombinant DNA technology?
Transfer of DNA fragments from one organism / species, to another
Recombinant DNA
The combined DNA of 2 or more organisms

Transgenic organisms / GMO organisms
Organisms that contain recombinant DNA
Why can transferred DNA be translated within cells of recipient (transgenic) organisms ?
1. Genetic code is universal
2. Transcription and translation mechanisms are universal
Give a use of recombinant DNA
Gene therapy; recombinant DNA used to alter a patient's genes to treat/cure a disease
What are the possible issues with recombinant DNA technology?
● Expensive
● Possible side effects associated with gene therapy / rejected by immune system / infection
● Ethical objections to altering genes.
What is the first step of sequencing the genome?
Cut up the DNA into smaller DNA fragments
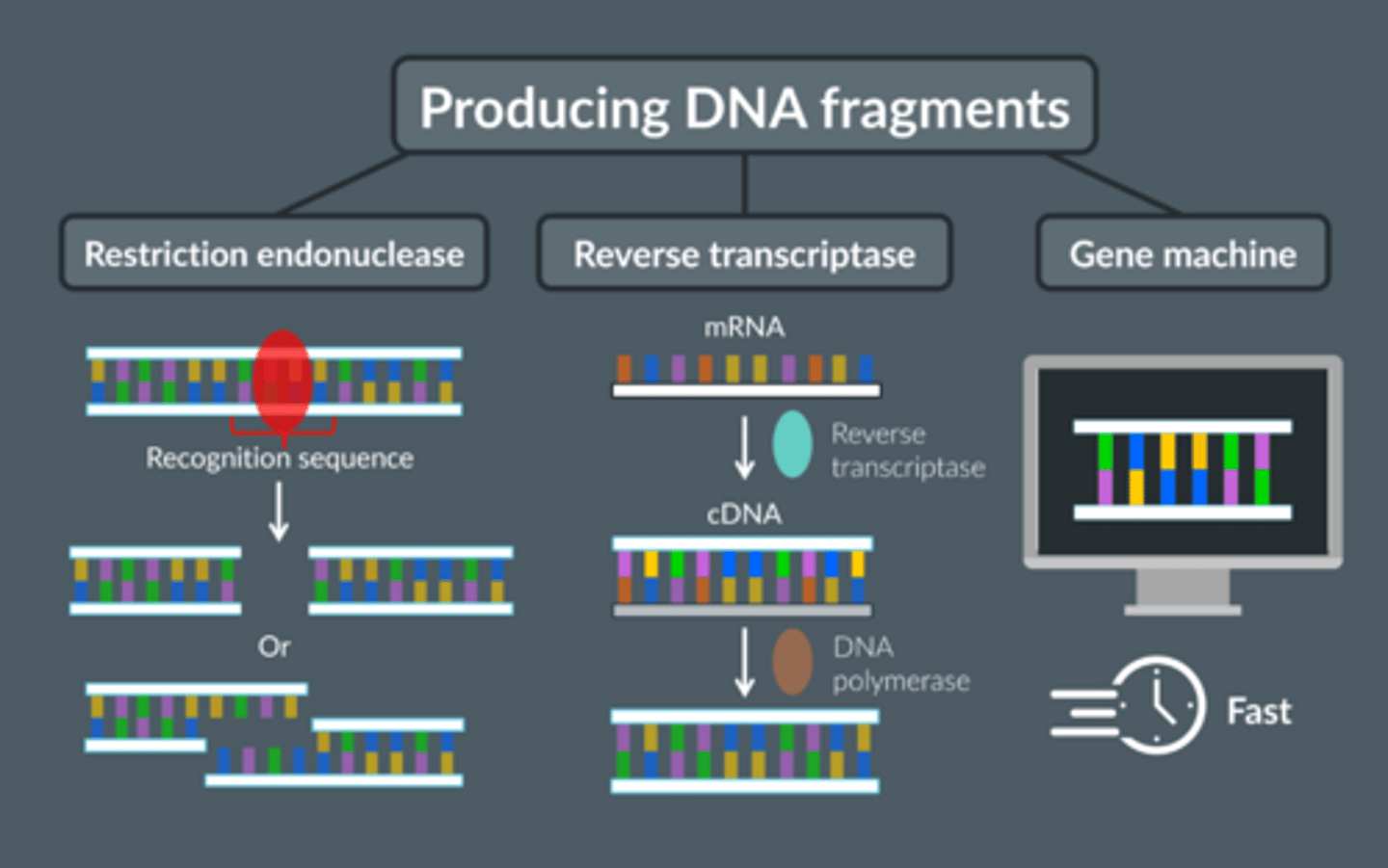
Give the 3 ways of isolating a gene
1. Reverse transcriptase to convert mRNA → cDNA
2. Restriction endonucleases to cut fragments containing the desired gene from DNA
3. Create gene in a gene machine
Name the enzyme used to cut open a plasmid.
Restriction endonuclease
Name the enzyme used to insert a gene into a plasmid.
DNA ligase
Describe how DNA fragments can be produced using restriction endonuclease (enzymes)
1. Restriction endonuclease cuts DNA at specific set of base 'recognition sequences' on either side of the desired gene
○ Breaks phosphodiester bonds between adjacent nucleotides
○ 2 recognition sequences → DNA fragment
○ Shape of recognition site complementary to active site
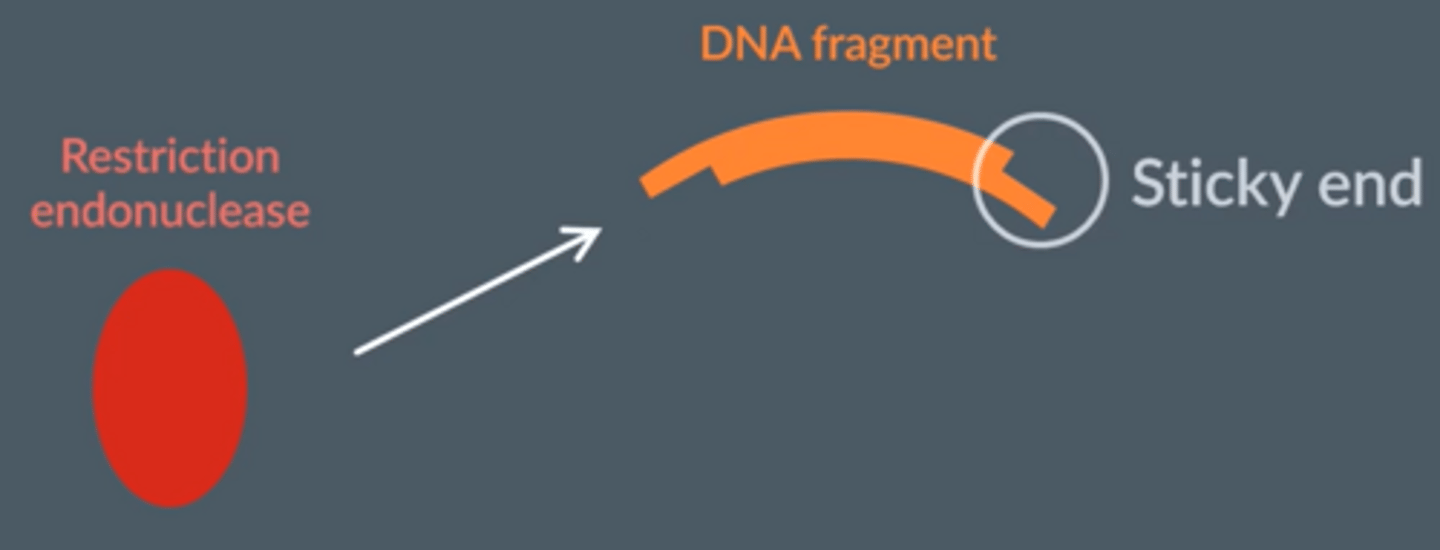
2. Many cut in a staggered fashion forming 'sticky ends' (single stranded overhang) or straight edges (blunt ends)
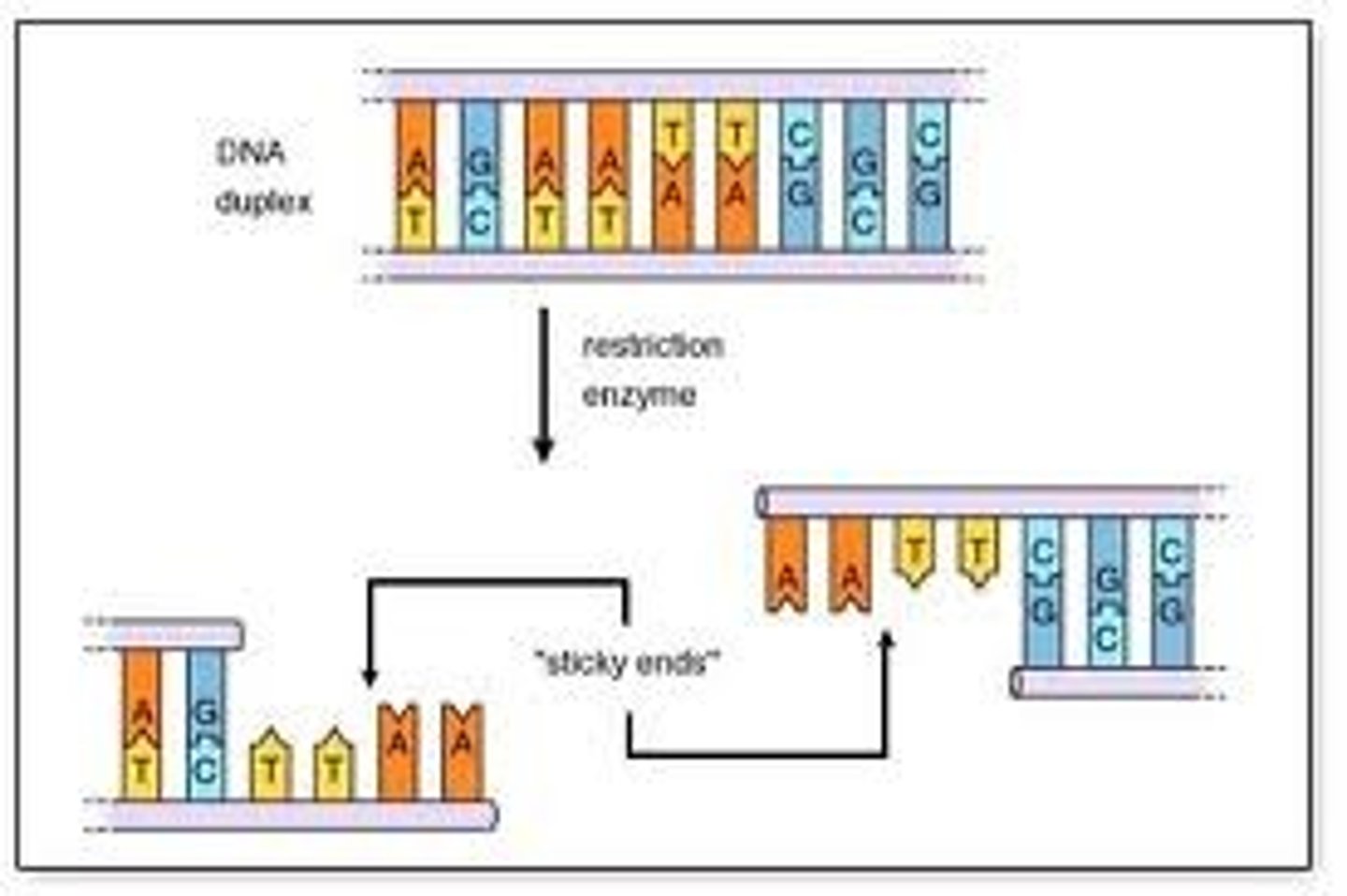
Describe restriction enzymes that hydrolyse the DNA to create blunt ends vs sticky ends.
● Some enzymes cut the DNA at the same location in the double strand between 2 opposite base pairs, leaving 2 straight edges → blunt end
● Others cut the DNA in different locations in the double-strand, creating palindromic staggered ends and exposed DNA bases, "sticky ends."
Palindromic sequence
Nucleic acid sequence: read the same antiparallel (5'-3' same as complementary 5'-3')
The recognition sequence of both blunt and sticky ends are palindromic.
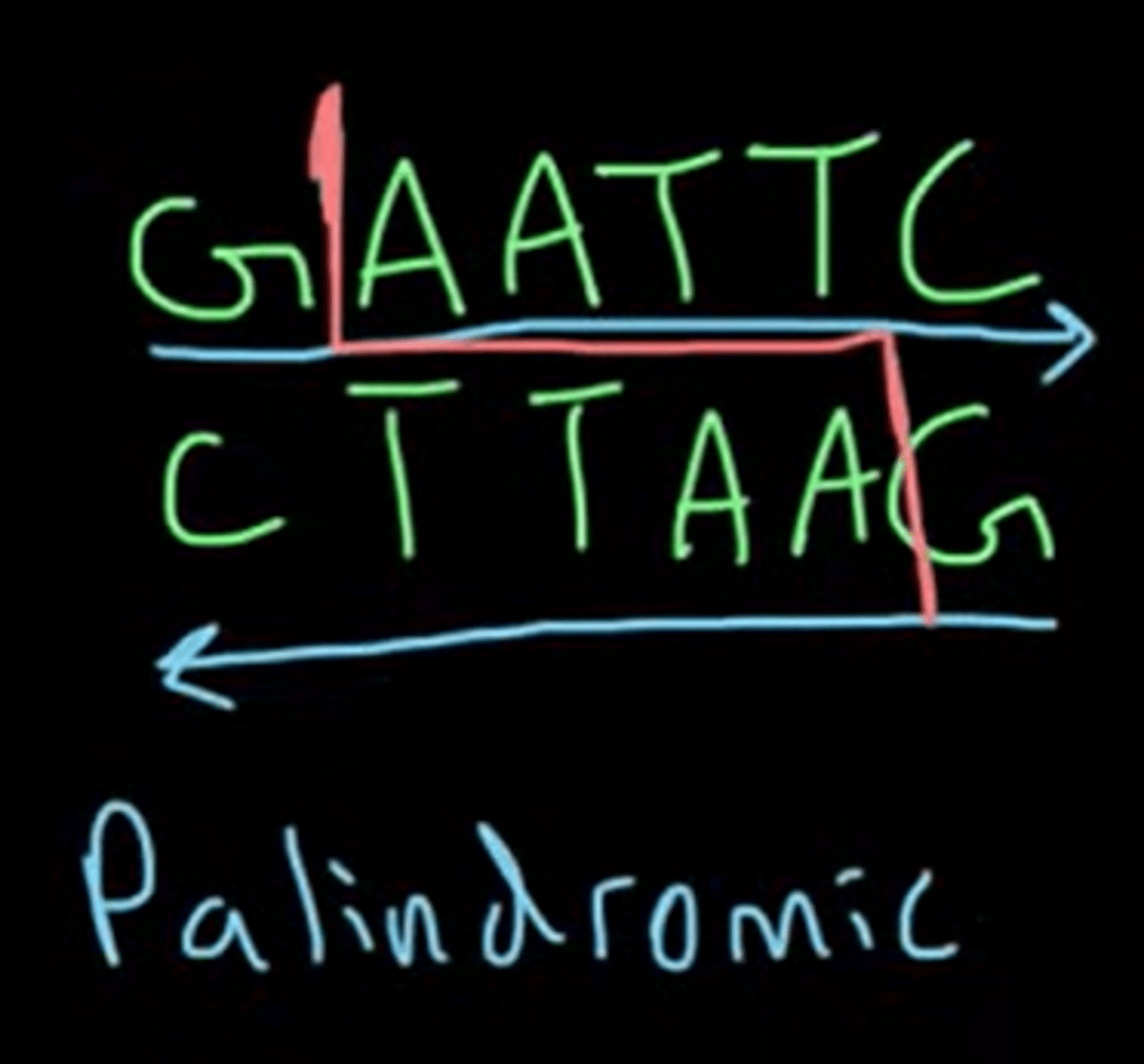

Give the importance of sticky ends
● The cut ends of the DNA double strand are left with a single strand which is a few nucleotide bases long
● They can bind (anneal) the DNA fragment with other pieces of DNA with complementary base pairs.
When would it NOT be useful to produce a DNA fragment using a restriction endonuclease ?
What ARE they useful for ?
When the bacteria are used to make functional proteins
As they cut out DNA containing introns, which cannot be spliced by bacteria, so would end up with a non-functional protein
Useful for gene cloning, genetic fingerprinting, making DNA probes.
Describe how DNA fragments can be produced from mRNA
1. Isolate mRNA from a cell that readily synthesises the protein coded for by the desired gene
2. Add DNA nucleotides to mRNA strand and reverse transcriptase → reverse transcriptase uses mRNA as a template to synthesise a single strand of complementary DNA (cDNA)
3. DNA polymerase can form a 2nd strand of DNA using cDNA as a template.
Suggest 2 advantages of obtaining genes from mRNA rather than directly from the DNA removed from cells
● Much more mRNA than DNA in cells making the protein → easily extracted
● No introns in mRNA as been removed by splicing (in eukaryotes) whereas DNA contains introns
○ So can be transcribed & translated by prokaryotes who can't remove introns by splicing
Describe how fragments of DNA can be produced using a gene machine
● Amino acid sequence of protein determined, so DNA base sequence can be established
○ Synthesises fragments of DNA quickly & accurately from scratch without need for a DNA template
(AA sequence → mRNA codons → complementary base sequence)
● No introns so can be transcribed & translated by prokaryotes
What happens before the gene machine assembles the fragments of DNA ?
Sequence is checked for biosafety and biosecurity to ensure it's safe and ethical to produce
Advantages of the gene machine
● Quick (doesn't rely on enzymes)
● Very accurate
● No introns so can be transcribed and translated in prokaryotic cells
● Any sequence of nucleotides can be produced
Advantages of each technique of isolating a gene
Reverse transcriptase: mRNA present in cell is from actively transcribed genes, so lots of the mRNA of interest available to make cDNA
Restriction endonuclease: Sticky ends on DNA fragment make it easier to insert to make recombinant DNA
Gene machine: Can design exact DNA fragment you want, with sticky ends, labels and preferential codons
Disadvantages of each technique of isolating a gene
Reverse transcriptase: more steps so time consuming, technically more difficult
Restriction endonuclease: still contains introns
Gene machine: need to know the sequence of amino acids or bases
Tissue plasminogen activator (tPA) is a protein that breaks down blood clots. It can be given to patients through an IV in the arm.
In order to make enough tPA for medical use, geneticists take the gene coding for tPA and place it into a bacterium. The bacterium then produces tPA which is collected.
Here 3 possible methods for obtaining the tPA gene.
1. Use a restriction enzyme to cut out a DNA fragment containing the gene
2. Use reverse transcriptase to convert the mRNA coding for tPA into cDNA
3. Use a gene machine to create the tPA gene
Explain why the bacterium wouldn't be able to make tPA using the DNA produced by method 1.
● The DNA fragment produced in method 1 would contain introns.
● Bacteria unable to carry out splicing, so these introns would be present in the final mRNA strand.
● Translation of this mRNA strand would result in a non-functional protein.
Name an IN VITRO & IN VIVO technique used to amplify DNA fragments
● In vitro (in glass) - polymerase chain reaction PCR
● In vivo (inside a living organism) - culturing transformed host cells e.g. bacteria
Polymerase chain reaction (PCR)
An automated method for amplifying (copying) DNA fragments in a continuous cycle
What does the reaction mixture/master mix contain for PCR ?
1. DNA fragment
2. DNA polymerase (taq polymerase)
3. Primers
4. DNA nucleotides
Inside a thermocycler
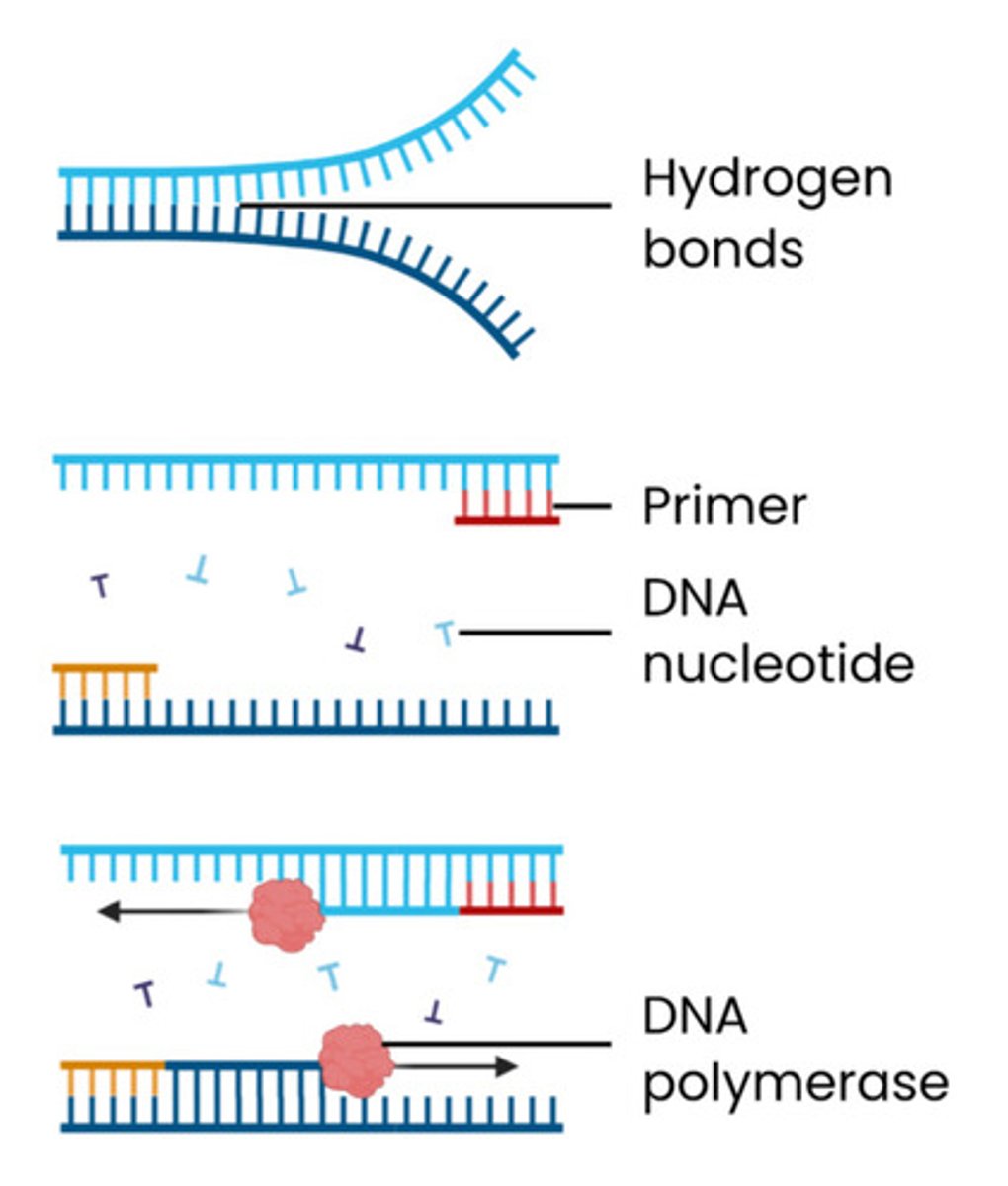
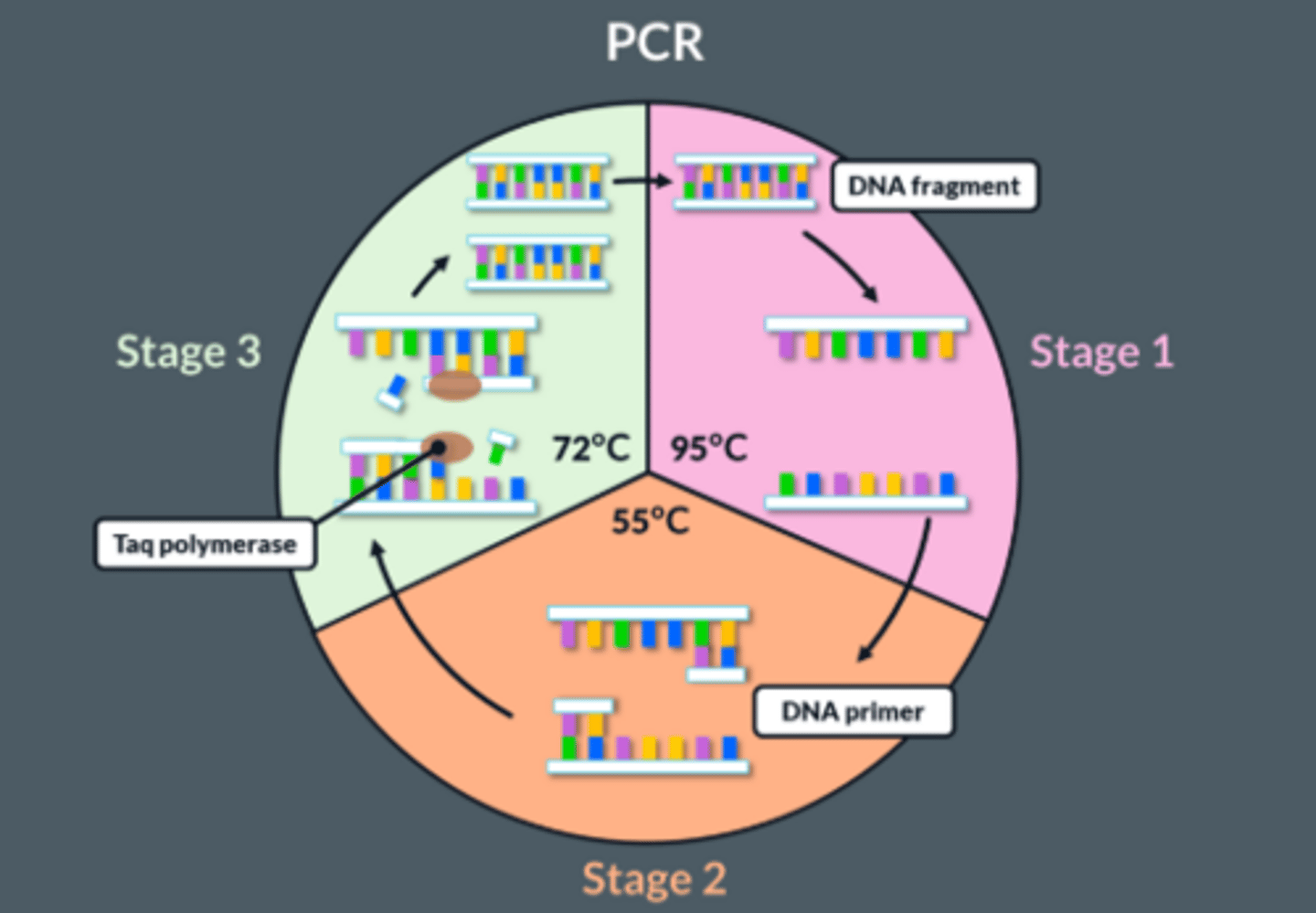
Explain how DNA fragments can be amplified by PCR
1. Mixture heated to 95°C
● This separates the 2 strands of DNA fragments
● Breaking hydrogen bonds between bases
2. Mixture cooled to 55°C
● This allows primers to anneal (bind) to their complementary bases at end of the DNA fragment
● By forming hydrogen bonds between complementary bases
3. Mixture heated to 72°C
● Nucleotides align next to complementary exposed bases
● Optimum temperature for taq polymerase which joins adjacent DNA nucleotides, forming phosphodiester bonds
● Begins at primer on both strands
Cycle is repeated - in every cycle, the amount of DNA doubles causing an exponential increase (2ⁿ)
6 marker PCR
● Add primers and nucleotides
● Heat DNA to 95
● To break hydrogen bonds / separate strands
● Cool to 50
● To allow primers to anneal to complementary bases by complementary base pairing
● Heat to 72
● DNA (taq) polymerase
● Adds complementary nucleotides to the separate DNA strands → produces 2 new identical DNA fragments are formed
● Cycle is repeated many times
Explain the role of primers in PCR
● Primers are short, single stranded DNA fragments
● Complementary to DNA base sequence at ends of region to be copied / start of desired gene
● Allowing DNA polymerase to bind to start synthesis (can only add nucleotides onto pre-existing 3' end)
Explain why 2 different primers are required
● 2 different primers (forward and reverse) are required
● As base sequences at opposite ends of the 2 strands of DNA are different
Suggest one reason why DNA replication eventually stops in PCR
Limited number of primers and nucleotides, eventually used up
Evaluate PCR
Advantages:
● Faster than in vivo. Can make 100 billion copies of DNA within hours
● Doesn't require living cells
● Efficient. No need to isolate DNA fragment/gene from the host DNA (as with in vivo cloning);
● Automated → efficient + rapid
Disadvantages:
● Contaminated DNA also multiplies
Vector
Transfers DNA fragments into a host cell
E.g. plasmids

Summary of in vivo gene cloning
Isolation of DNA;
Insertion into vector;
Transformation into host;
Identification by gene markers;
Growth/cloning of host cell population
What kind of DNA fragments are required for in vivo gene cloning ?
● DNA fragments produced by restriction endonucleases, which have sticky ends
● The same restriction endonuclease is then used on the plasmid, producing sticky ends complementary to the DNA fragment
Explain why the same restriction endonuclease must be used on both the DNA fragment and the plasmid during in vivo gene cloning.
● The restriction endonuclease cuts both the DNA fragment and the plasmid at the same recognition sequence.
● So the sticky ends on both are complementary to each other.
● So, the DNA fragment and the plasmid can slot together with hydrogen bonds between the complementary bases.
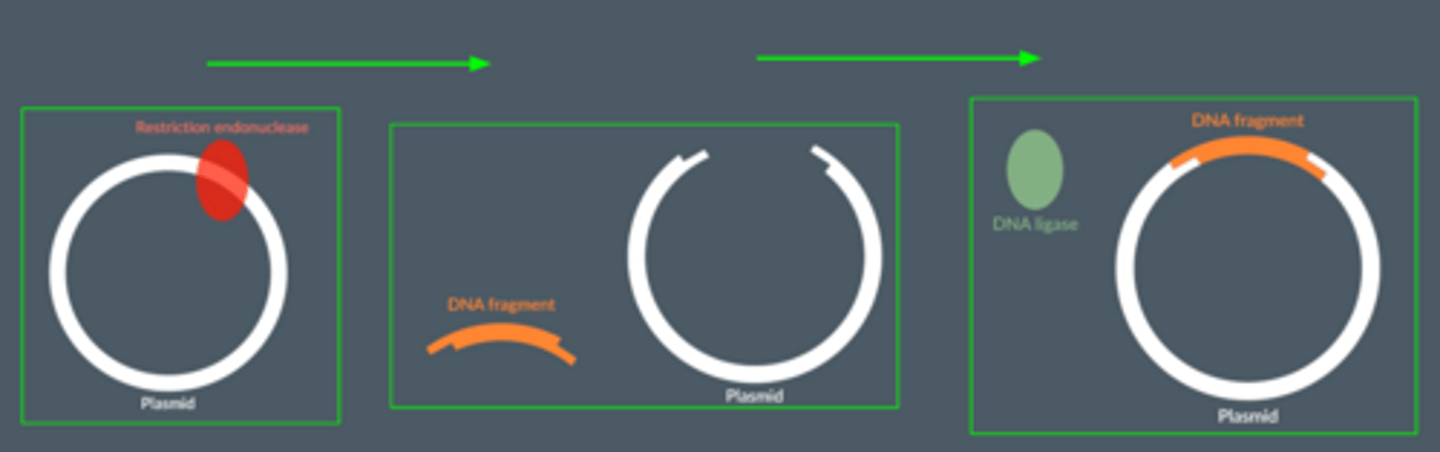
What is the role of DNA ligase ?
● Joins together the nucleotides of the DNA fragment with the nucleotides of the plasmid
● by catalysing the formation of phosphodiester bonds via condensation reactions.
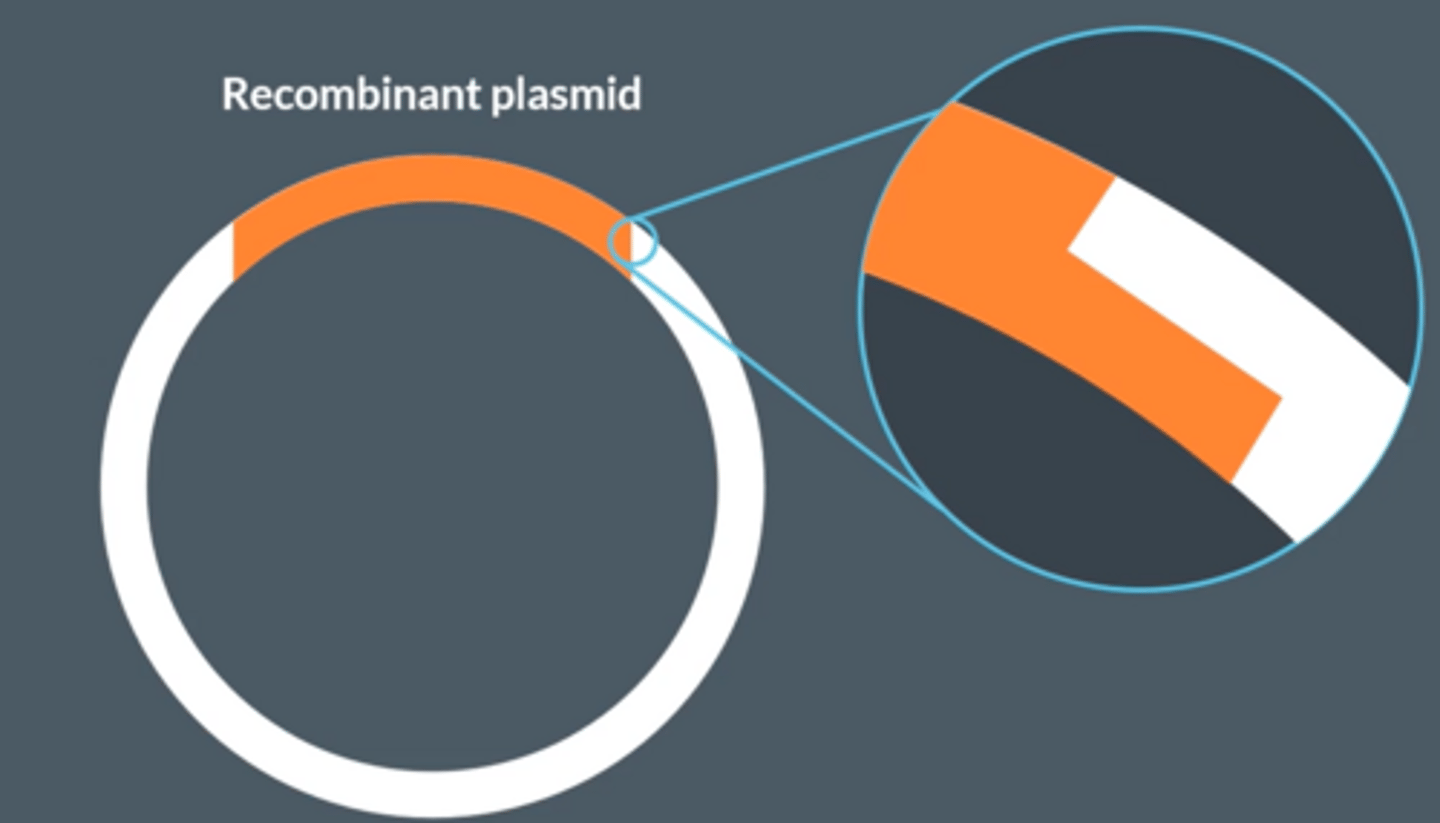
The sticky ends of the recombinant plasmid looks like blunt ends
Promoter
● A sequence of DNA bases that acts as a binding site for RNA polymerase and transcription factors.
● PROMOTES transcription
Terminator
● A sequence of bases that acts as a detachment site for RNA polymerase and transcription factors
● TERMINATES transcription at the appropriate spot.
Transformation
Adding a plasmid to a host cell
1. Add calcium ions
2. Heat shock
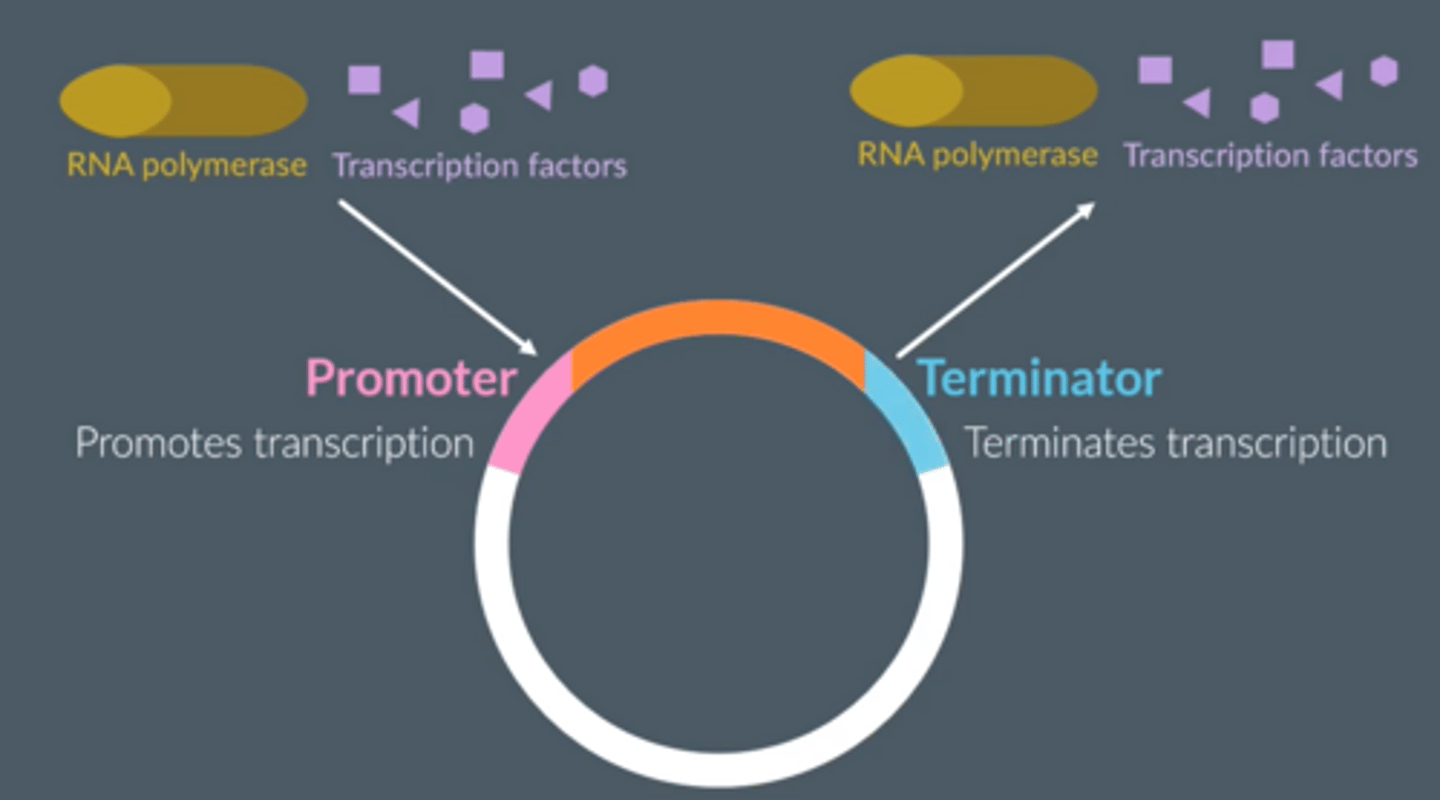
After transformation, why might the bacteria not contain a recombinant plasmid ?
● Plasmids will rejoin without incorporating a DNA fragment.
● DNA fragments will join to each other, rather than a plasmid.
● Bacteria won't take up a recombinant plasmid during transformation.
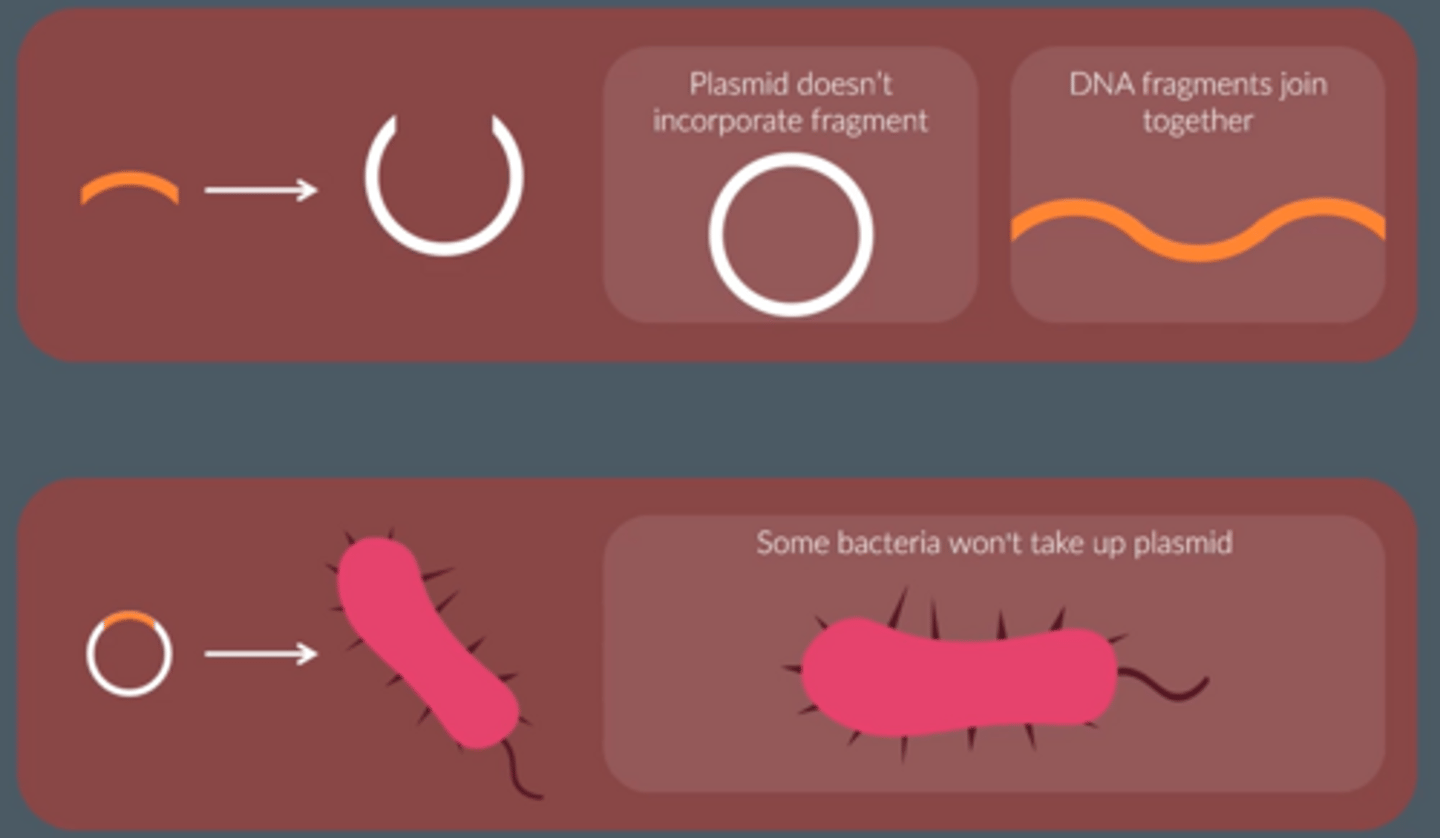
3 possible bacteria produced
● No plasmid
● Plasmid
● Recombinant plasmid
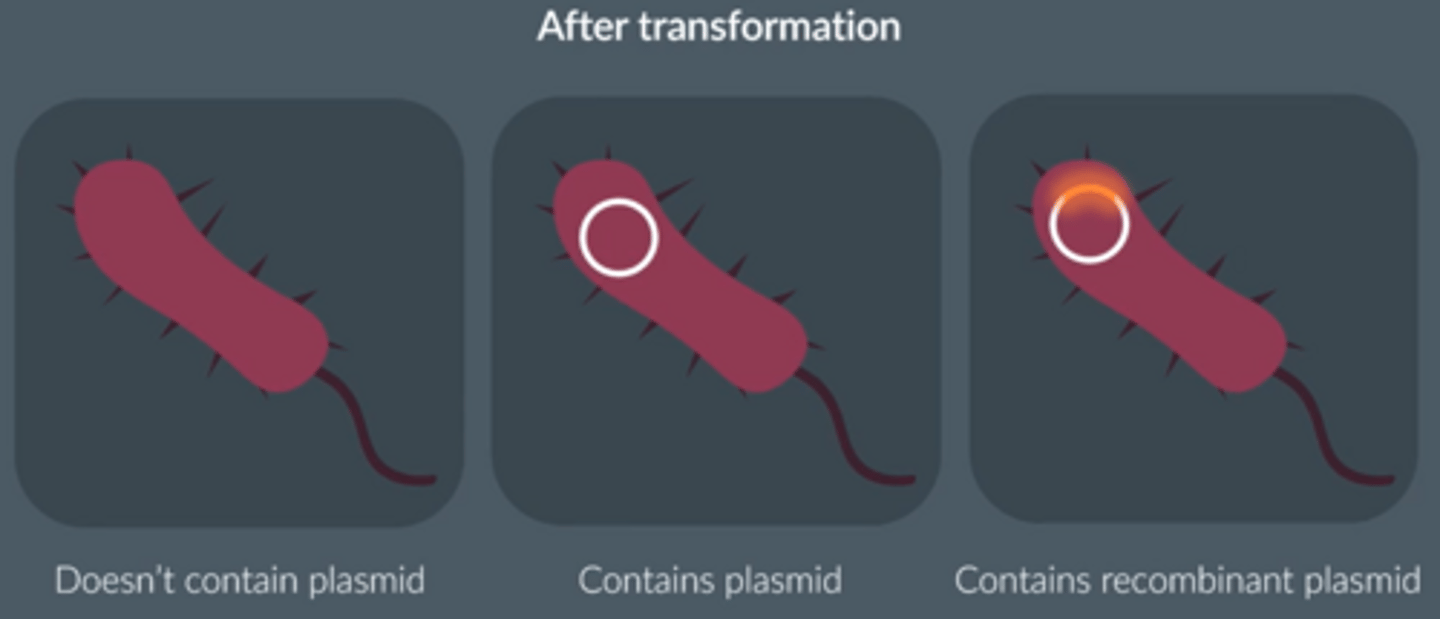
What would a scientist add to a plasmid to identify whether a DNA fragment has been incorporated into the plasmid?
Marker gene
After transformation...
Scientists use a plasmid that contains an antibiotic resistance gene (MARKER GENE) to identify which bacteria have taken up a plasmid,
Bacteria are placed in antibiotic, any bacteria without a plasmid will die
A 2nd antibiotic resistance marker gene is used. The addition of the DNA fragment into the plasmid disrupts the function of the antibiotic resistance gene so any bacteria that contain a recombinant plasmid (i.e. contains the DNA fragment) die.
Replica plating: The addition of the DNA fragment into the plasmid disrupts the function of the ampicillin resistance gene
Alternatives to replica plating
Fluorescent marker genes
Fluorescent and enzyme marker genes provide observable properties in the bacteria.
Bacteria which have taken up the recombinant plasmid...
DON'T display the observable property.
(Like adding random words into the middle of a sentence; it won't make sense anymore.)
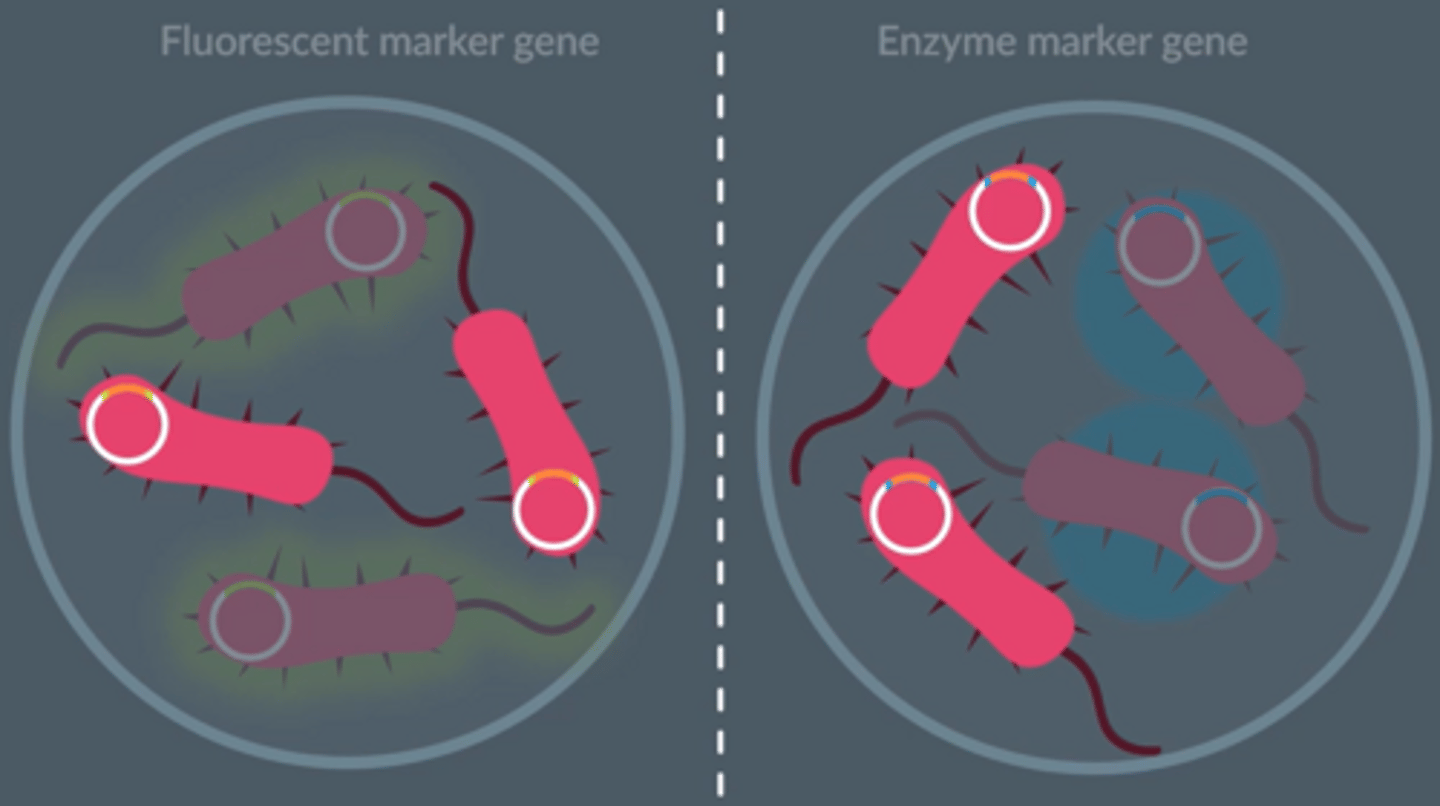
Benefit of using a fluorescent marker gene over an antibiotic resistance marker gene
● Faster; no need for replica plating
● Fluorescence is an observable property.
● There is no risk of antibiotic resistance being passed onto other bacteria.
● More economical.
Summarise the steps involved in amplifying DNA fragments in vivo
1. Add promoter and terminator regions to DNA fragments
2. Insert DNA fragments & marker genes into vectors (e.g. plasmids) using restriction enzymes and ligases
3. Transform host cells (e.g. bacteria) by inserting these vectors
4. Detect genetically modified (GM) / transformed cells / organisms by identifying those containing the marker gene (e.g. that codes for a fluorescent protein)
5. Culture these transformed host cells, allowing them to divide and form clones
Following this, DNA can be extracted from the host cells if needed or the host cells can produce a protein coded for by a gene in the DNA fragment
Explain why promoter and terminator regions are added to DNA fragments that are used to genetically modify organisms
Promoter regions
● Allow transcription to start by allowing RNA polymerase to bind to DNA
● Can be selected to ensure gene expression happens only in specific cell types
○ Eg. in gland cells of a mammal so the protein can be easily harvested
Terminator regions
● Ensure transcription stops at the end of a gene, by stopping RNA polymerase
How can promoter regions be used to control exactly which of the animal's body cells produce the protein ? Why is this useful ?
● Promoter regions that are only activated in specific cell types
● If the protein is only produced in certain cells it can be harvested more easily.
● Producing the protein in the wrong cells could damage the organism.
What are the role of vectors in recombinant DNA technology?
To transfer DNA into host cells / organisms e.g. plasmids or viruses (bacteriophage)
Explain the role of enzymes in inserting DNA fragments into vectors
1. Restriction endonucleases cut vector DNA
○ Same enzyme used that cut the gene out so vector DNA & fragments have 'sticky ends' that can join by complementary base pairing
2. DNA ligase joins DNA fragment to vector DNA
○ Forming phosphodiester bonds between adjacent nucleotides
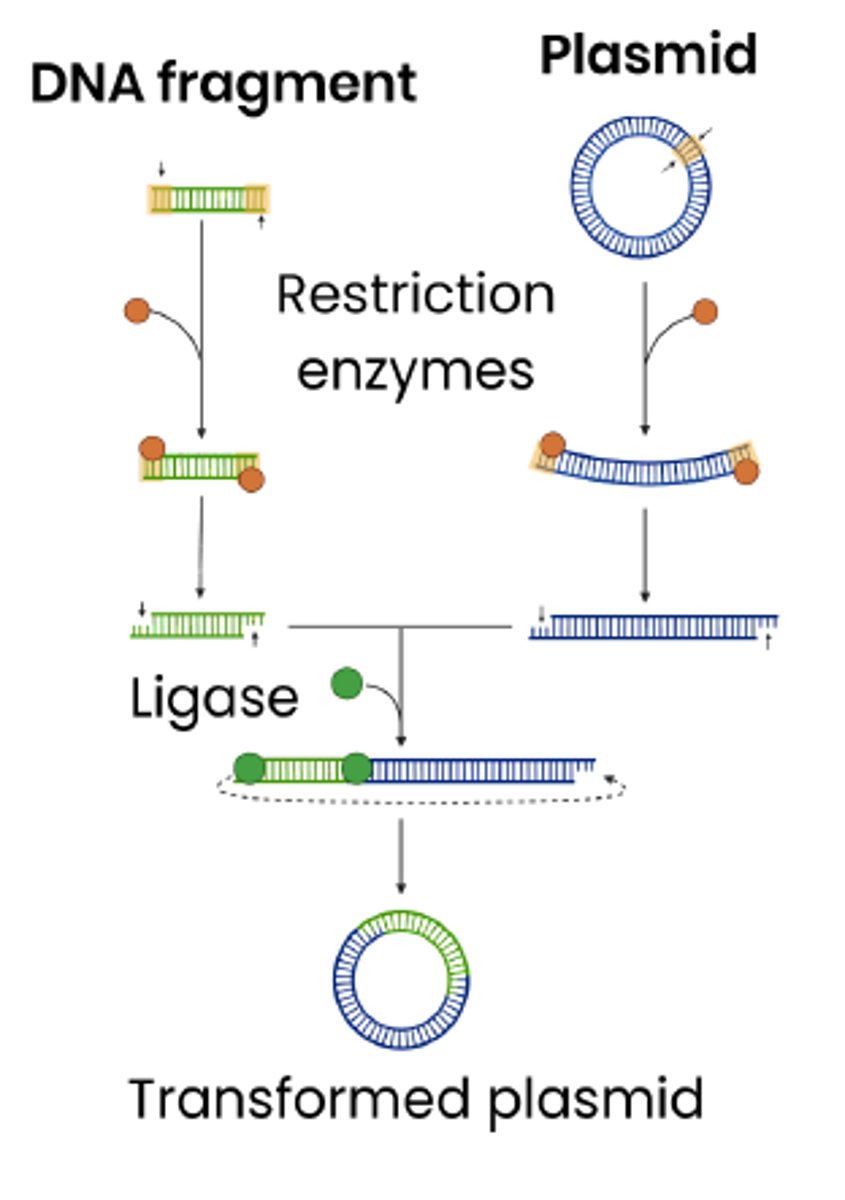
Describe how host cells are transformed using vectors
● Plasmids enter host cells
● Following heat shock in a calcium ion solution
● Viruses inject their DNA into cells which is then integrated into host DNA
How are host cells transformed?
Cell membrane of the host cell must be made more permeable
By mixing with calcium ions and heat shocked (sudden increase in temperature)
Explain why marker genes are inserted into vectors
● To allow detection of genetically modified / transgenic cells / organisms
○ If marker gene codes for antibiotic resistance, cells that survive antibiotic exposure = transformed
○ If marker gene codes for fluorescent proteins, cells that fluoresce under UV light = transformed
● As not all cells / organisms will take up the vector and be transformed
Suggest and explain one reason why bacteria might not be able to produce every human protein.
● Cannot splice (pre-mRNA), so cannot remove introns
● Do not have Golgi apparatus, so cannot process / modify (proteins);
● Do not have (required) transcriptional factors so cannot carry out transcription/produce mRNA;
Suggest how recombinant DNA technology can be useful

Describe gene therapy
● New (recombinant) DNA introduced to cells, often containing healthy / functional alleles
● To overcome effect of faulty alleles in people with genetic disorders e.g. cystic fibrosis
Note - if body cells are altered, changes are not heritable. Gene therapy in gametes is currently illegal.
Students should be able to relate recombinant DNA technology to gene therapy.
Suggest some issues associated with gene therapy
● Effect is short lived as modified cells (eg. T cells) have a limited lifespan → requires regular treatment
● Immune response against genetically modified cells or viruses due to recognition of antigens
● Long term effect not known - side effects e.g. could cause cancer
○ DNA may be inserted into other genes, disrupting them → interfering with gene expression
Suggest why humanitarians might support recombinant DNA technology
● GM crops increase yields → increased global food production → reduced risk of famine / malnutrition
● Gene therapy has potential to cure many genetic disorders
● 'Pharming' makes medicines available to more people as medicines cheaper
Suggest why environmentalists and anti-globalisation activists might
oppose recombinant DNA technology
● Recombinant DNA may be transferred to other plants → potential herbicide resistant 'superweeds'
● Potential effects on food webs eg. affect wild insects → reduce biodiversity
● Large biotech companies may control the technology and own patents
Students should be able to evaluate the ethical, financial & social issues associated with the use and ownership of recombinant DNA technology in agriculture, in industry & in medicine AND balance the humanitarian aspects
of recombinant DNA technology with the opposition from environmentalists & anti-globalisation activists.
Exam insight: common mistakes ❌
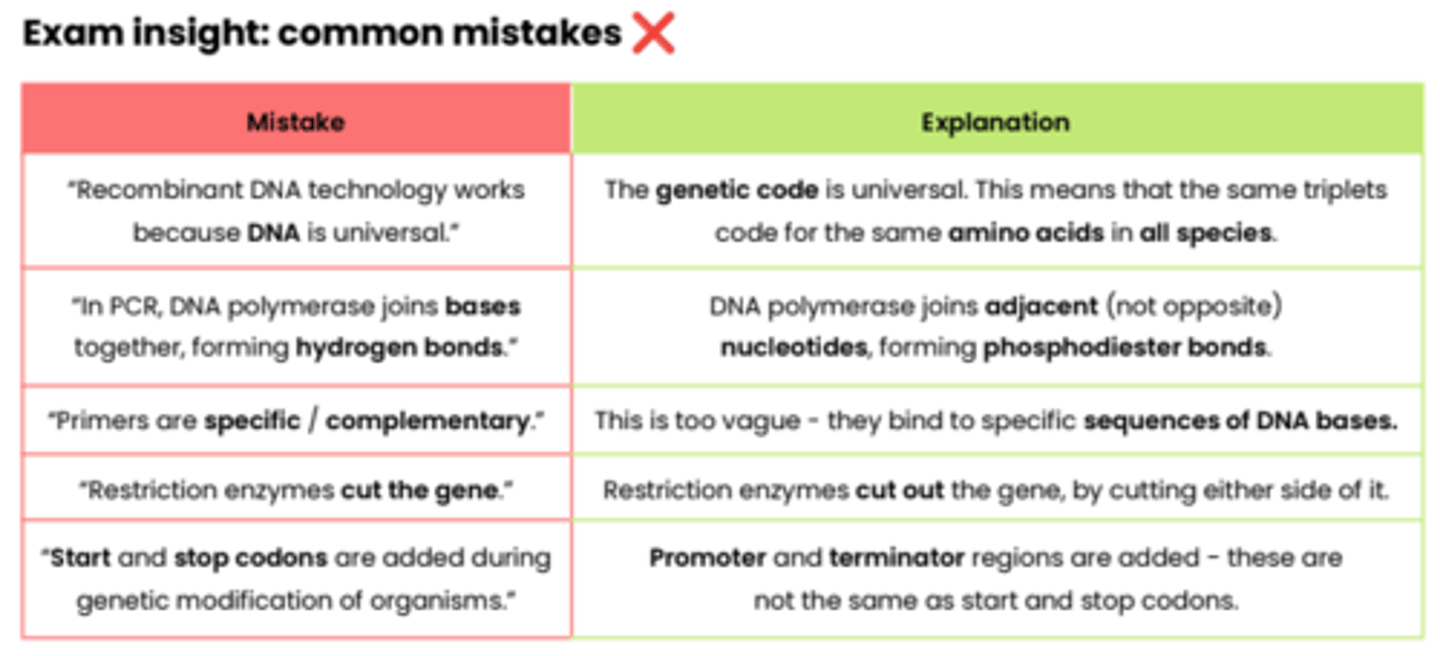

8.4.2 Differences in DNA between individuals of the same species can be exploited for identification & diagnosis of heritable conditions
What are DNA probes?
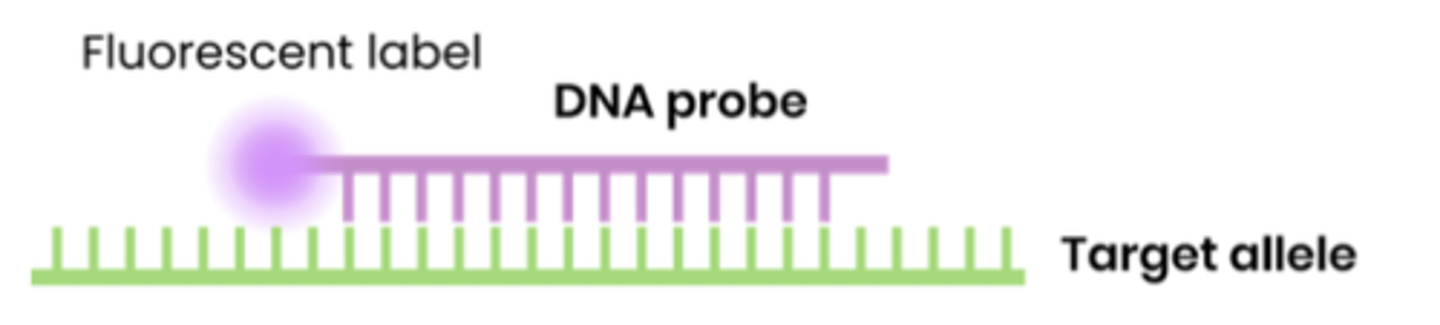
● Short, single stranded pieces of DNA with a base sequence complementary to bases on part of a specific allele / region
● Usually labelled with a fluorescent or radioactive tag for identification
Suggest why DNA probes are longer than just a few bases
● A sequence of a few bases would occur at many places throughout the genome
● Longer sequences are only likely to occur in target allele
What is DNA hybridisation?
● Binding of a single stranded DNA probe to a complementary single strand of DNA
● Forming hydrogen bonds between comp base pairs
How are DNA probes produced ?
DNA probes are made from DNA fragments.
1. Using restriction endonuclease
2. Using reverse transcriptase
3. Using a gene machine
Describe the process of DNA hybridisation
1. Multiply DNA probe using PCR
2. Heat DNA sample, to separate strands as hydrogen bonds between complementary DNA bases break
3. Add DNA probes and cool the mixture. Cooling allows new hydrogen bonds to form between DNA probe + DNA sequence
4. Wash DNA sample to remove any unbound DNA probes.
5. Check for possible attachments of DNA probe to the DNA sample. Using scanner / UV light (radioactive or fluorescent)
Why must the DNA probe first be multiplied by PCR?
The 2nd DNA strand is also complementary to the 1st DNA strand and could bind to it, preventing the DNA probes from binding to the 1st DNA strand.
Sickle cell anaemia is a genetic disorder that affects red blood cells. The disorder is due to a single mutation in the gene encoding the β globin protein in haemoglobin, as seen in the image below.
The normal allele of the gene is called A, and the mutated gene is called S.Individuals with one copy of the S allele and one copy of the A allele are called carriers, and do not show symptoms of sickle cell disease.
In order to have the symptoms of sickle cell anaemia, an individual must carry two copies of the S allele.
The fluorescent label of the DNA probe glows under a special light only when the DNA probe is attached to DNA. Explain why the DNA probe will only detect the S allele.
Suggest how the scientists could adapt their method to determine whether the individual is a carrier or whether they have two copies of the S allele.
The base sequence of the probe is complementary to the DNA of the S allele. The probe binds by forming hydrogen bonds so only the DNA of the S allele, if it is present, will be hybridised by the probe and glow under the special light.
The scientists could design another DNA probe to use on the individual's DNA sample, complementary to the A allele. If the individual's DNA sample also hybridises with the DNA probe complementary to the A allele, this means that they are a carrier. If there are no A alleles present in their DNA, it means they must carry two copies of the S allele.

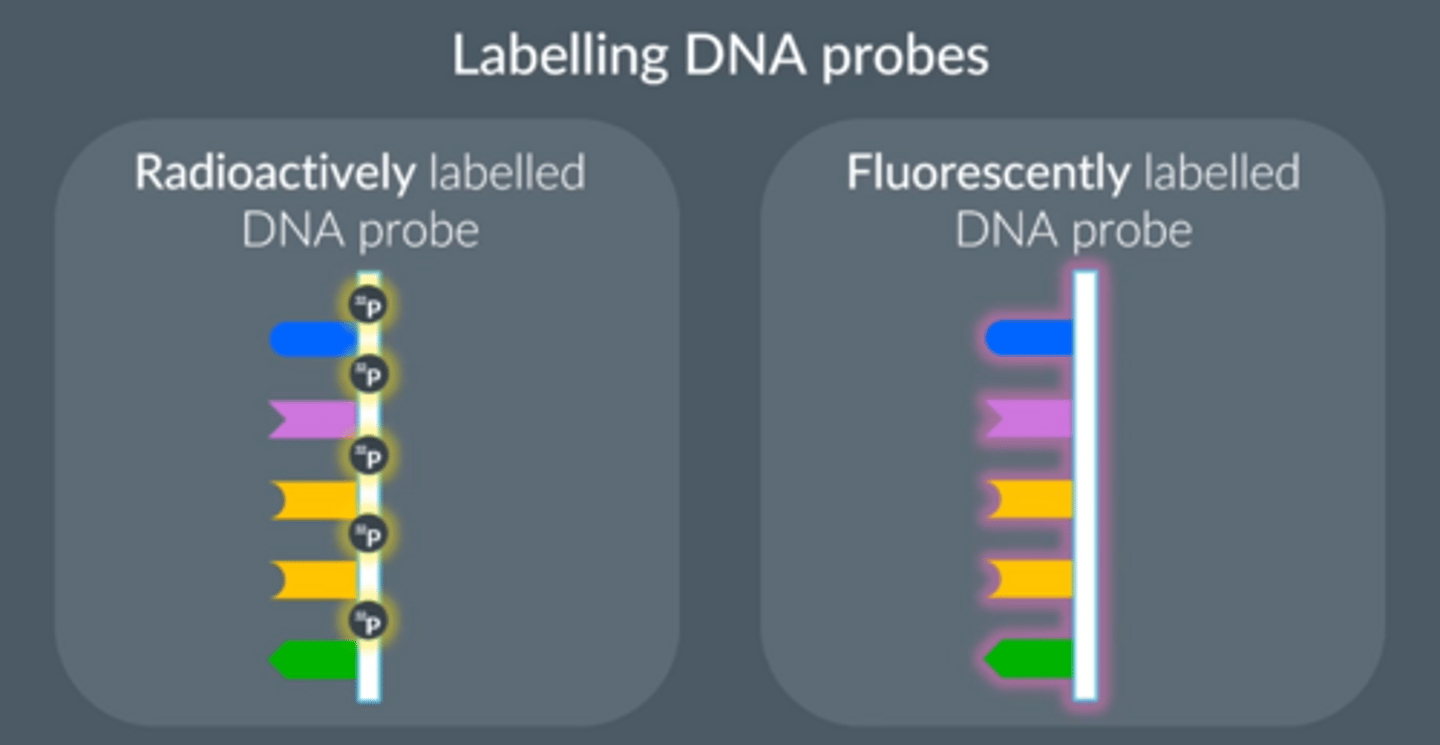
2 methods of labelling DNA probes
1. Radioactively labelled probes (radioactive form of phosphorus)
● Exposing radioactive probes to X-ray film (autoradiography)
● More dangerous
● Replace part of the probes’ molecular structure
2. Fluorescently labelled probes
● Use a special light
● Safer
Genetic fingerprinting
● Technique to compare 2 DNA samples through analysis of VNTR DNA fragments (non-coding, repeating sequences in the genome) that have been cloned by PCR.
● Creates a unique pattern that can distinguish one person from another.
(ie determine the genetic identity of an organism).
What is gel electrophoresis?
● A method used to separate nucleic acid fragments (DNA / RNA) OR proteins
● According to length/mass AND charge
Gel electrophoresis separates DNA fragments by length/mass and charge. Explain what is meant by length/mass and charge.
Length = number of bases / amino acids
Charge = DNA is negatively charged due to phosphate groups, and protein charge varies based on amino acid R groups
Gel electrophoresis
1. Add each sample to a different sample well.
2. Apply voltage to the gel.
3. This causes the DNA/RNA fragments to move down the gel, forming bands
4. Transfer the fragments to a membrane to preserve them.
5. Add a DNA / RNA probe with a stain to make the bands easier to see.
6. Take a photo to preserve the results for longer
Explain how genetic screening can be used to locate specific alleles of genes
1. Extract DNA and amplify by PCR
2. Cut DNA at specific base sequences (either side of target gene) using restriction enzymes
3. Separate DNA fragments / alleles (according to length) using gel electrophoresis
4. Transfer to a nylon membrane and treat to form single strands with exposed bases
5. Add labelled DNA probes which hybridise with target alleles (& wash to remove unbound probe)
6. To show bound probe, expose membrane to UV light if a fluorescently labelled probe was used OR
use autoradiography (expose to X-ray film) if radioactive probe was used
Gel electrophoresis separates DNA fragments by size (and charge). A DNA probe is then added, which binds to specific VNTR sequences, producing a pattern of bands which forms the genetic fingerprint.
What can this tell us?
The pattern produced is unique to each individual
Except for identical twins, who have the same pattern.
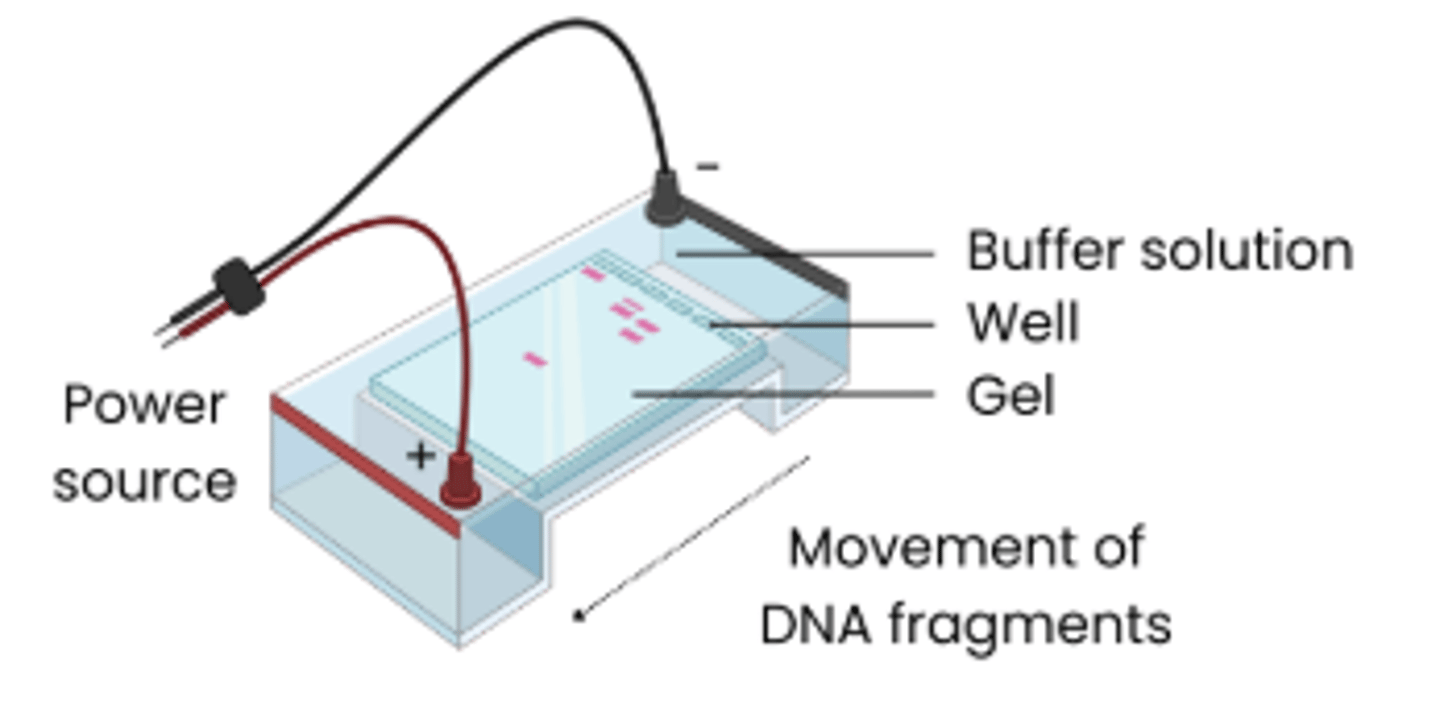
Explain how gel electrophoresis can be used to separate DNA fragments
1. DNA samples loaded into wells in a porous gel and
covered in buffer solution (which conducts electricity)
2. Electrical current passed through → DNA is negatively
charged so moves towards positive electrode
3. Shorter DNA fragments travel faster so travel further
Why do we need to do PCR before gel electrophoresis?
● PCR amplifies the DNA/RNA fragment sample so there are now many copies
● So MANY fragments of the same length build up at specific positions on the gel, forming visible bands
● Individual fragments are too small to see with the naked eye, so if we had just a few fragments on the gel, we wouldn’t be able to see anything at all.
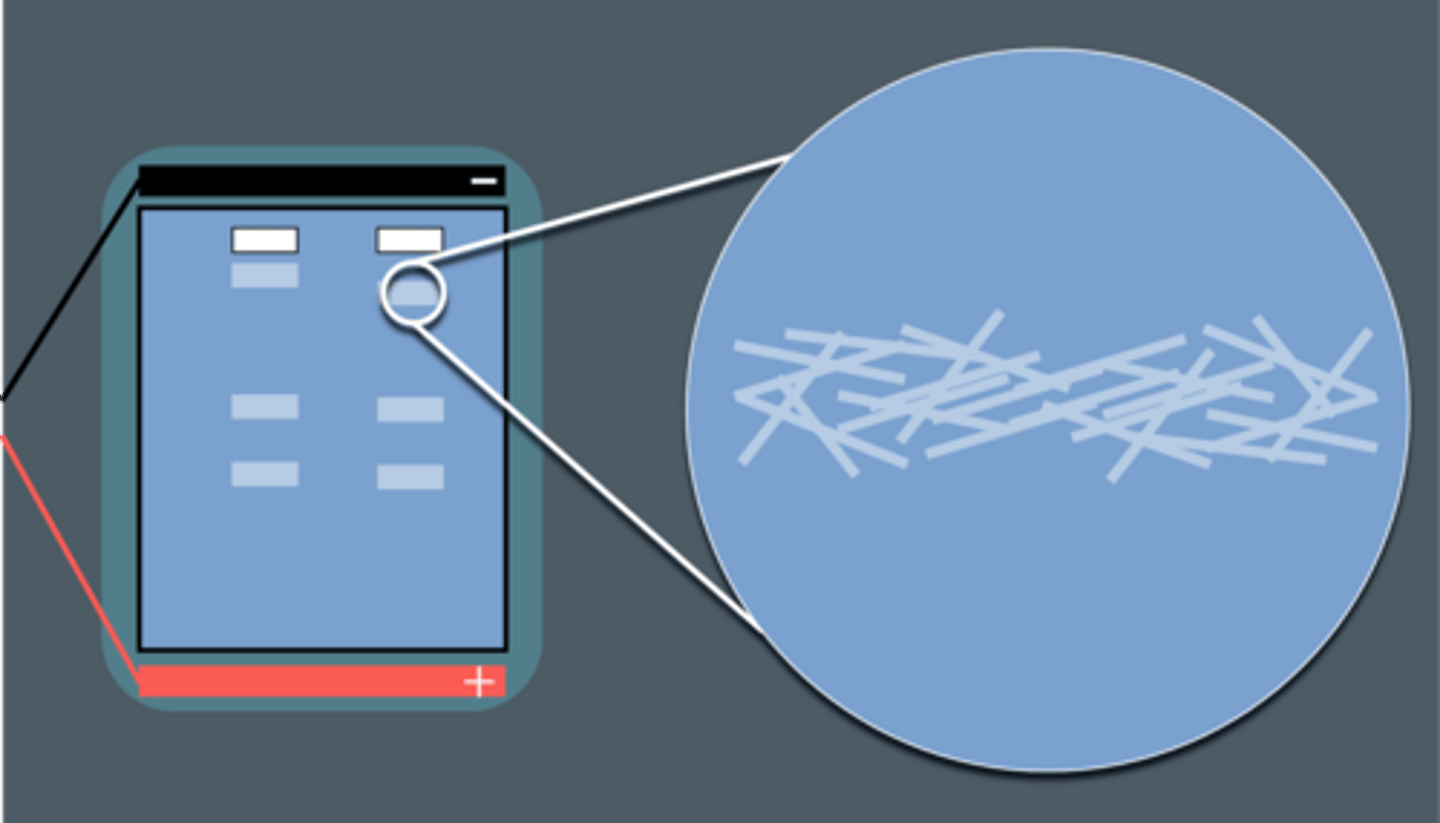
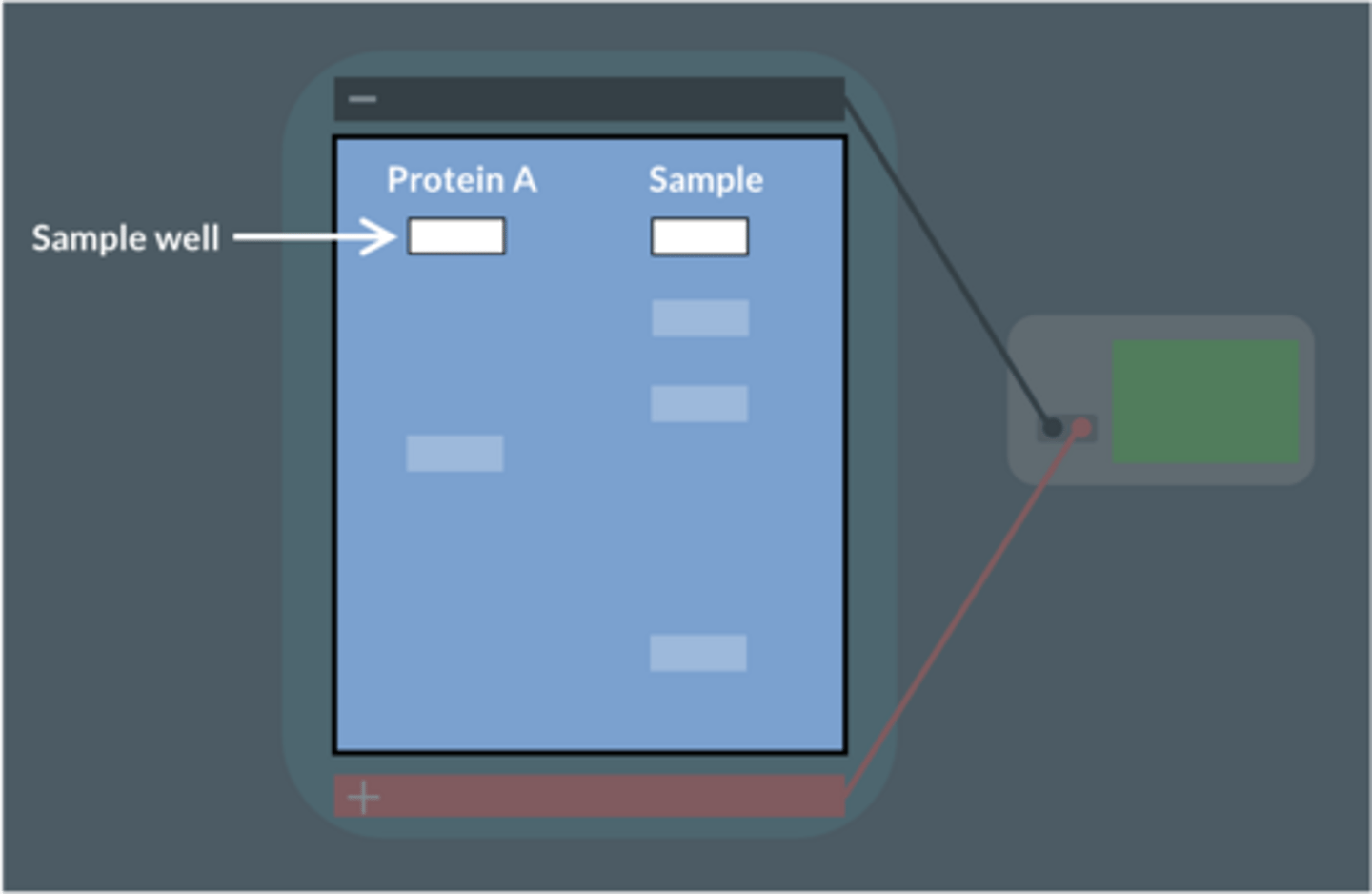
A scientist wants to find out if a sample contains a protein called 'Protein A'.
She uses gel electrophoresis to test this.
A pure sample of Protein A is placed in the first well.
The sample being tested is placed in the second well.
The results of the gel electrophoresis are shown below.
No.
The sample does not produce a band in the same position as protein A. This means protein A is not present in the sample.
How can data showing results of gel electrophoresis be interpreted?
● Run a standard with DNA fragments / proteins of known lengths under the same conditions
● Compare to position of unknown DNA fragments / proteins to estimate their size
● Shorter DNA fragments/ proteins travel further
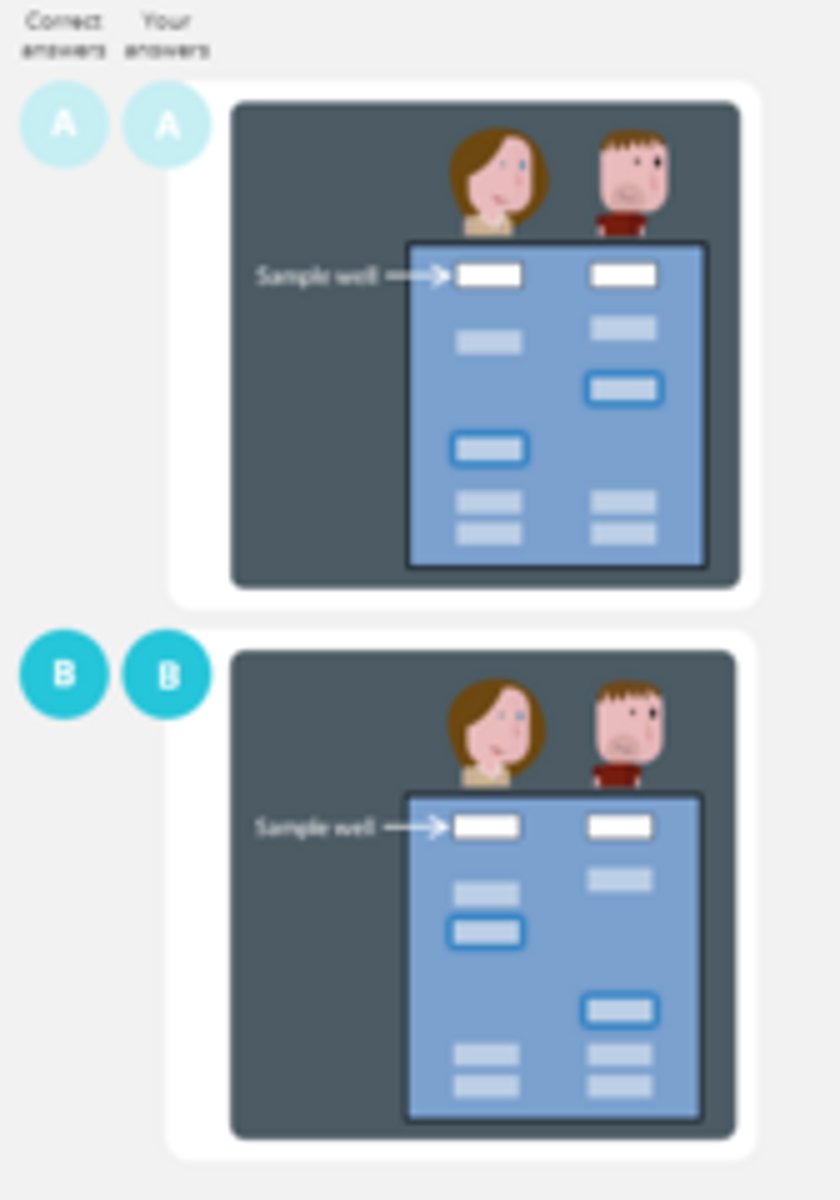
Sarah has more repeats in a particular non-coding sequence than Bob.
What are the fragments associated with Sarah's sequence be like ?
We carry out gel electrophoresis on samples from Sarah and Bob .The sequence mentioned above is highlighted blue.
Which diagram shows the correct result ?
Sarah's sequence will have more nucleotides and so longer fragments. More bands near the top as they travel slower.
What else can gel electrophoresis be used for ?
To identify particular proteins in a sample
Add your sample to a dent in the gel
Add a known sample of a protein you want to identify in a well alongside this
How are proteins separated by gel electrophoresis ?
● Length of amino acids (shorter travels faster)
● Charge - R groups can be +ve or -ve charge (greater -ve charge travels further down, as anode is usually on bottom)
Describe examples of the use of labelled DNA probes
● Screening patients for heritable conditions (e.g. cystic fibrosis)
● Screening patients for drug responses (some alleles code for enzymes involved in drug metabolism that enable better responses to certain drugs)
● Screening patients for health risks (some alleles predispose patients e.g. to high blood cholesterol)
Describe the role of a genetic counsellor
1. Explain results of genetic screening, including consequences of a positive result
2. Discuss treatments available for genetic condition
3. Discuss lifestyle choices / precautions that might reduce risk of a genetic condition developing e.g. regular screening for tumours or a mastectomy
4. Explain probability of condition / alleles being passed onto offspring → enable patients to make informed decisions about having children
What is personalised medicine?
● Medicine tailored to an individual's genotype / DNA
● Increasing effectiveness of treatment eg. by identifying the particular mutation / allele causing cancer and treating it with tailored drugs