VDJ joining
1/47
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
48 Terms
If humans dont have 10 million different genes, how can we have Ig to 10 million different epitopes?
Through several mechanisms of genetic diversity, a small set of gene segments can generate a total potential range exceeding 1011 different antibody possibilities
Describe splicing (RNA), what does this allow B cells to do?
Occurs at RNA after transcription by removing introns and joining exons to form a functional mRNA.
→It allows a B cell to make different forms of a protein from the same DNA sequence. Expressing both surface IgM or IgD or choosing between membrane-bound and secreted antibody forms
Describe recombination in B cells
Occurs at DNA level by rearranging and joining separate gene segments (variable, diversity, and joining) to create a functional gene irreversibly. Its responsible for making the vast diversity of the antibody repertoire.
What are the 3 basic stages of B cell development in a cellular level
→Begins in the bone marrow to the B cell lineage, which produces high affinity antibodies of various isotypes through class switching.
Bone marrow (antigen-independent)
Mature B cell
Peripheral lymphoid organs (antigen-dependent)
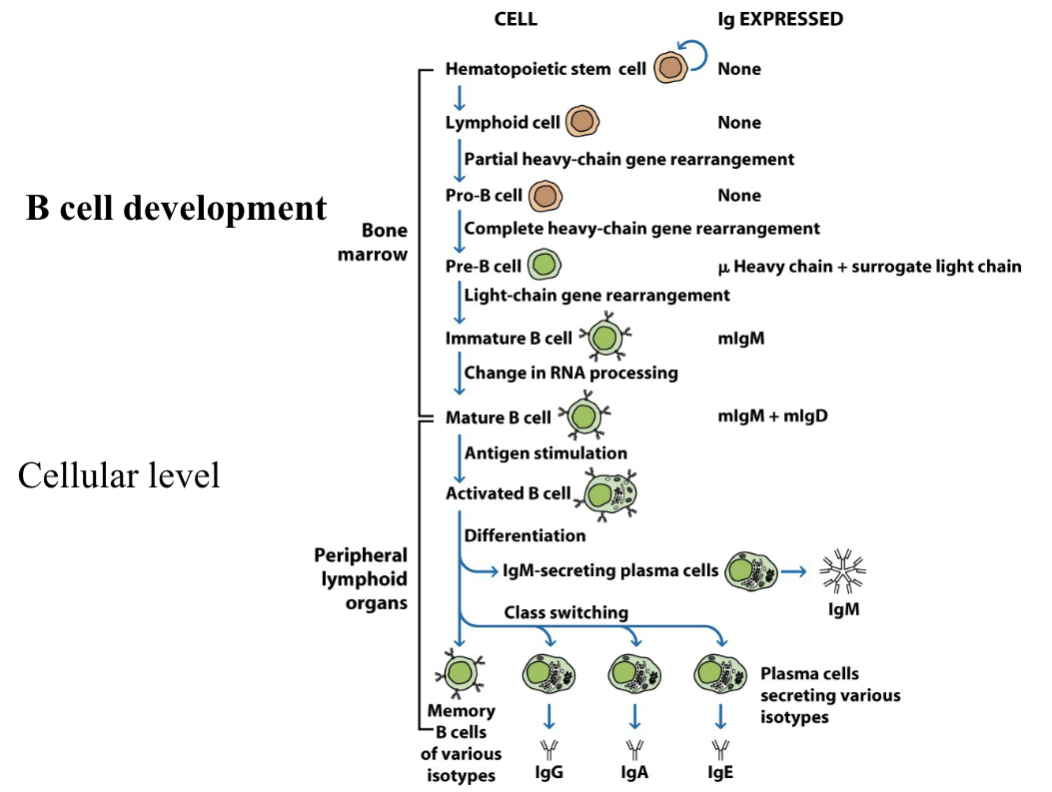
Describe the bone marrow stages (5) of B cell development (antigen-independent)
Hematopoietic stem cell: Resides in the bone marrow and express no immunoglobulin
Lymphoid cell: Resides in the bone marrow and express no immunoglobulin. Initiates the partial heavy chain gene rearrangement, joining a diversity (D) segment to a joining (J) segment.
Pro-B cell: Completes heavy chain gene rearrangement by joining a V segment to the fused DJ complex. No Ig is expressed.
Pre-B cell: They express a pre-BCR that has a μ heavy chain paired with a surrogate light chain. They begin light chain gene rearrangement (joining V to J segments), which has an intracellular μ heavy chains that cannot express Ig.
Immature B cell: Light-chain rearrangement is complete, can now express a monomeric mIgM on its surface. Undergoes a change in RNA processing (like splicing the heavy chain).
→If they recognize self-antigens, they may undergo apoptosis or receptor editing, which is the main mechanism for correcting self-reactivity.
Describe the maturation stage in B cell development
Mature B cell: Must undergo a change in RNA processing (alternative splicing) to transition to a mature naive B cell, which migrate to peripheral lymphoid organs. Once matured, the B cell can co-express mIgM and mIgD on its surface. Both receptors share the same VDJ segment and have the same antigen specificity.
Describe the peripheral lymphoid organ stages (3) of B cell development (antigen dependent)
Activated B cell: BCR binds to a specific antigen, which provides the signal for activaton/survival. Results in proliferation and differentiation.
IgM secreting plasma cells: Forms when activated B cells turns mIgM (receptor) to secreted IgM (effector) by changing mRNA of IgM heavy chain.
Class switch recombination (CSR): Mediated by AID, the VDJ segment is joined with a different antibody CH, changing from IgM to IgG, IgA, or IgE. Now secreting antibodies of the new isotype.
→Or turn into long-lived memory B cells of various isotypes, they avoid apoptosis and have high affinity for their specific antigen.
What are the 3 Murine Ig gene segments?
The murine (mouse) immunoglobulin (Ig) system has 3 distinct loci, which are specific locations on chromosomes containing clusters of gene segments.
λ Light-chain Locus (Chr 16): Has a few V and J segments paired with Cλ1, Cλ2 ,Cλ3, Cλ4 (constant) genes.
κ Light-chain Locus (Chr 6): Has 85 V segments, 5 J segments (including one pseudogene), and one Cκ gene.
Heavy-chain Locus (Chr 12): Has 134 V segments, 2 D segments, 4 J segments, and C genes (Cμ, Cδ, Cγ3, Cγ1, Cγ2b, Cγ2a, Cϵ, Cα).
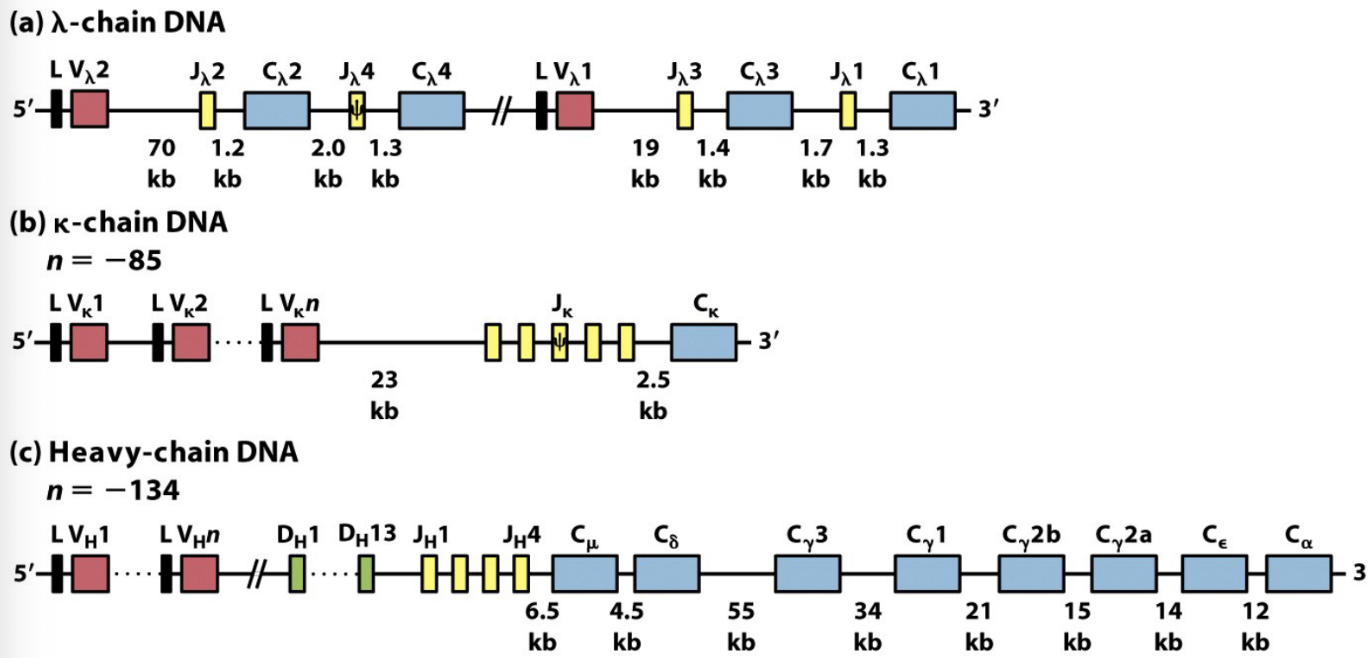
Define
ψ (Psi) = Pseudogenes:
L (Leader Sequence):
V, D, J (Variable, Diversity, Joining):
ψ (Psi) = Pseudogenes: Non-functional gene segments that cant be expressed due to genetic defects.
L (Leader Sequence): Encodes a signal peptide to guide the protein into the ER, cleaved before the antibody is matured.
V, D, J (Variable, Diversity, Joining): Separate gene segments that randomly recombine to form the unique variable region of an antibody. D segments are unique to the heavy chain.
Define
C (Constant) Genes:
Cμ,Cδ,Cγ,Cϵ,Cα:
Cγ1,Cγ2a,Cγ2b,Cγ3:
C (Constant) Genes: Determine the isotype (class) and biological effector functions of the antibody.
Cμ,Cδ,Cγ,Cϵ,Cα: These represent the constant regions for IgM, IgD, IgG, IgE, and IgA, respectively.
Cγ1,Cγ2a,Cγ2b,Cγ3: Specific constant genes for the four different subclasses of murine IgG.
Define Locus and Gene
Locus: a specific physical location on a chromosome
→ex: the H chain locus on chr 12
Gene: a specific unit within a locus or the unit made after the rearrangement of V, D, and J segments.
→ex: the Cμ gene for IgM
What are the 3 types of recombination
Combinatorial V(D)J joining
Class Switch Recombination (CSR)
Receptor Editing (Re)
Explain Class Switch Recombination (CSR)
A mechanism where the activated B cell’s VDJ is joined with a different antibody constant region (CH) of the heavy chain. This changes the antibody isotype from IgM to IgG, IgA, or IgE without altering antigen specificity. AID mediates this process to create dsDNA breaks, which are then repaired and rejoined by repair enzymes, effectively deleting the DNA between them to change the antibody’s function.
→AID targets highly repetitive Switch (S) regions located upstream of the Constant (C) region genes. The conversion of C to U in these switch regions leads to the formation of double-strand DNA breaks. Deletes DNA between two targeted switch regions, moving the VDJ segment next to a new constant region to change IgM to IgA, IgE, or IgG.
→CSR allows immune system to make antibodies suited to fight specific pathogens.
What is recombination signal sequences (RSS)
Theyre conserved DNA motifs that flank the V, D, J segments, which consist of a heptamer and monomer. The 2-turn RSS has a 23bp spacer, while the 1-turn RSS has a 12bp spacer. Ensures correct arrangement with the 12/23 rule.
→Mechanism: Theyre recognized by the RAG-1/2 complex, bringing together one 12bp spacer and one 23bp spacer, introducing nicks in the DNA to ensure correct arrangement. (HMGB1/2 stabilizes RAG binding)
Explain the productive and non-productive rearrangements of B-cells
→Productive rearrangements: Produces a functional gene sequence that leads to the expression of a working antibody. Enables B cells to express IgM on their surface and proceed with development. Has high antibody diversity in CDR3.
→Non-productive rearrangements: Produces a non-functional receptor when the addition/deletion of nucleotides causes a frameshift (nonsense) or introduces a premature stop codon.
What is the success and failure outcomes of rearragement in B cells?
The B cell has two alleles to achieve a productive rearrangement for each chain.
→Success (allelic exclusion): If the 1st allele rearranges productively, the resulting heavy-chain protein (μ) signals the cell to stop rearranging the second allele, a process called allelic exclusion.
→Failure/Apoptosis: If the first rearrangement is nonproductive, the cell attempts to rearrange the second allele. If both alleles fail to produce a functional rearrangement, the pre-B cell cant express a pre-BCR. Without survival signals provided by the receptor, the cell undergoes apoptosis.
Why is allelic exclusion important, what is it the foundation of?
It ensures the expression of only one functional receptor thats specific to one antigen. This occurs in productive rearrangement, sending a signal to inhibit/silence the rearrangement of the other chain if the first rearrangement is successful.
→This 1-cell-1-receptor rule is the foundation of the clonal selection theory.
What are the 7 mechanisms of Ig diversity
Combinatorial V(D)J joining
Junctional flexibility
P-nucleotide addition
N-nucleotide addition
Somatic hypermutation (SHM)
Ag binding domain combo
Receptor editing
Describe Combinatorial V(D)J joining
Combinatorial joining occurs in the pro-B cell (VDJ joining) and pre-B cell (VJ joining). It combines one of each segment randomly for antigen receptor diversity.
→Mechanism: RAG-1/2 complex recognizes specific RSS flanking the segments via the 12/23bp rule and introduces a double-stranded break (DSB) and non-homologous end joining as NHEJ repairs the DNA, removing/deleting intervening sequences permanently.
-Heavy chains: D to J joining occurs, follwed by V to DJ joining.
-Light chains: V to J joining occurs
Describe Junctional flexibility
Junctional flexibility is the imprecise joining between the V, D, J gene segments (occurs in pro-B and pre-B cells) to increase anitgen receptor diversity in the CDR3 region, which is the part of the V domain that makes most contact with the antigen. Its mediated by DBSK and NHEJ.
→During recombination, DSBR joins segments at varying nucleotide positions (P-nucleotide & N-nucleotide) which can cause a frameshift or change the amino acids at the junction. This can lead to productive or non-productive rearrangements to create anitbody diversity.
→involves RSS and RAG-1/2 complex
What are the 6 steps of joining mechanism by RSS in junctional flexibility
Recognition: Recognition of RSSs by RAG-1/2 complex and synapsis of 12/23bp spacers
DNA nicking: RAG-1/2 complex makes single strand nick at 5’ end of heptamers, making a 3’-OH group on coding end
Hairpin formation: 3’-OH group attacks phosphodiester bond, creating a closed DNA hairpin at the coding ends and leaving signal ends blunt.
Hairpin cleavage and P-nucleotide addition: Artemis opens the hairpin asymmetrically, resulting in single stranded overhang being filled in by Polλ using the other strand as a template, creating P-nucleotide.
N-nucleotide addition (Heavy chain only): TdT randomly adds non-templated N-nucleotides to the ends of the heavy chain gene segments to further increase diversity.
Ligation: DSBR and ligase IV join the coding ends to form a coding joint, ligating blunt signal ends to form a signal joint (usually circular piece of deleted DNA)
Describe P-nucleotide addition
It occurs in both pro-B cell and pre-B cell, because its a part of VJ and VDJ joining. Occurs when Artemis asymetrically cleaves the hairpin formed by the RAG-1/2 complex at a random, and DNA repair enzymes fill in the overhangs, resulting in P nucleotides to join, follows strand as a template.
→Resulting in many different rearrangements each time two segments are joining.
Describe N-nucleotide addition
This occurs in the pre-B cell, bc its only in VDJ joining (heavy chain). TdT randomly adds 1-12 non-templated nucleotides to the end of joints after P-nucleotide addition occurs. They add entirely new genetic information to the antibody’s binding domain.
Describe Somatic hypermutation (SHM)
This occurs in the activated B cell stage, increasing the diversity of the BCR repertoire. It leads to affinity maturation in the lymph nodes where cells with the highest binding affinity survive (clonal selection).
→Mediated by AID, which causes a single bp mutation in the DNA of the Ig variable regions which code for the antibody’s antigen-binding site.
Describe Ag binding domain combo
Its the combination of 3 CDRs from the light V chain and 3 CDRs from the heavy V chain (the epitope-binding section of an Ig). This increases the repertoire of antibody specificities.
Describe receptor editing
A tolerance mechanism where self-reactive immature B cells rearrange their light chain genes to alter their specificity, replacing self-reactive receptors with non-self-reactive ones. Allows self-reactive B cells to be rescued and become functional non-autoreactive lymphocytes, preventing autoimmunity.
→Mechanism: When an immature B cell strongly binds to a self-antigen (autoreactive), it downregulates the self-reactive receptor (instead of dying) and expresses the RAG-1/2 complex genes to rearrange (VJ) light chain gene segments. The AID enzyme causes a DSB inside the recombine variable region (VJ or VDJ). If the newly edited receptor is still self-reactive, the B cell may edit again or undergo apoptosis.
What region is greatest contributor to idiotype diversity?
The CDR3 is the greatest contributor to idiotype diversity in Ig. Its part of the V domain that makes the most contact with the antigen.
What mechanisms in the AID enzyme involved in, what does it do?
Receptor editing: Makes a double-stranded DNA break inside the recombined variable region (VJ or VDJ), bc the two C residues are across both strands.
Somatic Hypermutation (SHM): Causes single base pair mutations (C to either A, G, T) on Ig loci, altering Ig affinity.
Class Switch Recombination (CSR): Makes a double-stranded DNA break, deleting DNA between two targeted switch regions, moving the VDJ segment next to a new constant region to change IgM to IgA, IgE, or IgG.
Explain the steps of entering B-cell population to exiting B-cell population
Entering B cell population: Mature B cells are activated via antigen stimulation and enter germinal center.
The DARK zone: Activated B cells (centroblasts) undergo SHM, generating BCRs with varying affinities (high, low or nonfunctional)
The LIGHT zone: The mutated cells (centrocytes) undergo selection process, competing for limited antigens. Cells with high binding affinity survive and proliferate.
→Centrocytes with low to no affinity fail to bind, thus undergo apoptosis.
Exiting the B cell population: They undergo class switching (IgM to IgA, IgG, or IgE) to secrete antibodies or turn into memory cells.
Why is class switching not considered as diversity?
It changes the constant domains, NOT the variables. It still binds to the same epitope. This occurs late in primary response, AFTER all diversity mechanims have occured.
→IgM to IgG changes the isotype, not the idiotype
Why is there no CSR and SHM in KO mice
They dont produce the AID enzyme, which means DNA cant be deleted during recombination.
→CSR cannot occur, so they can only express IgM and IgD.
→SHM cannot occur, so their antibody affinity doesnt improve.
What are 5 facts about Re
RAG gene expression stops after successful recombination
RAG re-expression can occur to:
→Eliminate self-reactive BCR which escaped detection in bone marrow and were made by SHM
→Works with SHM in the process of affinity tuning of activated B cells (in peripheral lymphoid)
RAG is involved in VDJ and VJ recommbination, and receptor editing
Re is stimulated during B cell selection in the bone marrow in cells with autoreactive BCR
Autoreactive B cells are stimulated to undergo apoptosis or undergo Re
What are 3 facts about the AID enzyme
AID is involved in SHM, CSR and Re
S regions in the Ig locus are mutated, then CSR occurs and intervening DNA is deleted
E proteins may bind to sites near the Ig locus and direct AID to that region
What are 4 similarities betwen Ig and TCR genes
Ig & TCR genes are made from V, D, and J segments to make functional receptors.
Both mediated by RAG-1/2 complex
Both follow the 12/23 rule
Both generate variabliity in the CDR3 region
What are 4 differences betwen Ig and TCR genes
Only Ig undergoes SHM for affinity maturation and class switching
Ig antibodies can have membrane bound or secreted forms. TCRs are always membrane bound
TCR repertoire is larger than Ig’s
Ig can recognize soluble, native antigens. TCR only recognizes peptides via MHC
Provide a comparison of gene organization of Ig and TCR
Feature | Immunoglobulin (Ig) | T-Cell Receptor (TCR) |
|---|---|---|
Heavy/Primary Chain | Heavy Chain (V,D,J) | β and δ chains (V,D,J) |
Light/Secondary Chain | κ and λ chains (V,J) | α and γ chains (V,J) |
Maturation Site | Bone Marrow (B cells) | Thymus (T cells) |
What are the TCR multigene families in humans?
1. The α/β TCR Multigene Families (most common)
→α-Chain Family (Chr 14): Undergoes VJ joining
→β-Chain Family (Chr 7): Undergoes VDJ joining
The γ/δ TCR Multigene Families
→γ-Chain Family (Chr 7): Undergoes VJ joining
→δ-Chain Family (Chr 14): Theyre interspersed within the α-chain locus. Undergoes VDJ joining, the rearrangement of the α locus leads to the deletion of the δ locus on that chromosome.
Why are δ genes within the α locus?
To ensure a T cell expresses either an αβ receptor or a γδ receptor, but not both.
-The δ chains are interspersed between the Vα and Jα segments on Chr 14, the rearrangement of the α locus leads to the deletion of the δ locus on that chromosome. Enforcing the exclusion of the γδ TCR once a T cell has committed to the αβ lineage.
Explain how the αβ heterodimer is made on T cell surface
The protein heterodimer is formed by the rearrangement of the α and β chains bc theyre encoded at different genome loci.
Germ-line β-chain DNA to Rearranged DNA: Its the heavy chain equivalent bc it contains D segments. One Dβ segment randomly joins with one Jβ segment. Then a Vβ segment joins to the combined DJβ segment to form a complete VDJ rearrangement. This rearranged VDJ segment is transcribed along with a Cβ gene, RNA splicing removes the remaining introns to create the final β-chain mRNA.
Germ-line α-chain DNA to Rearranged DNA: Its the light chain equivalent bc it doesnt have D segments. A Vα segment joins a Jα segment (VJ joining), and deletes the interspersed δ genes. The rearranged VJ segment is transcribed with the Cα gene, and is spliced into the α-chain mRNA.
Protein assembly: The separate α and β chains move to the ER, which are joined by disulfide bonds to form the αβ heterodimer expressed on the T-cell surface.
Describe the H-chain primary transcript
The H-chain primary transcript is produced after the successful rearrangement of heavy-chain DNA in a B cell. In a mature, naive B cell, this transcript serves as the precursor for both IgM and IgD. It has 4 distinct poly-A sites, site 1 is used for secreted IgM while site 2 is used for mIgM.
→Coexpression: By splicing the same VDJ segment to either the Cμ or Cδ exons, they can coexpress both mIgM and mIgD on its surface simultaneously.
Why do plasma cells look like a mass of ER?
Bc theyre specialized, high-capacity antibody factories. Their primary biological function is the massive synthesis and secretion of Ig proteins. Bc of the high volume of protein being produced, the cell relies on robust ER quality control machinery.
→BiP is a major ER chaperone that binds to antibodies to ensure they fold and assemble correctly. If theyre folded incorrectly, theyre retained in the ER by BiP and exported to the cytosol for degradation.
What are the 5 Steps of the ER Quality Control Process
Nascent H & L chains are translated into the ER, where their leader sequences are cleaved.
The proteins attempt to fold and assemble into functional heterodimers. If a protein fails to fold correctly, its retained in the ER bound to BiP
The misfolded/incomplete antibody complexes are disassembled while still in the ER
Once labeled defective, they’re exported into the cytosol to be tagged with ubiquitin molecules
The ubiquitinated proteins are delivered to the proteasome to be broken down and degraded.
What is oct-1 and oct-2?
Theyre nuclear transcription factors that help regulate Ig gene expression. They recognize and bind to octamer motifs found within the promoters located upstream of each V segment. They facilitate the recruitment of the transcriptional machinery,
→Oct-1: A ubiquitous TF found in many different cell types
→Oct-2: A B cell specific TF, its presence in B cells is the reason why Ig genes are expressed in the B cell lineage and not in other tissues.
What are the 5 functional consequences of B cell activation
1. Increased Survival & Proliferation: The initial activation signals induce the expression of proteins that promote survival and cell cycling. This results in the rapid proliferation and clonal expansion of the antigen specific B cell population.
2. T-Cell Interaction: Activated B cells increase B7 expression which provide secondary signals to TH cells. They also do antigen presentation, both resulting in interactions with TH cells.
3. Responsiveness to Cytokines: Activation leads to increased expression of cytokine receptors on B cell surface, which makes the B cells responsive to cytokines (IL-2, IL-4..) secreted by TH cells, which drive B cell proliferation and their differentiation.
4. Migration to T-Cell Zones: Activated B cells increase expression of CCR7, which causes the B cells to migrate from the lymphoid follicles to T-cell zones within peripheral lymphoid organs, facilitating their interaction with TH cells.
5. Antibody Secretion: Activation generates plasma cells (antibody factories), the first antibody produced and secreted is the IgM. Signals from TH cells (w CD40 ligand) induce heavy-chain isotype switching, allowing B cell to produce antibodies with different functions for IgA, IgG, or IgE while maintaining the same antigen specificity.
Explain the role of innate immune signals in B cell activation.
Innate immune signals provide essential "second signals" that work in concert with antigen recognition to ensure robust B cell activation, particularly in response to microbial pathogens. This process involves two primary innate mechanisms: the complement system and Toll-like receptors (TLRs).
Activation via the Complement System: Microbe activates alternative complement pathway and C3d attaches to the microbes surface. B cell binds to microbial antigen through its BCR while binding the attached C3d to its CR2 receptor (recognizes tagged microbes). The CR2 brings a signaling complex close to the BCR to initiate activation.
Activation via TLRs and PAMPs: BCR binds to specific antigen on microbe while PAMPs bind to the TLRs on the B cell. The TLRs trigger internal activating signals that synergize with the BCR signals, The signals from BCR and TLR triggers B cell to undergo proliferation, differentiation, and Ig secretion.
Explain the mechanism of TH cell-mediated activation of B cells (3 steps)
TH cells recognized peptide antigens presented by B cells.
The TH cells are activated to express CD40L ligand & secrete cytokines, binding to and activating the B cells.
The B cells can now proliferate and differentiation into plasma cells or memory cells.
Provide differences between Primary response and Secondary response
Features: Lag phase, Peak response, Primary isotype, Avg affinity, longevity
Feature | Primary Response | Secondary Response |
|---|---|---|
Lag Phase | 5–10 days | 1–3 days |
Peak Response | Smaller | Larger |
Primary Isotype | Mainly IgM | Relative increase in IgG |
Average Affinity | Lower | Higher (Affinity Maturation) |
Longevity | Short-term (weeks) | Long-term (years) |
→Primary response: Naive B cells & T-independent
→Secondary response: Memory B cells & T dependent
What is the resolution phase? Provide mechanism
The resolution phase of a humoral immune response is regulated through antibody feedback, which terminates B cell activation after the infection has cleared.
→Mechanism: IgG antibodies bind to Fc receptor, which contain ITIMs. These ITIMs recruit enzymes to block activating signals from the BCRs ITAMs. Terminating B cell activation and antibody production.