D1.1
1/17
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
18 Terms
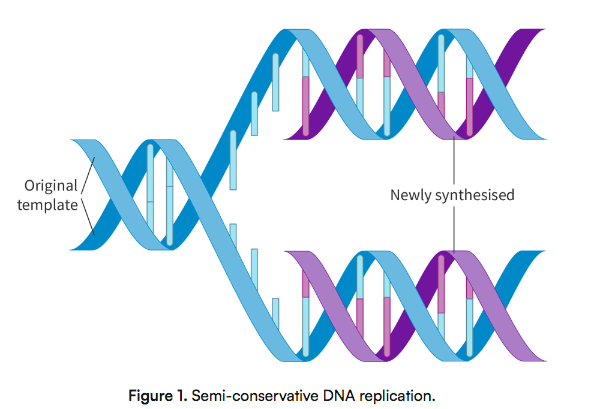
What does the semiconservative process of DNA replication mean? Why is it important?
Each new double stranded DNA has 1 strand OG DNA strand + 1 strand new DNA
Occurs because each strand of OG DNA is a template for new DNA
Allows a high degree in accuracy when coping base sequences and stability in passing down code
Define and summarize PCR and its materials needed
Def: a technique used to amplify small DNA fragments
Used to help clone genes, use crime DNA, sequence extinct DNA
Desired DNA is put into a reaction chamber, involving:
free nucleoside triphosphates, primers (single stranded, short polymers of 15-20 nucleotides that are complementary to nucleotides at the target DNA), taq polymerase
thermocycler
List the 3 steps in PCR
Denaturation
DNA is heated to break H-bonds (98C)
annealing
temp is lowered (60C) to allow primers to bind to complementary sequences in the DNA sample
extension
taq polymerase replicates DNA using primers as the starting point (72C)
Why are primers used in PCR?
Serves as a starting point for replication since DNA polymerases can’t add the first nucleotide but extend
Summarize gel electrophoresis and its purpose/applications
Uses electricity to move molecules through a semisolid medium to separate DNA by size + charge
helps identify key features of DNA
DNA profiling
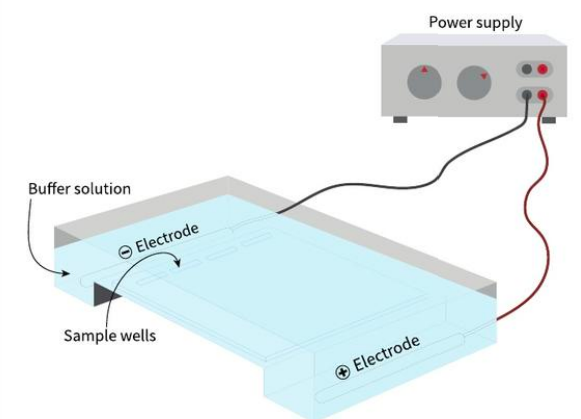
List 3 key steps in gel electrophoresis
to get proper sizes of DNA, its digested using restriction endonucleases (enzymes)
this cuts the backbone at specific sequences
samples of DNA fragments are loaded into wells at one end of gel
gel is soaked in buffer solution
electric current runs through gel
DNA moves to the positive side (since DNA is negatively charged)
Due to the porous gel, smaller and most charged DNA pieces travel further
Length of sample DNA fragments can be found by comparing it against DNA ladder
What substance is used to see DNA in gel electrophoresis
Ethidium bromide to help it fluoresce
What contributes to directionality of DNA replication?
The fact that phosphodiester bonds (occurring between phosphate group on 5’C and hydroxyl group on 3’C) are holding nucleotides
Which direction is new DNA built? Why?
From 5’ to 3’
Since DNA polymerase III has an active site complementary to a specific shape
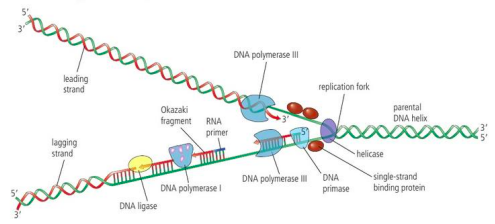
What is the lagging vs. leading strand? Why does this exist?
Both refer to the NEW DNA strands formed
Leading (the template strand is 3’ to 5’)
the strand replicated in the same direction as helicase moves
formed in continuous way (5’ to 3’)
thus, primer and primase are only needed once
formed quickly
Lagging
Doesn’t allow DNA polymerase III to move in same direction as helicase
Moves away from replication fork
is synthesized discontinuously in okazaki fragments, formed slowly
Thus, RNA primer needs to be added repeatedly
requires DNA ligase
This exists due to the antiparallel nature of DNA
List the 6 enzymes of DNA replication
Helicase
unwinds/unzips DNA by breaking H-bonds (moves one complementary base pair at a time)
forms a replication fork
Works with single strand binding proteins, preventing the strand from coming back together
Gyrase
Moves ahead of helicase
relieves the tension created by the unzipping of double helix
DNA primase
attaches RNA primers to the template strand
without it: DNA polymerase III can’t attach to DNA strand
allows DNA polymerase III to later assemble free nucleotides into a new DNA strand
For leading strand: 1 primer needed
For Lagging strand: primers need to be placed at regular intervals
DNA polymerase III
places free nucleotides in correct sequence based on complementary base pairing rule
replicates continuously on leading strand, discontinuously on lagging strand
DNA polymerase I
removes RNA nucleotides of primers and replaces with DNA nucleotides
DNA ligase
forms phosphodiester bonds between okazaki fragments
Allows the lagging strand to form one single strand
Which enzyme is needed for DNA proofreading?
DNA polymerase III (its additional function)
It proofreads newly formed DNA strand as its being built
it replaces incorrect nucleotides
What is each nucleotide made of
Deoxyribose (5C sugar)
phosphate group
nitrogenous base
What are 2 major molecules found in nucleus important for DNA replication?
Enzymes
free nucleotides (freely floating in nucleoplasm, like adenine, thymine, etc)
Describe the steps for continuous and discontinuous replication
Continuous
DNA primase adds RNA primer
DNA polymerase III adds nucleotides to 3’ end
DNA polymerase I replaces primer with nucleotides
discontinuous
DNA primase adds RNA primer in front of 5’ end
DNA polymerase III adds nucleotides
DNA polymerase replaces primer
DNA ligase attaches Okazaki fragment ot the lagging strand
List purposes of DNA primers in PCR
primers attach to the separated strands of DNA
primers prevent hydrogen bonds between the separated bases from reforming OR prevent original helix from recombining
provide a starting point for replication as Taq polymerase attaches the first nucleotide to the 3'-end of the primer
primers can be designed to target specific sequences in the genome
Outline how DNA profiling works
a. sample of DNA obtained from person/hair/blood/mouth/crime scene ✔
b. PCR used to amplify/make copies of DNA (in sample) ✔
c. using Taq DNA polymerase / using DNA polymerase from thermophilic bacteria ✔
d. tandem repeats amplified/used ✔
e. gel electrophoresis used to separate DNA (into bands) ✔
f. separation according to length of fragments/number of repeats
OR
fragments of same length/number of repeats travel same distance ✔
g. pattern of bands/numbers of repeats is the profile/is unique to the individual ✔
h. example of application/forensics/crime investigation/paternity ✔
how are tandem repeats used in DNA profiling
tandem repeats (at one locus) vary in number of times sequence repeats / represent different alleles for one locus;
DNA sample cut by restriction enzymes into fragments;
samples of DNA are amplified at specific genetic sites with PCR;
the fragments are separated by their size/number of repeats with gel electrophoresis;
fluorescent/radioactive label attached to different tandem repeats;
data from several loci at one time uniquely identify individuals / like a fingerprint, combinations of alleles are specific to an individual;
comparisons/similarities between fragment patterns to determine paternity/evidence match to a suspect’s profile / other example of comparison/similarity;