Biology - D1.2 Protein Synthesis
1/40
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
41 Terms
Two key stages of protein synthesis
Transcription and translation
Triplet code
3 bases code for one specific amino acid
Transcription
Synthesising RNA using a DNA template
Why is transcription important?
Proteins are made by the ribosomes outside of the nucleus but DNA, which codes for the proteins, can’t leave the nucleus so messenger RNA is a transporter of the code.
Gene expression and transcription
If a protein is needed by the cell, the gene for it is switched on and the gene is being expressed. The switching on of the gene is necessary for transcription as it allows the enzyme RNA polymerase to bind to the DNA.
Function of RNA polymerase
Binds at the promoter region and makes a single strand of mRNA.
RNA polymerase adds RNA nucleotides temporarily to the template strand of the unwound DNA using complementary base pairing.
Antisense vs sense strand in transcription
The antisense strand is the template strand, where the mRNA is made from.
The sense strand is the coding strand, it is not the template, it has the same code as the new mRNA strand (with uracil rather than thymine).
Importance of rewinding DNA after transcription
DNA is only unwound for a short period of time so that it is minimally vulnerable to mutations. DNA should be stable and unchanged for the life of the cell.
E.g. in somatic cells, body cells that never divide, nerve cells
6 steps of transcription
1) Gene is expressed and identified
2) RNA polymerase binds
3) The gene is unwound by the RNA
4) RNA polymerase adds RNA nucleotides temporarily to the template strand using complementary base pairings
5) RNA polymerase encounters a sequence that indicates the end of the gene (stop codon) and the mRNA strand is released
6) mRNA leaves the nucleus through a nuclear pore and the DNA is wound back up
4 steps of translation (SL)
1) mRNA attaches to the small subunit of a ribosome
2) The start of translation or initiation (1st and 2nd tRNA with a formation of 1 peptide bond)
4) Making of a polypeptide / elongation phase of translation
5) Completion of polypeptide / termination phase of translation
tRNA
Special transfer RNA molecules (tRNA) in the cytoplasm have 3 bases sticking out, forming an anticodon. On the other side, tRNA temporarily binds to an amino acid.
The amino acid the tRNA carries depends on its anticodon; there are 64 different tRNA molecules with the bases that differ and the amino acids that can be repeated.
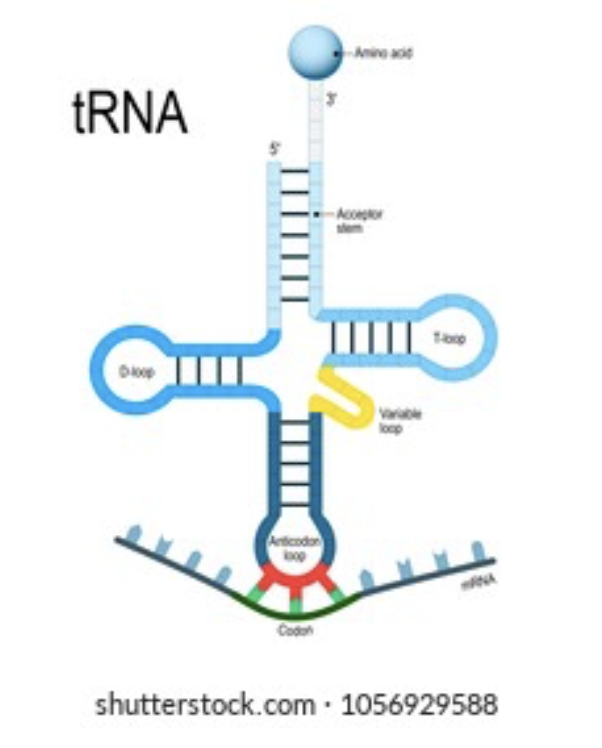
Anticodon
The three bases on a tRNA molecule which are complementary to the coding mRNA
Translation definition
The synthesis of polypeptides from mRNA
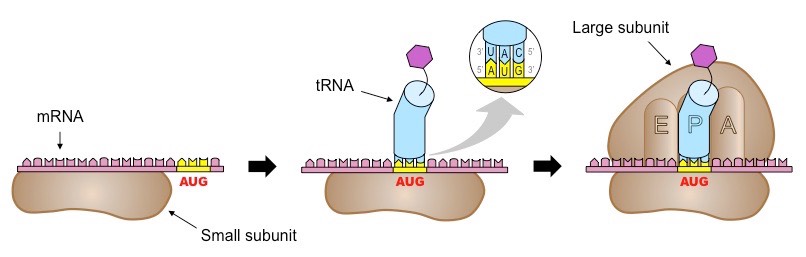
The start of translation
The ribosome, with the initiator tRNA already attached, moves along until it reaches a start codon
The initiator tRNA with complementary bases to the start codon (anticodon) binds to the middle tRNA binding site in the large subunit.
A second tRNA attaches to the adjacent binding site.
The transported amino acids are positioned so that a peptide bond forms between them
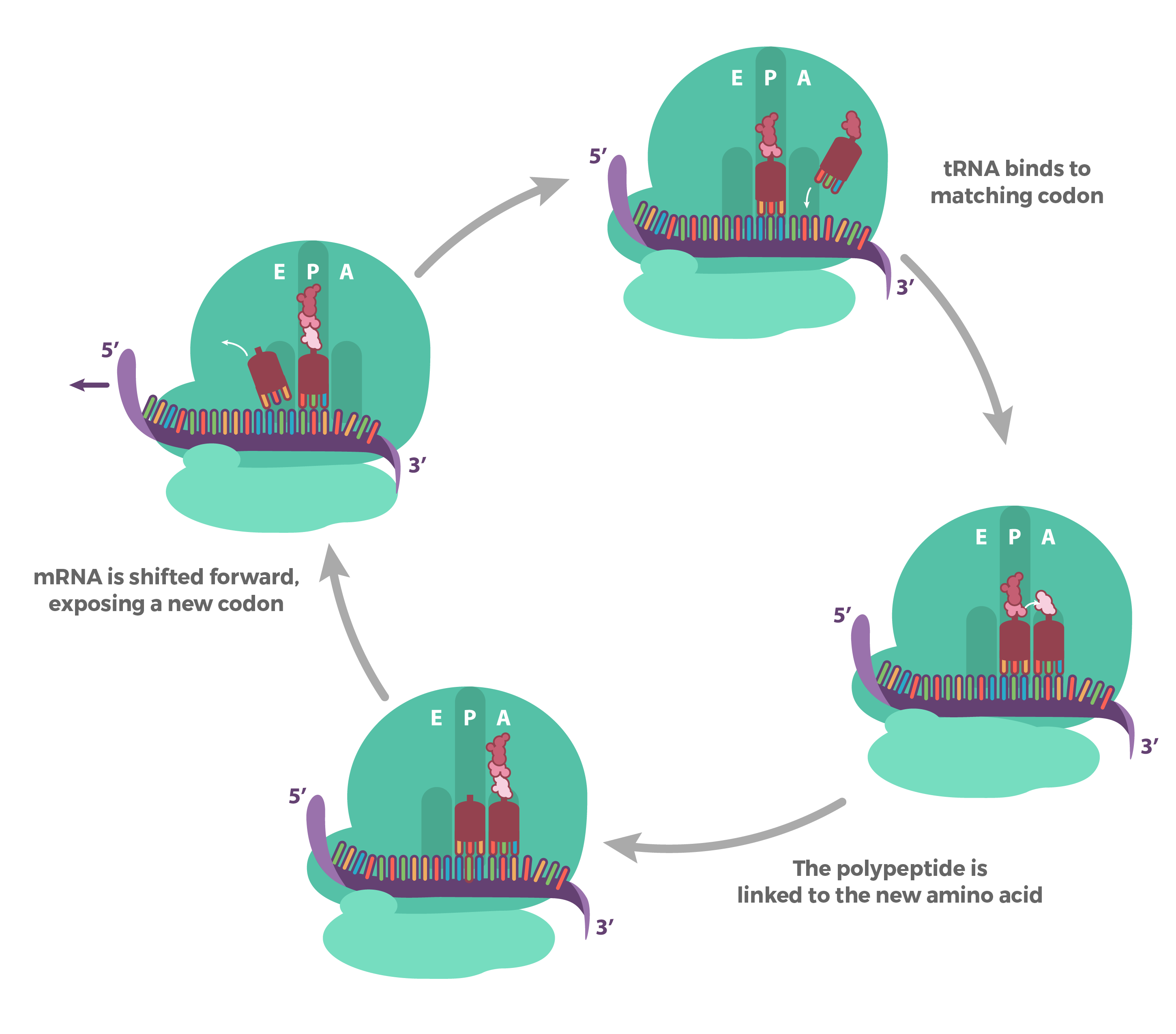
Making of a polypeptide / elongation phase of translation
Once the peptide bond has been formed, the ribosome moves to the next codon and the first tRNA is released from the E binding site.
The next tRNA molecule with the correct anticodon joins and a peptide bond is formed between the dipeptide and new amino acid.
REPEAT
Completion of polypeptide/ termination stage of translation
The ribosome encounters a stop codon and the polypeptide is released from the ribosome
The polypeptide can interact with the environment outside of the ribosome and coil up and finished off/packaged in the Golgi apparatus
The mRNA can be used again or broken up into its nucleotides
What strand is used to figure out the code for the amino acids?
the mRNA sequence / the coding/sense strand, the only difference is the uracil instead of thymine.
Degeneracy
The aspect of the code where many codons can code for the same amino acid
Universal genetic code
The code coding for amino acids is the same for all living organisms
Gene mutation
A change to the base sequence of a gene
Effect of a single base mutation
A change in one base can be drastic/large effect as it can result in a different amino acid being coded for
On the other hand, the change may not be too significant as the code is degenerate
Sicke cell anaemia
Is an example of an inherited point mutation. It is a single base substitution in the 6th codon, GAG to GTG, no longer glutamic acid instead now have valine.
This mutation cause the haemoglobin to link together in chains at low oxygen levels. This means the blood is trapped in capillaries and the red blood cells life's are shortened by 4 day, no more replication and thus anaemia.
Directionality of mRNA transcription
The nucleotides are added to the mRNA strand in a 5’ to 3’ direction. So the next nucleotide is added at the sugar
Promoter region
Important section of DNA (between 100 ad 1000 bases long) that is normally located just before a gene and has a base sequence that allows RNA polymerase to bind to the DNA.
Promoters differ slightly for each gene but all have a TATA box
Transcription factors definition
Many proteins inhibiting or allowing RNA polymerase to bind. Within or associated with the promoter region. Influence gene expression.
Enhancer sequence
AKA Activator sequence, as it allows transcription factors such as activator proteins to bind to the sequence. This switches the gene on, allowing or encouraging the RNA polymerase to bind to the promoter region.
Silencer sequence
AKA Repressor sequence as it allows transcription factors such as repressor proteins to bind which prevent the binding of RNA polymerase and switch the gene off temporarily.
Non-coding sequences
Do not directly code for polypeptides and make up 98%-99% of DNA
Base sequences involved in regulation of gene expression e.g. enhancers and silencers
Base sequences coding for tRNA and rRNA (ribosomal RNA)
Telomeres
Introns
Introns
Base sequences that are transcribed but edited out of the mRNA during post-transcriptional modification. They are sections of mRNA usually between 20 and 200 nucleotides long.
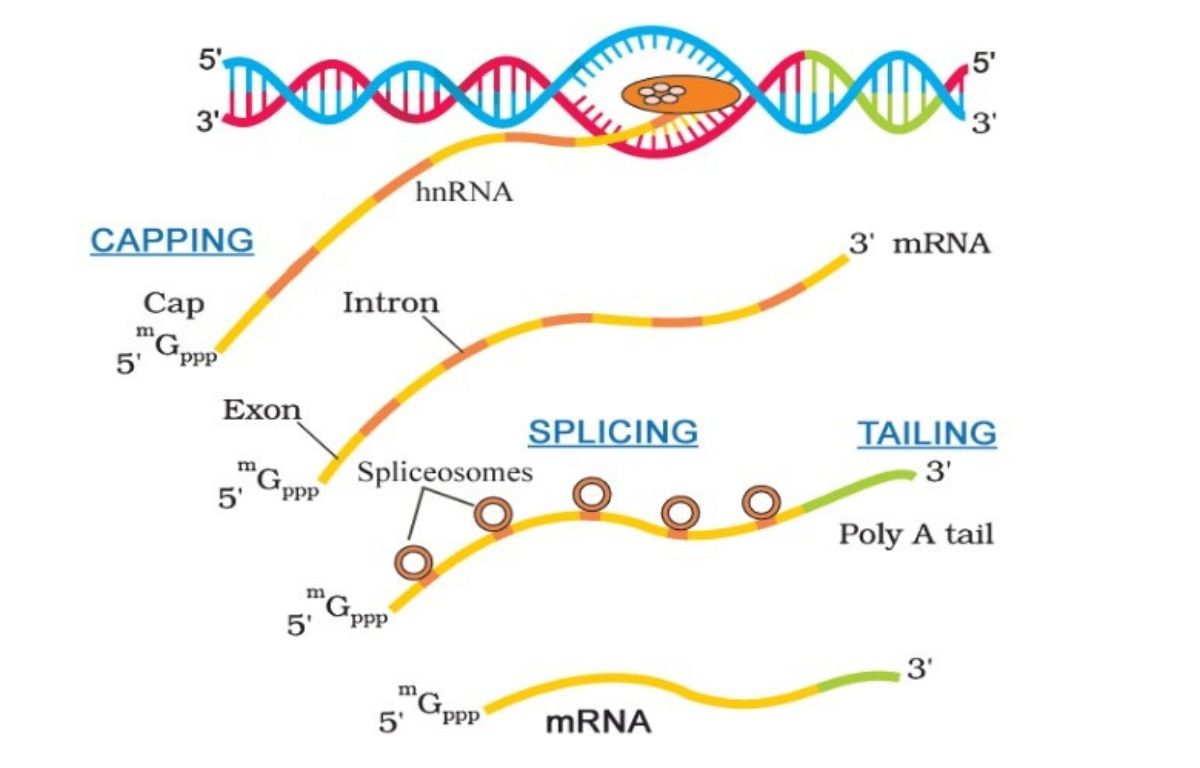
List of post-transcriptional modifications
Adding a 5’ cap
Adding a poly A tail
Removal of introns
After, the mRNA is ‘mature’.
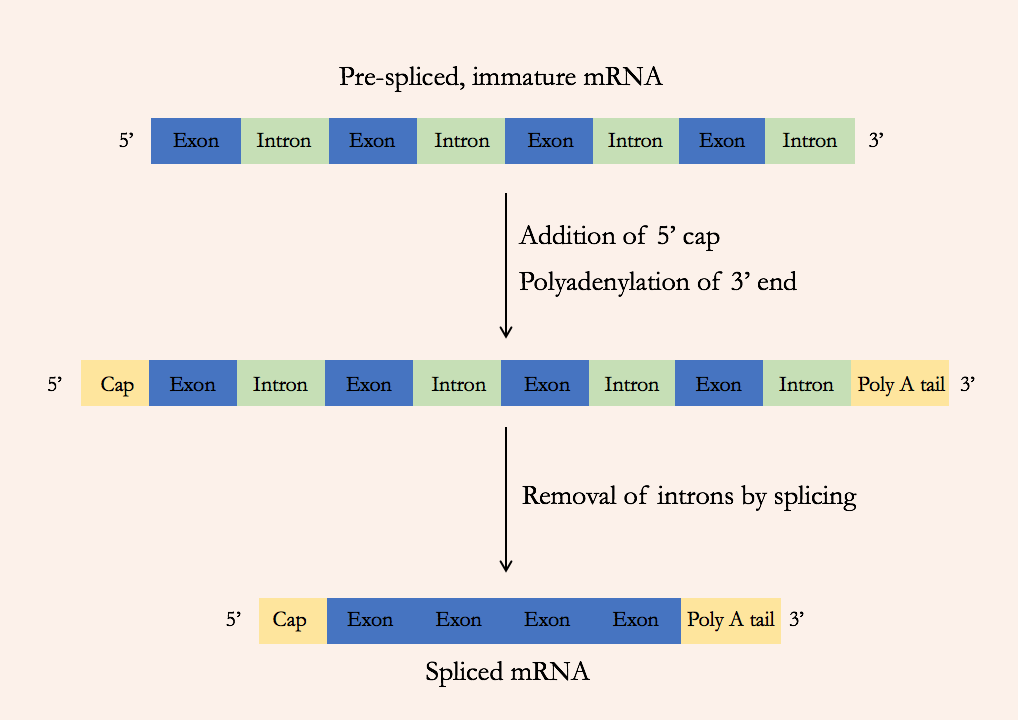
Post-transcriptional modifications definition and function
Once transcription has taken place, the mRNA formed is modified before it leaves he nucleus.
This is done to stabilise the transcripts and protect the mRNA and encourage removal of introns.
5’ cap addition as post-transcriptional modification
A modified guanine nucleotide with three phosphates is added to the 5’ end of the mRNA
Addition of Poly A Tail as post-transcriptional modification
Between 100-200 adenine nucleotides are added to the 3’ end of the mRNA
Removal of introns as post-transcriptional modification
The introns between the exons in the initial mRNA are cut out using spliceosomes. The exons are spliced back together, potentially in a different order depending on the protein.
Alternative splicing
The order in which the exons are spliced back together can vary, and sometimes some are even omitted. Useful in diversity and storage, as one gene can code for different polypeptides.
Three binding sites in the large subunit of a ribosome
the 3 binding sites are: A, P and E.
P is where the initiator tRNA binds
A is where the second or following tRNA binds
E or the exit site is where they exit, a tRNA molecule never occupies it
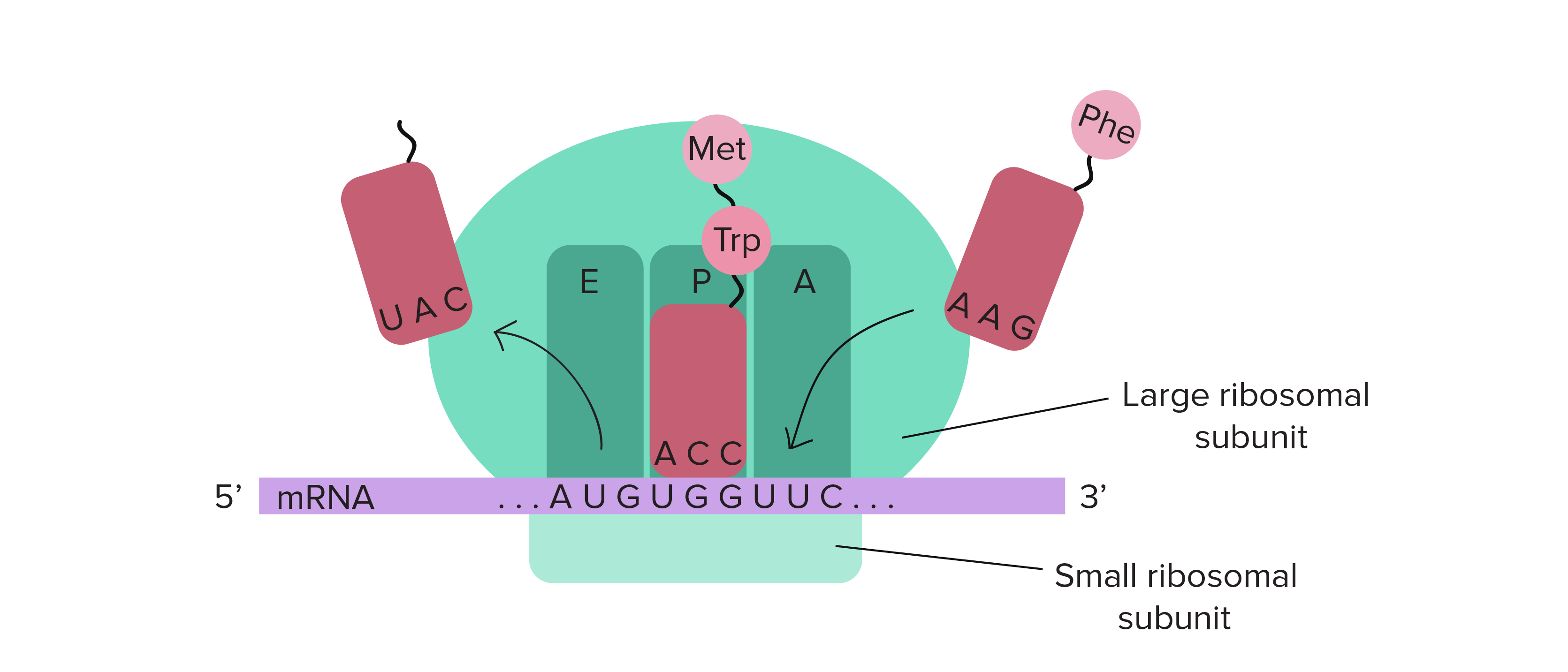
Initiator tRNA
Initiator tRNA is the first tRNA with an anticodon that is complementary to the start codon. It temporarily binds to the amino acid methionine. It binds to the A site in the ribosome before the mRNA binds to the smaller subunit of the ribosome.
Modifications of polypeptides into their functional state
Folding up, also while being made (secondary structure & tertiary structure)
Addition of another polypeptide or a prosthetic group (quaternary structure)
Cutting a polypeptide up (propeptide into mature peptide)
Removal of amino acids
Changes to the R-groups of amino acids
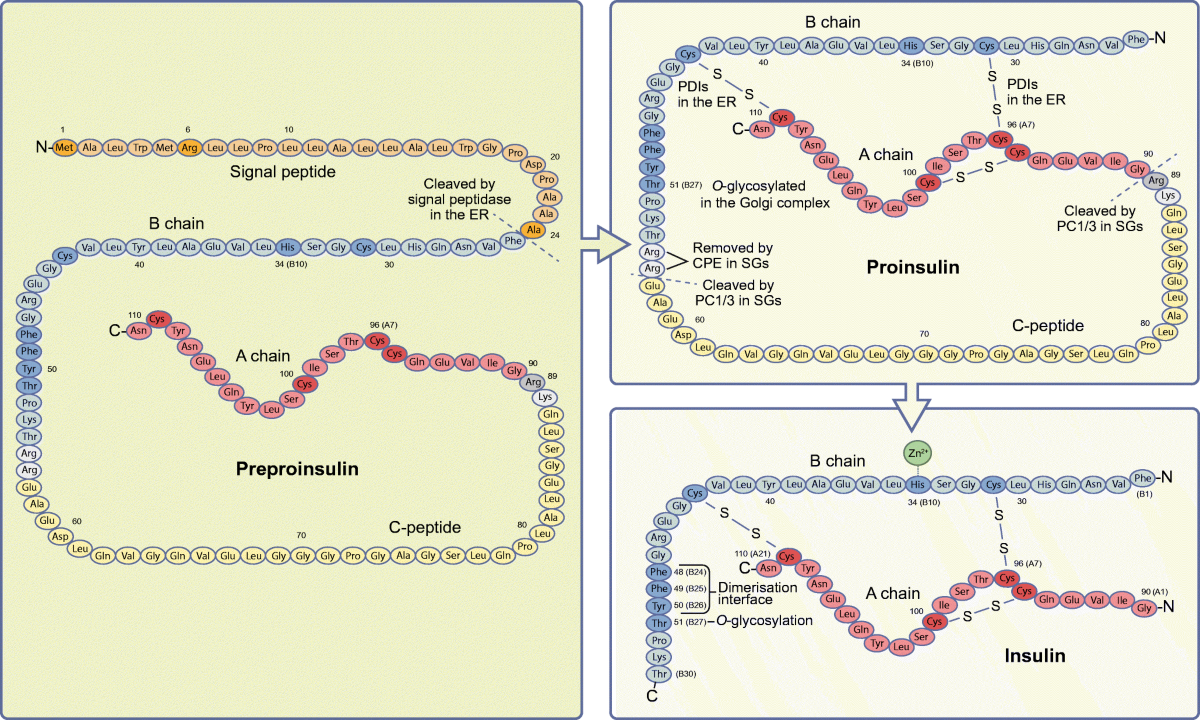
Two-stage modification fo pre-proinsulin into insulin
1) To make proinsulin: signal peptide is removed by an enzyme
2) To make insulin: C-peptide is cut out, 3 disulfide bonds are formed to make the tertiary structure, lastly 2 amino acids (arginines) are removed from the B chain.
Process of recycling of proteins using proteasomes
1) The damaged or now useless proteins are identified and tagged with a small protein called ubiquitin.
2) Proteasomes recognise ubiquitin and unfold the protein and feed the protein into its core
3) In the core the multiple protease enzymes can break the protein down into smaller chains of amino acids.
4) The smaller chains exit into the cytoplasm and are broken down further there into individual amino acids.
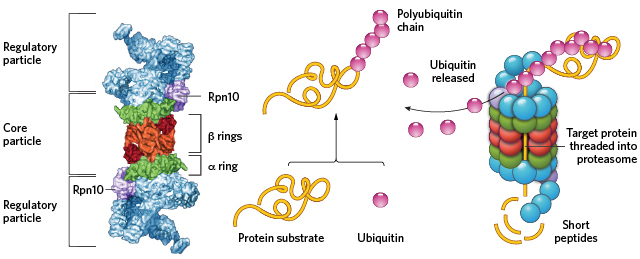
Proteasome
Proteasomes recognise proteins tagged by ubiquitin and feed it into their core where the proteins are broken down into smaller chains of amino acids by the protease enzymes in the core.