Nicotinic Acetylcholine Receptor
1/27
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
28 Terms
Cholinergic Systems
Acetylcholine binds to its receptor which is either a nicotinic or muscarinic Ach receptor which have differential paharmacology
Nicotinic receptors activated by nicotine, muscarinic receptors activated by muscarine
Alpha-bungarotoxin is an inhibitor of the nicotinic Ah receptor, atropine binds to the muscarinic receptors and acts as an inhibitor
Nicotinic Rs are ionotrophic found primarily in the sympathetic and parasympathetic NS, allowing them to have a rapid response time (microsecond/millisecond)
Muscarinic Rs are G protein coupled receptors found in the parasympathetic NS and act a lot slower (milliseconds to seconds)
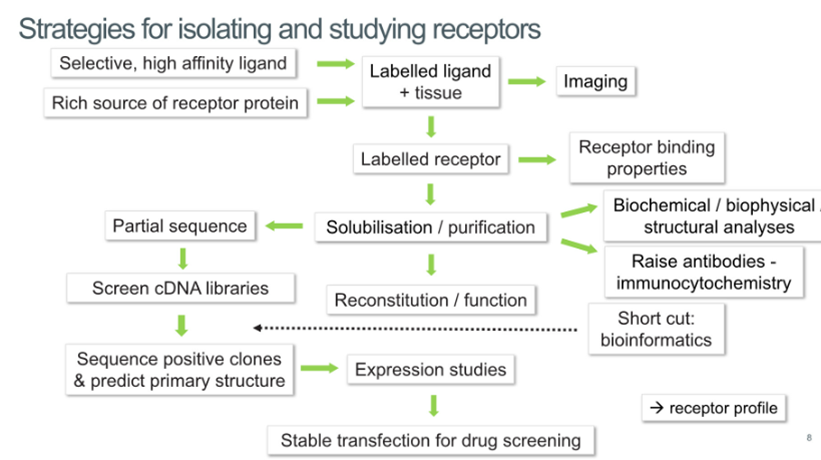
Strategies for isolating and studying receptors
In history receptors needed to be abundant to study which is why these receptors are good targets
Receptors initially defined by their pharmacology, so people able to identify high affinity ligands which can help them purify them
Need to find a rich source of receptor which allows you to label ligand with radioisotopes, label the tissue that the receptor is present in which allows you to localise it in the body
Once the receptor has been labelled allows you to purify it and assess the binding properties
Purification of the receptor allows you to understand the molecular change, create agonists for the receptor and to obtain a partial sequence
Nicotinic Ach receptor isolation
Can find a rich source of protein in electric fish organs called torpedo nobiliana (can obtain several bands)
Has this electroplax tissue on the back of its head which allows it to shock its prey
Can very carefully stimulate this tissue and homogenise the tissue then load it onto a sucrose gradient
Layers of different concentrations of sucrose, as you go down the tube the concentration of sucrose increases
Samples placed on top and spun in a centrifuge and the receptor membranes form a band partway down the tube depending on the amount of protein in the sample
Allows you to obstain bands of receptor membrane
Can take the receptor fraction and run it on an SDS gel which will give us bands for the alpha, beta, gamma and delta subunits → expect a thick band of alpha as the receptor comprised of 2 alpha subunits, one beta, one gamma and one delta
Protein based sequence
Before we could sequence the whole genome, we could obtain a sequence of the protein via Edman degradation
Involves labelling the AA at the N terminus with a colourful molecule, then cleave it off and work out what it is, then cleave the next amino acid to resolve the N terminal sequence
Now, instead of this technique we would excise the protein from the gel and do mass spectrometry analysis which involves fragmenting the protein and comparing the mass spectrometry to a database
If we know what the protein sequence then we can work backwards and work out the DNA sequence
From partial agonist to full sequence
The sequence of Aas can be mapped across to the different codon possibilities
We can then screen cDNA libraries from humans and sequence the positive clones
This allows us to pull out the sequence for each of the NicAchR subunit
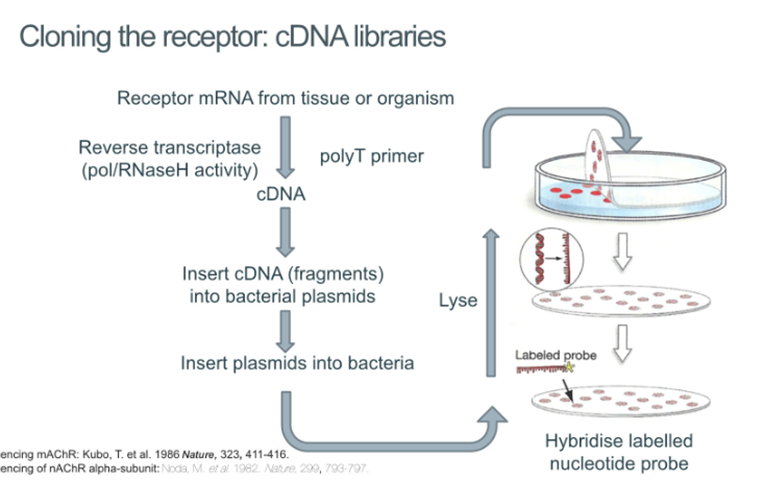
Cloning the receptor
Take the mRNA from the tissue or organism, convert this into cDNA and insert the cDNA into bacterial plasmids
Insert plasmids into bacteria and plate the bacteria, lyse them and hybridise the DNA using a labelled probe which allows you to work out which bacteria contain the cDNA of the receptors that we’re interested in
Sometimes the numbering of Aas can vary considerably in the older literature because they haven’t worked it out from the gene sequence but from the protein sequences
What can we learn from the sequence
We realise from the sequence that all of the different subunits are relatively homologous and structurally the subunits are likely to be similar
Can also predict what post translational modifications like glycosylation and phosphorylation from the sequence —> this allows us to predict where the protein might be situated within the cell (glycosylated will be outside the cell)
Can also do a hydropathy plot analysis which allows us to identify hydrophobic regions → hydrophobic regions 21-24 residues in length indicate a transmembrane pass which is what we can see from the hydropathy plot and can see 4 TM passes
Topology and orientation of nAchR subunits
So far had predicted: an N terminus region rich in beta strands, 4 alpha helical TM domains and a glycosylation site with an EC disulphide bond
However, this doesn’t help you understand the pharmacology of the receptor
Now start to think about expressing this protein in model organisms such as E. coli, xenopus, insect cell culture or tissue culture → opens up more types of analysis
Homologous expression in Xenopus oocytes
Inject the 4mRNAs that you have into the eggs
Allows you to:
follow this with electrophysiology
Observe how the subunits come together
Identify binding sites for drugs that we’ve identified
Mutate the mRNA inserted to observe the functional relevance of certain parts of the protein
Electrophysiological studies of nAchR in xenopus
2 electrodes into the egg and either plant the current or the voltage and look at the response (change in voltage or current) to certain pharmacological stimuli
Can then remove parts of the subunits, mutate parts of the mRNA and observe the response to see how these perturbations affect the pharmacology
Eukaryotic membrane protein expression in E.coli
However, expression of these proteins in bacteria can be problematic because they don’t have the same machinery that humans have
Start to encounter problems with:
Different codon usage
Limited membranes (no ER)
Different membrane protein insertion machinery compared to eukaryotes
No post translational modifaction
Lipid composition differs
Over expression of heterologous protein may lead to inclusion bodies
Multi-subunit proteins difficult to assemble
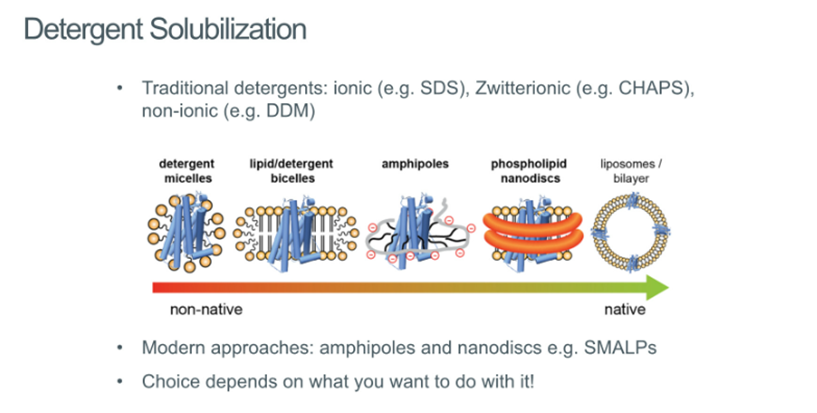
Detergent solublisation
Need to be able to take them out the membrane and purify them
Was initially problematic, but now we’ve been able to come up with solutions around these problems:
Detergent micelles
Lipid/detergent bicelled
Amphipoles
Phospholipid nanodiscs
Liposomes/bilayer
All can be used to extract the receptor out of the membrane so that it can be studies
Separation of membrane proteins
Once isolated, needs to be purified
Involves chromatography and binding a ligand to a chromatography column and passing the sample over this
Affinity chromatography: may involve his tags or maltose binding protein
Ligand affinity chromatography: may involve using an agonist or antagonist like alpha bungarotoxin
Combination: uses a tag-based system for crude purification and functional selection with an agonist-based column
Modern approaches may utilise the his tags attached to maltose binding protein, ligand affinity chromatography older technique and requires you to know what can bind to the receptor
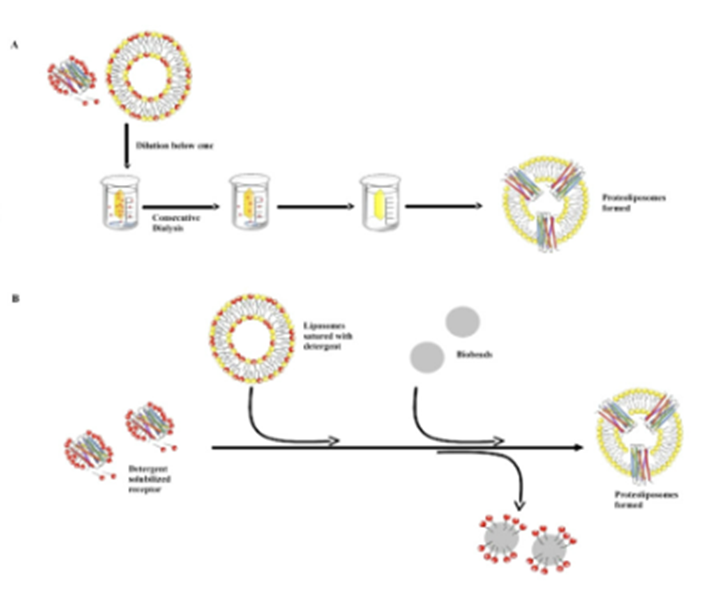
Reconstruction into lipid vesicles
If you want to study the function of the protein you need to return them to the bilayer (reconstitution)
This allows us to understand how fast things are transported across the membrane to understand the accessibility and the binding sites
Challenges involve removing the detergent molceules
Do this by mixing protein with lipids, then can do
Dialysis: if you have low conc of detergent its now a monomer (not in a micelle) and monomers leave the dialysis bag → protein is taken up into the lipid vesicle
Biobeads: polystyrene balls with ahigh affinity for the detergent molecules and pull them out of solution
Dilution
Chemical labelling for the identification of binding sites
Can look at what pharmacophores involved:
ammonia group often onvolved in binding
Has acetylcholine group which contains oxygen. Oxygen great at binding so suggests that this part may be involved in binding
Developed a class of different compounds like
Ach mustard which has a ring under tension → can bind this with the receptor to identify which part of the molecule is interacting with the receptor
BromoAch which has a bromine group which can be reacted with cysteine to identify where the cysteines are in the protein
MBTA: has an ammonium group which localises the reactive functional group to the binding site
DDF has a triple bonded nitrogen which are highly reactive and will react with the binding site
Now that we can localise the binding site, we can label them with 14C or tritium and can now track which subunits these molecules are binding to by running them on a gel and tracking the radioactivity
Found out that all of these different compounds are preferentially reacting within different groups within the alpha subunit
Assuming that all of these different molecules are binding to the alpha subunit, suggests that the alpha subunit forms the primary binding site
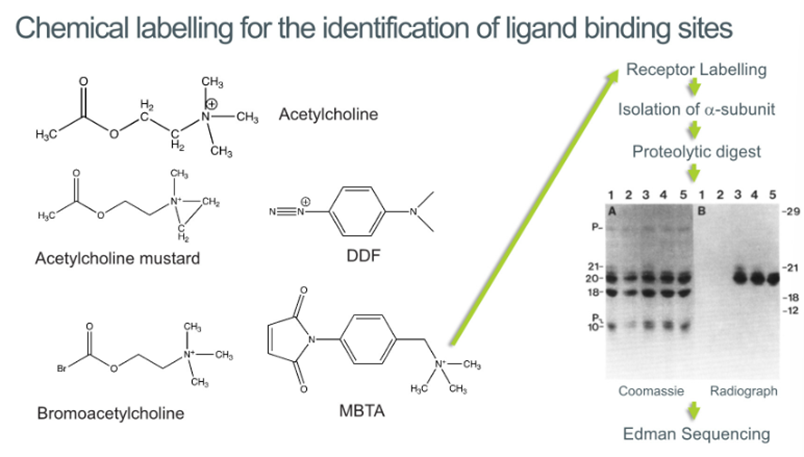
Protease digestion
Digest this with various subunits on the gel and get these different fractions
Can now localise this radioactivity (due to binding of ligand) to a band of 20kD
We now know where the protease will cleave this protein and can do Edman sequencing
The benefit if this is that if you keep on cleaving off individual Aas at some point you will reach your amino acid which you initially radiolabelled and will indicate that you’ve reached the binding site
Can obtain sequence specific information about the localisation of the binding site —> allows you to know the specific Aas involved in binding
However, to avoid using radioactivity, we can take the same approach with mass spec → fragment the protein and work out which protein has an elevated Mw
Identification of Ach binding sites
You would expect the quaternary ammonium group to bind to a negative charge and things that can form hydrogen bonds
But in fact it bonds to tryptophan and tyrosine (aromatic residues) → this is useful because if you want to design a drug, you would know to include these aromatic AAs
Did confirm that the alpha subunits is the primary binding site, however, also showed that there was binding in the gamma and the delta subunits → part of the binding site is facing towards this complementary subunit
We now know that the binding site is between the alpha and the gamma and the alpha and the delta subunit
Structural characterisation of the nAchR
Observed tubular structures if left at -80 for 2 years → these turned out to be an array of AchRs
Gave all the information to reconstitute the structure of the AchR
Used electron diffraction where you defocus the beam of electron microscopy and get a diffraction pattern which can be used to generates an electron density
Then managed to generate a structure of the AchR containing:
Ligand binding domain (extracellular)
Transmembrane domain
Intracellular domain
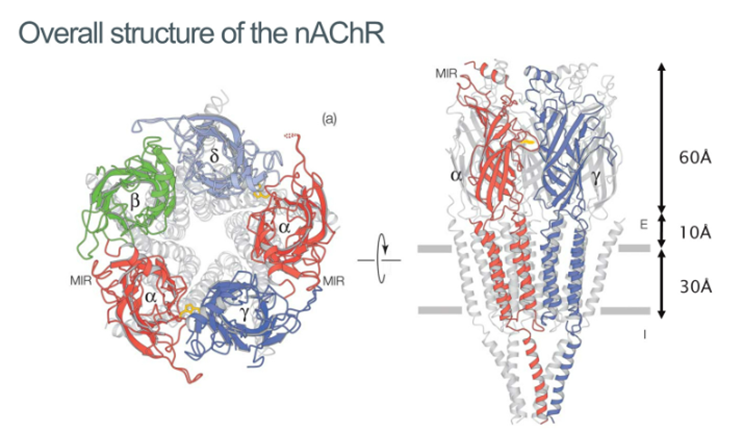
Intracellular domain
This is important because the function of the receptor is modulated by the lipid environment
Intracellular domain is involved in localising with the receptor through interaction with the cytoskeleton
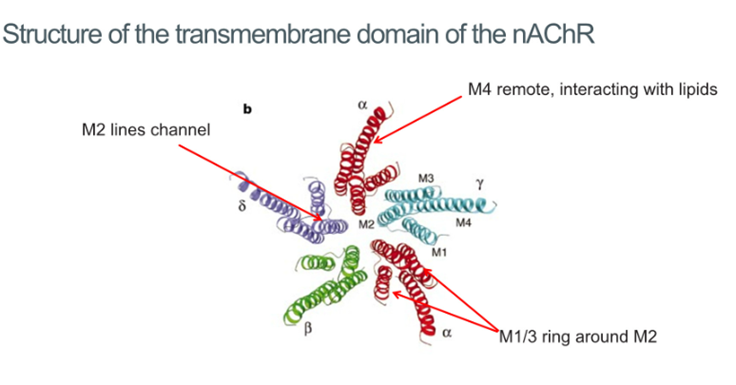
Transmembrane domain
5 subunits all come together and contribute their second TM domain to form this pentameric structure which is a channel going through the membrane
M2 domain conveys selectivity: possesses a negative charge which selects for positive ions
M1 and M3 surrounding this in a ring
M4 TM domain makes the interaction with the lipid bilayer. This receptor is very sensitive to lipid environments and cholesterol allows it to be activated
No activation of the receptor in membranes devoid of steroids because there’s no interaction of the M4 TM domain and so prevents activation
Anaesthetics are hydrophobic and hypothesised that they bind to sites at the lipid protein interface which highlights the functional importance of these domains
Ligand binding domain
The residues involved in binding the Ach referred to as the beta sandwich: 2 beta sheets packing together on top of each other, C loop with Cysteine involved
Within this ligand binding domain the interface region made up of a cys loop and a beta1 and beta2 loop are very important for receptor activation once the ligand has been bound (perturbing this rotation)
Chemical labelling data proved that the binding site was located between the alpha and the delta and the alpha and the gamma subunit as hypothesised
Also Confirmed that there were 4 TM domains per subunit
C loop suggested through chemical labelling that this region interacts with the ligand when bound
Because it’s a channel, a structural change must be involved In fluxing ions → suggested that this is the role of the beta sandwich
Binding site made up of main subunit and complementary residues
C loop fold over the molecule binding
Pharmacophores for mapping Ach binding
Ach is a very flexible molecule that will fit the shape of the receptor and so is not very useful for understanding the binding of this receptor
However, can use other molecules which are a lot less flexible to study binding site
So far we know that Ach has quaternary ammonium group which is +vely charged and the Acetly group which has potential for hydrogen bonding
Strychnine has O atom which is separated by 4.97A from the N atom and matches the distance for binding to the Ach receptors
Pancuronium binds with high affinity to Ach receptor and suggests that there's a hydrogen bond acceptor which is separated by 5.06A from the quaternary ammonium group
A model of the Ach binding pocket
Solid state NMR allows you to determine the structure of the receptor when Ach is bound based off the resonance of particular atoms → Tells you about the local electrostatic environment
We have a positive charge and so it was expected that the binding site would have a negative charge but all they found is a big shell of aromatic groups
NMR studies labelling the molecule (13C) when bound to the receptor caused a shift in peak of the quaternary ammonium group
Position of the group moved by 1.5 ppm and showed that it was binding to this quaternary ammounium group → Ach being bound by this aromatic cage
NMR is very sensitive to aromatic groups and results in these ring current effects
Identified one of the most common binding interactions: extensive cation-pi interactions
Aromatic rings made up of double bonds are highly polarisable due to their fluctuating ring structure and form the basis of cation-pi interactions
As the positive charge approaches, one side of the ring acquires a positive charge and the other acquires a negative charge
Docking studies
Usually rely on a flexible structure binding to a rigid protein but this time held the structure of the ligand rigid and allowed the protein to flex around it
Found the C-loop folding over the ligand
Docking studies made use of an Ach binding protein which is a soluble ligand binding protein which can be isolated from snails
At the time thought that carbonyl group was important for high binding however, was consistently found net to this leucine molecule incapable of H bonding but Ach binding protein mimics the pharmacology of the NachRs in the brain, not the ones that are found in torpedo nobilliana or the NMJ
Lower affinity for Ach in the receptors in the brain due to lack of H bonding between the carbonyl group and the
Snail Ach binding proteins which have a high degree of similarity but instead of leucines, NMJ receptors have threonines which are capable of H bonding which gives higher affinity binding
Ach binding proteins
Once the crystal structure had been defined, tried to exploit this to generate crystal structures of when antagonists and agonists are bound
When the antagonist is present, the C loop seems to be pushed back from the binding site
But in the presence of an agonist, we see the C loop wrap around the agonist
Lots of aromatic groups involved in binding which promotes the idea of cation-pi interactions and confirms the correct binding sites
Gives insight into the type of interactions that these receptors make → useful for trying to optimise interactions between these scaffolds and receptor binding site when developing drugs
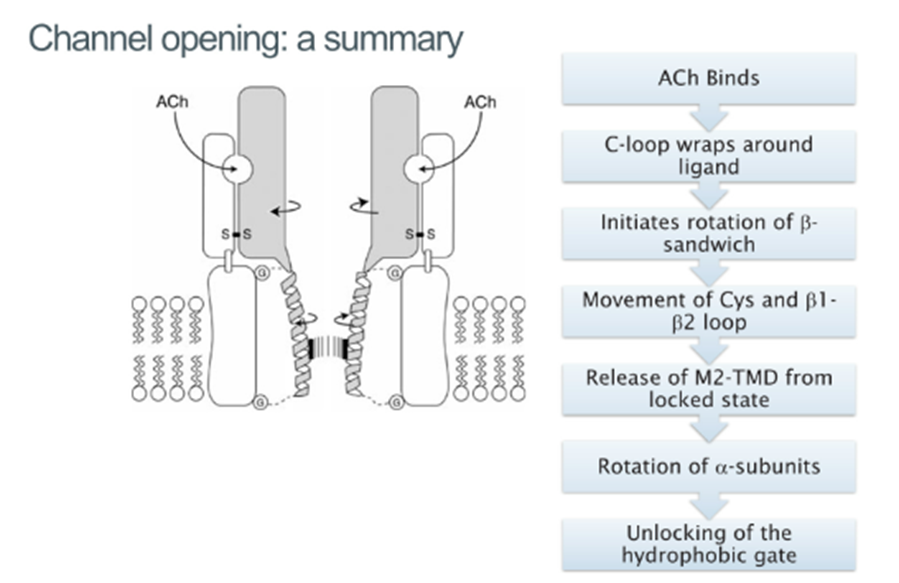
How does movement of the C-loop propagate to the change in structure of the ion channel?
Activation involves ligand binding sequentially of 2 Ach molecules (microsecond timescale), the channel remains open for a few milliseconds and rapidly, the receptor becomes desensitized and the channel closes and so difficult to study the open structure as its very transient
Receptor dropped from a height and atomised Ach sprayed as its falling to activate it, before falling into liquid ethane which freezes it and keeps it in this open conformation
Movement of the C-loop in the ligand-binding domain acts like a lever and pulls the rigid beta sandwich across
The Cys loops and the beta1 and 2 loops form the interface between ligand binding and TM domain —> are pulled away from the top of the TM domains
The AchR acts in a prestressed conformational state, which is being held under tension in the closed state
The moment the loop regions are pulled as a result of agonist binding, then you can release the energy of the Tm doamins and can open up the channel of the bilayer
These loops interact with the M2 TM domains which are involved in lining and opening the centre of the channel
In the closed conformation all the Lecuines are facing the centre, but when the channel opens, they get replaced with these serine molecules which allow the flux of ions
Summary
Prokaryotic ligand-gated ion channels
Ligand gated ion channels in bacteria which are more amenable to crystallography
Don’t bind Ach
ELIC and GLIC:
ELIC has a closed state conformation, while GLIC has an open state conformation
Allows us to get a crystal structure associated with the open/closed LGIC
Proposed a revision to the original model and suggested the opening and closing was driven by the beta1 and 2 loop as opposed to the Cys loop
Causes a rotation of beta sandwich and results in the release f the M2 TM domain and interaction f the channel
Instead of the rotation of the M2 TM domain, there was a pulling apart of the M2 domains of 12A to allow the flux of ions