D3.2 Inheritance HL
1/18
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
19 Terms
segregation
the pair of alleles of each parent separate and only one allele passes from each parent on to an offspring, determined by random orientation of chromosomes in metaphase I
occurs independently in unlinked genes found on different chromosomes
dihybrid cross
investigates inheritance of two traits controlled by two genes, whether inherited together or independently
application of law of segregation and independent assortment
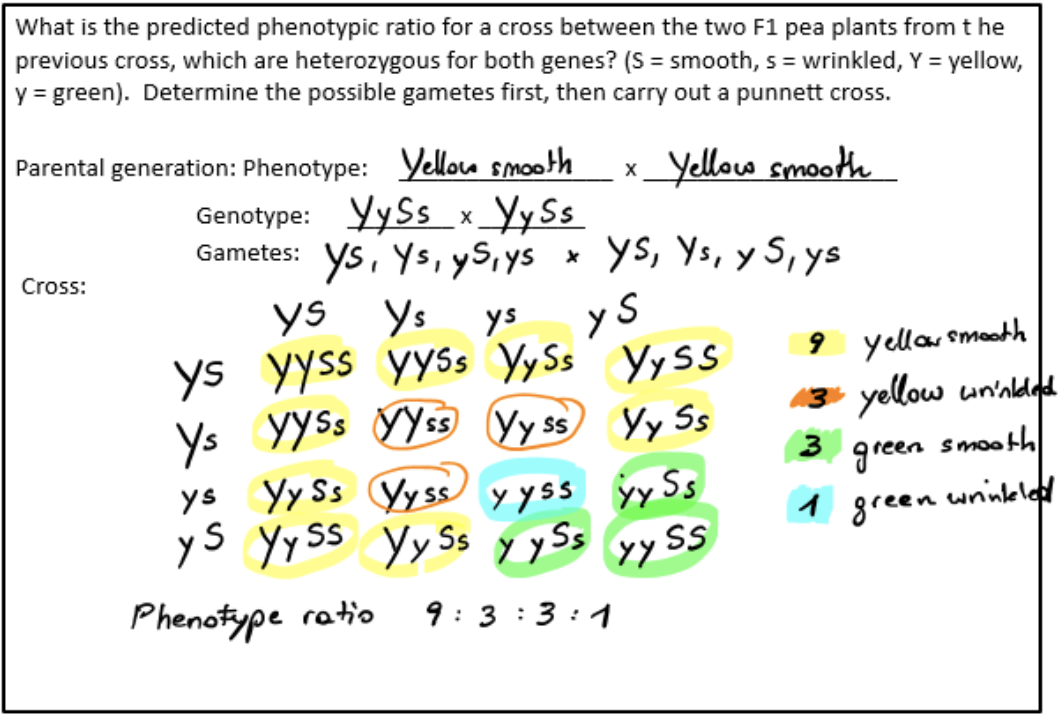
Mendel’s law of independent assortment
assortment of one pair of genes into gametes is independent of the assortment of another pair of unlinked genes, determined through performing dihybrid crosses
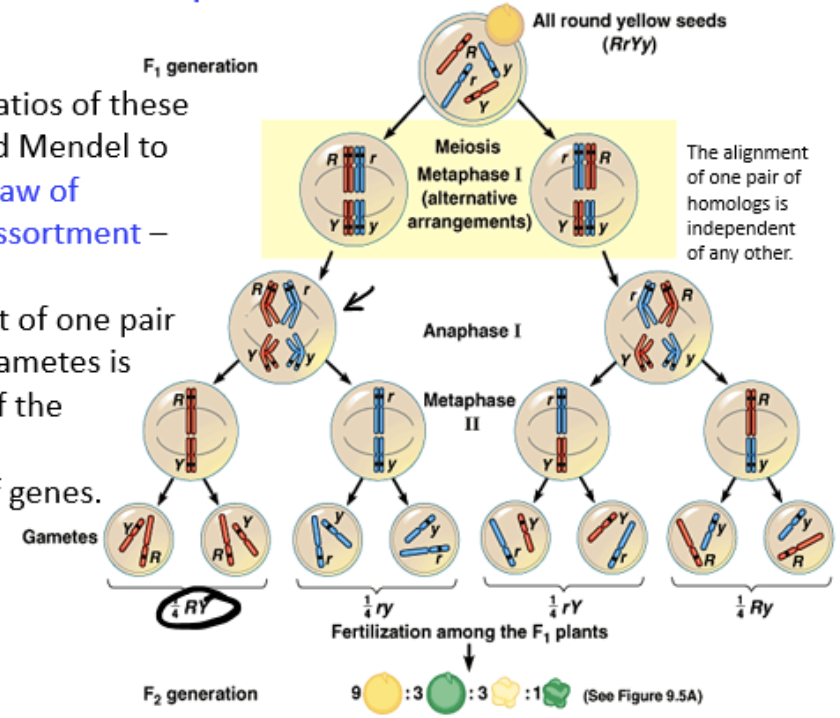
9:3:3:1 ratio
received when parents in a dihybrid cross that are heterozygous for two genes are crossed together
reasons for irregular ratios after dihybrid cross
genomic imprinting
codominance
sex-linked genes
linked genes (carried close together on the same chromosome)
non-heterozygous parents
gene interactions (one gene affects expression of another)
locus
a gene’s specific position on one of 22 types of autosome or one of two types of sex chromosome
linkage group
all genes with loci on the same chromosome
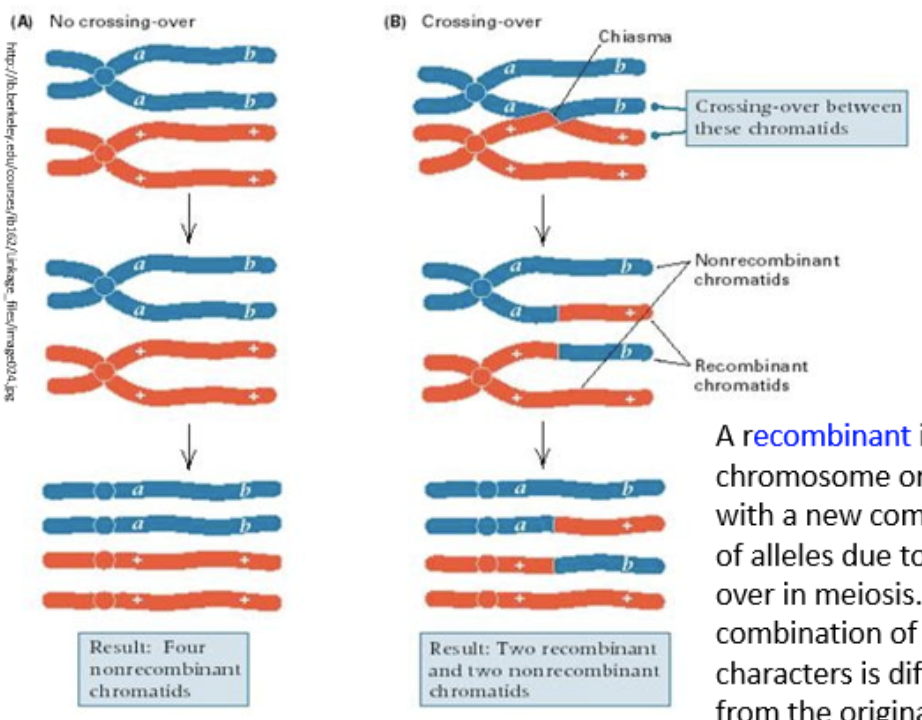
crossing over
two homologous chromosomes, one bearing alleles A and B and the other alleles a and b are paired at prophase I
two nonsister chromatids undergo crossing over, causing portions of each to exchange places
results in two recombinant chromatids, with alleles A and b on one chromatid and a and B on the other, leads to added genetic variation
closer proximity of linked genes
lower likelihood of transferring alleles from one chromosome to another and creating recombinant gametes
recombinant
chromosome or DNA with a new combination of alleles due to crossing over in meiosis, different from that of original parents
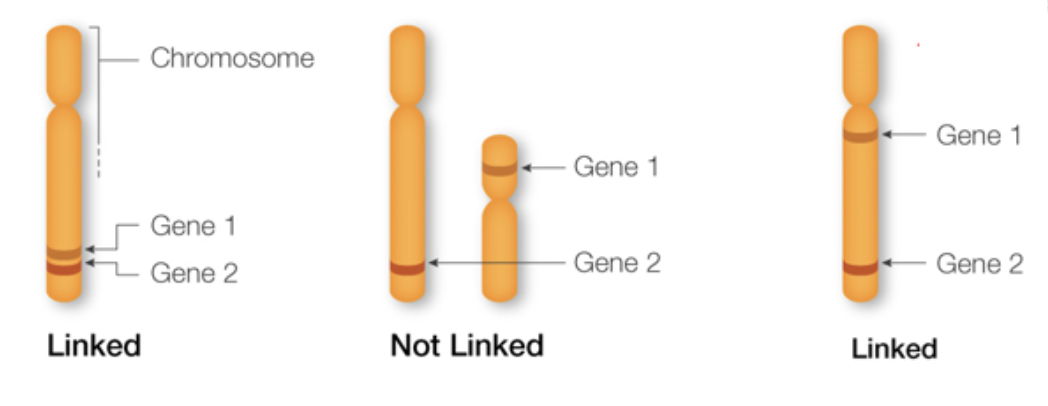
IB gene linkage notation
line represents chromosomes
locus 1
locus 2
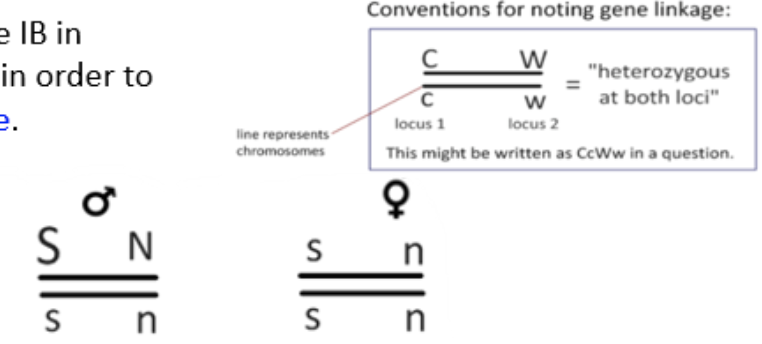
backcross/test cross
tests frequency of recombination between two genes by crossing heterozygous individual with homozygous recessive
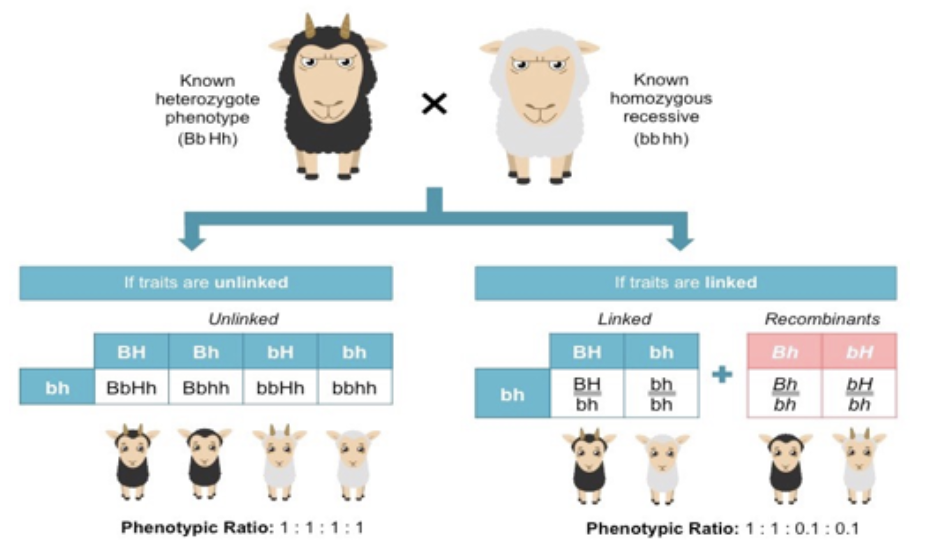
backcross of unlinked genes
equal ratio of four potential phenotypes
backcross of linked genes
two phenotypes in high amounts, two phenotypes in low amounts
chi-squared test
calculate expected frequencies, assuming independent assortment, for each of the four phenotypes (expected frequency = expected probability * total)
determine degrees of freedom
calculate critical value using table and 0.05 confidence
calculate chi squared value
if chi squared value greater than critical value, null hypothesis rejected
chi-squared test null hypothesis
alleles assort independently and are not linked, statistically insignificant
rejected if chi-squared value greater than critical value
chi-squared test alternative hypothesis
alleles do not assort themselves independently, genes are linked
accepted if chi-squared greater than critical value
chi squared degrees of freedom
total number of classes - 1
chi-squared value
∑(observed-expected)²/expected