genetics unit 3 + cumulative
1/103
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
104 Terms
the _______ making up DNA contains the information for making proteins
linear sequence of deoxyribonucleotides
mRNA from DNA
one of the two strands of dna serves as a template for the information to be transcribed into mRNA
mRNAs associate with ribosomes to ultimately produce proteins
the genetic code is written in linear form using the ribonucleotide bases that compose mRNA molecules as ____
“letters”
the sequence of RNA is derived from
the complementary bases in the DNA
in mRNA triplet codons each specify
one amino acids
the code in mRNA contains what kind of signals?
stop and start
certain codons are necessary to initiate and to terminate translation
why is the code triplet for 20 amino acids?
4³ = 64 which is just enough with a few extra
4² would be too little and 4^4 would be too many
the genetic code is
non overlapping
comma-less
degenerate
unambiguous
nearly universal
unambiguous vs degenerate for the genetic code
unambiguous = each triplet specifies only a single amino acid
degenerate = more than one triplet can code for the same amino acid
what infers that the genetic code is non overlapping and comma less
the fact that the code reads three nucleotides at a time in a continuous, linear manner
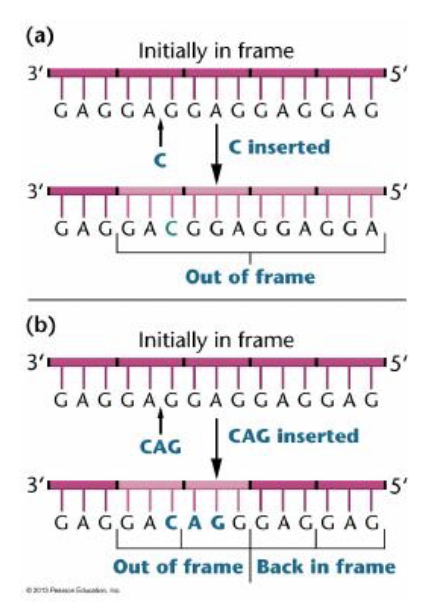
this represents
frameshift mutations
RNA homopolymers are
RNA nucleotides with only one type of nucleotides
who discovered / how did they discover which amino acids were encoded by specific codons
Nirenberg and Matthaei
added homopolymers to an in vitro translation system
poly U directed the incorporation of only phenylalanine - they discovered that UUU codes for phenylalanine
poly a and poly c
poly A= lysine
polt c= proline
the genetic code is degenerate, what does this mean?
many amino acids specified by more than one code
64 different triplet codes but only 20 amino acids
what are the only two amino acids that are encoded by a single codon
tryptophan and methionine
AUG - met
UGG- trp
the wobble hypothesis
proposed by Francis crick
predicts that hydrogen bonding between codon and anticodon at the third position is subject to modified base pairing rules
what three codons serve as termination codons and do not code for any amino acid?
UAG, UAA and UGA
what are some exceptions to the universal genetic code?
mitochondrial DNA
mitochondrial DNA can express UGA as
tryptophan instead of termination
CUA can be altered in mitochondrial dna to be read as
threonine instead of leucine
AUA can be read as what in mitochondrial DNA
methionine instead of isoleucine
AGA can be read as what in mitochondrial dna
termination instead of arginine
AGG can be read as what in mitochondrial dna
termination instead of arginine
rna synthesis overview
no primer is required for initiation
the enzyme uses ribonucleotides instead of deoxyribonucleotides
transcriptions begins with template binding by RNA polymerase at a promoter
the o subunit is responsible for promoter recognition
prokaryotic transcription begins at the transcription start site where
the dna double helix is unwound to make the template strand accessible
e.coli promoters that have two consensus sequences _______ and _____ positioned at -35 bp and -10 bp with respect to the transcription initiation site
TTGACA TATAAT
in Ecoli once initiation has been completed with the synthesis of the first 8-9 nucleotides ….
sigma dissociates and elongation proceeds with the core enzyme
at the end of the gene in e coli, transcription terminates when a termination sequence is reached at ~40 bp and
transcribed into rna, forming a hairpin structure (self complementary)
transcription in eukaryotes
occurs in the nucleus and not coupled to translation
requires chromatin remodeling
promotors, enhancers also influence transcription regulation
what is required of eukaryotic mRNAs to produce mature mRNAs?
processing
what is the shortened version of RNA polymerase II?
RNP II
difference between start codons in prokaryotes and eukaryotes
both use AUG
prokaryotes it represents n-formylmethionine (feet)
eukaryotes, unformulated methionine is used
what are the three termination codes and what do they stand for
UAG, UAA. UGA
they do not code for any amino acids
What are the key components of the RNA Polymerase holoenzyme in prokaryotes?
core enzyme: responsible for elongation
sigma subunit: responsible for promoter recogninition and template binding
no primer is required for rna synthesis
identify the two consensus sequences in E. coli promoters
-35 region:TTGACA
-10 region: TATAAT (pribnow box)
describe the mechanism of prokaryotic termination
it occurs when a termination sequence (approximately 40 bp) is reached
the transcribed RNA forms a hairpin structure due to self-complementarity which causes the polymerase to dissociate
how does the location of transcription differ in prokaryotes from eukaryotes?
in prokaryotes: translation and transcription are coupled in the cytoplasm
how does eukaryotic transcription differ from prokaryotic?
eukaryotic occurs in the nucleus
requires chromatin remodeling to access dna
involves regulation with both promoters and enhancers
produces an initial transcript hnRNA that requires extensive processing
what does the wobble hypothesis suggest?
the first two positions of the codon require strict base pairing but the third position of the anticodon interaction is subject to modified base-pairing rules, allowing one tRNA to recognize multiple codons
what is the initial transcript for eukaryotes that requires extensive processing
hnRNA (heterogeneous nuclear RNA)
describe the components of a eukaryotic core promoter?
TATA Box: located at -35 bp (TATAAAA)
facilitates DNA denaturation to determine the start side
CAAT Box: located -80 bp
influences the efficiency/ rate of transcription
What are enhancers and where are they located?
they are cis-acting elements that modulate transcription from a distance
found upstream, within introns or downstream of gene
distinguish between general and specific transcription factors
general factors: absolutely required for RNP II transcription (TFIIA, TFIIB, TFIID)
Specific factors: influence the rate and efficiency of transcription for specific genes or groups of genes
what transcription factor specifically binds to the TATA sequence?
TFIID
CAAT box
promoter element
-80 upstream
helps efficiency
Describe trans acting factors that facilitate RNP II binding
usually a protein coded for by another gene
gen transcription factors are required for transcription by RNP II
specific TFs influence rate and efficiency of transcription and may be specific to gene or group of genes
in humans what are the transcription factors?
TFIIA, TFIIB, TFIID
which transcription factor binds directly to the TATA sequence?
TFIID
cis- promoter elements that interact with RNP II - how does it begin transcription?
RNP II recognizes a core primer element in the DNA sequence that will determine where RNP II binds and begins transcription
TATA box in eukaryotes
core promoter element located at -35 bp upstream from the start of transcription
consensus sequence TATAAAA
helps facilitate denaturalization of helix so helps determine transcription site
in transcription in eukaryotes the first rna produced is
hnRNA (heterogeneous nuclear RNA)
also called primary transcriptt
what does RNA polymerase ii do?
makes mRNA
Core promoter=
the exact spot where transcription begins
introns in rna are removed by
splicing
in rna processing what does the 7-methylguanosine cap do?
protects the 5’ end of mRNA
transports mRNA out of the nucleus
what does the poly-a tail do in rna processing
has up to 250 residues
protects RNA from degradation
introns and rna
removed from rna
they are not expressed in the amino acid sequence of the protein
give overview of RNA processing
dna template → (transcription)→ pre mRNA→ (cap and tail added)→ processed pre-mRNA → spliced mRNA
exons and introns in mature mRNA
introns removed, exons joined together
therefor, mRNA is usually smaller than initial rna
translation
the biological polymerization of amino acids into polypeptide chains
translation requires
amino acids
mRNA
ribosomes
tRNA
ribosomes
large, intricate and numerous cellular structures
each has a large sub unit and a small subunit
eukaryotic vs bacterial cells and ribosomes
one bacterial contains about 10,000
eukaryotes have many times more
what are ribosomes composed of
ribosomal proteins and rRNAs
____ provide for important catalytic functions associated with translation
rRNAs
what do ribosomal proteins do?
promote binding of various molecules involved in translation?
rRNA genes are also called
ribosomal DNA
rRNA genes are ______ repeated
moderately repetitive and tandemly
ecoli genome and rRNA
has 7 copies of a single sequence that encodes all 3 rRNA components
what are the three rRNA components for prokaryotes?
16S, 23S, 5S
Eukaryotes and rDNA
MANY more copies than prokaryotes
present in clusters at various chromosomal sites
a single transcript of rDNA encodes for
three of the components
28S, 18S, 5.8S
In eukaryotes there is a unique component of rRNA that is encoded by a distinct gene cluster and located separately - not part of the larger transcript. what is it?
5S rRNA
trna function can be described as
adapter molecules to adapt the triplet codons in mRNA to the correct amino acid
tRNAs overview
composed of only 75-90 nucleotides
identical structure in bacteria and eukaryotes
transcribed as larger precursors but cleaved into mature 4S molcules
several nucleotides are modified through post transcriptional modification
what activates tRNAs with the appropriate amino acid?
Aminoacyl tRNA synthetase
how many different enzymes are each responsible for linking a different amino acid to its corresponding tRNA molecule?
20!
what are the three steps of translation?
initiation
elongation
termination
what does initiation require?
small and large ribosomal subunits
GTP
charged initiator tRNA
mg2+
initiation factors
initiation of translation steps
1. mrna and IF1 bind to small subunit
2. IF2 and IF3 bind to complex while initiated tRNA binds to mRNA codon in the P site
3. large subunit binds to complex, IF factors are released EF-Tu binds to tRNA, facilitating entry into A site
what does elongation require in translation?
both ribosomal subunits assembled with the mRNA to form the peptide site and aminoacyl cite
elongation in translation steps
trna enters A site
peptide bond forms
ribosomal shifts (translocation)
what happens when tRNA enters the a site in elongation
a charged tRNA with an amino acid matches its anticodon to the mRNA codon
what happens after the trna enters the A site in elongation of translation
the amino acid in the Psite is transferred to the amino acid in the A site, growing the chain
describe translocation in elongation
ribosome moves 1 codon forward via gTP
trna moves A site to P site and p site to e site
describe all the sites of elongation in trna elongation
a site= arrives p site= peptide bond e site= exit
termination is signaled by what in the A site for translation
stop codons
what are the stop codons
UAG, UAA, UGA
what helps cleave the polypeptide chain from the trna to release it from the translation complex?
GTP dependent release factors
eukaryotes and translation
ribosomes are larger than in bacteria
transcription and translation are spatially and temporally separated
requires more factors for initiation, elongation, termination than bacteria
ribosomes are associated with the endoplasmic reticulum
in prokaryotes ribosomes are
free floating
proteins and diseases through translation
accumulation of intermediates due to metabolic deficiencies (Archibald Edward Garrod)
inborn errors of metabolism
incest related
beadle and Tatum
showed rear nutritional mutations in the bread mold neurospora caused the loss of enzymatic activity that catalyzed an essential reaction in wild type organisms
neurospora
a fungus
in the wild type like bacteria can be grown on minimal media
can be used to study genetic mutations of nutritional mutants
mutations can be generated at high frequency using radiation
studies of neurpspora led us to the one gene on enzyme hypothesis, describe it
not all proteins are enzymes and some proteins have more than one subunit
this led to us actually ending up to one-gene:one polypeptide chain
how can you tell if a neospora has a mutation?
does not grow on minimal medium
grows on complete medium
neospora can tell us what genetic mutation has occurred by
adding each nutritional value to a minimal media until it grows
shows which exact mutation occurred
spontaneous mutations
happen naturally and randomly
are usually linked to normal biological or chemical processes in the organism