Regulatory RNAs
1/16
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
17 Terms
Recall: Information Content of an Organism’s Genome…
the information content of an organism’s genome (as measured in genes) is not proportional to its complexity (as measured in cell number)
differences must lie within non-coding regulatory regions
these non-coding RNAs have a role in the regulation of protein expression
protein coding sequences, rRNA and tRNA account for only 1.5% of genome
combinatorial control and alternative splicing accounts for another 24% (introns and non coding DNA regulatory sequences)
as of 2012, the ENCODE project has determined that ~75% of the human genome is transcribed
what accounts for the remaining transcripts?
Previously known types of RNAs
mRNA (protein coding)
rRNA (ribosomal RNA)
tRNA (transfer RNA)
snRNAs: components of splicing factors
snoRNAs (small nucleolar RNA): guide modification of snRNA, tRNA and rRNA
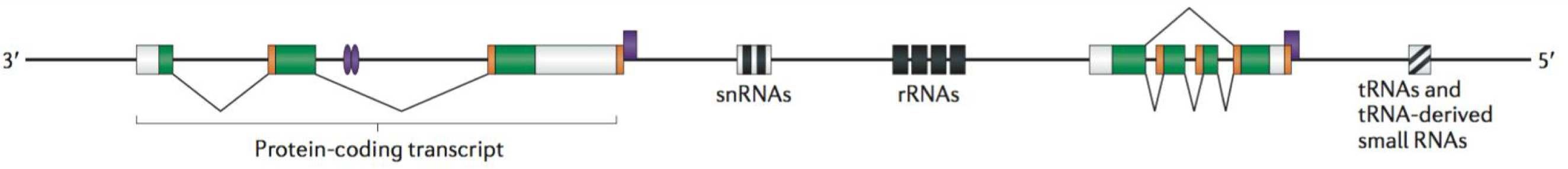
Types of non coding RNAs
microRNAs (miRNA or miR)
small interfering RNA (siRNA)
long noncoding RNA (lncRONA)
piwi-interacting RNA (piRNA)
promoter-associated small RNAs
terminus-associated small RNAs
tRNA-derived small RNA (ts RNA)
tRNA-derived stress-induced RNA (tiRNA)
tRNA-derived fragment
splice site RNA (spliRNA)
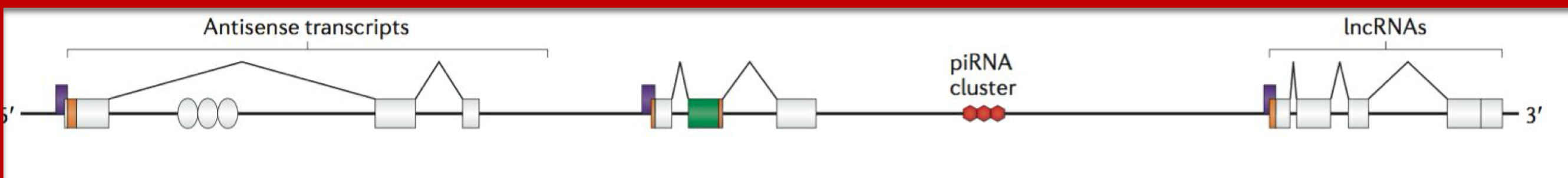
Bias of common RNA sequencing methods
Not all RNA gets a poly(A) tail (e.g. noncoding RNAs, unstable RNAs, and many regulatory RNAs)
Only many mature mRNAs have a poly(A) tail
most standard RNA methods only “catch” poly(A)+ RNA (e.g. oligo(dT) selection)
we end up missing a lot
microRNA (miRNA or miR)
a new control for regulating mRNA translation into a protein
microRNAs were first discovered thru genetic studies but their importance as a general regulatory mechanism was not appreciated
it was only after sequencing genomes and sequencing the transcriptome (i.e. all the RNA transcripts produced in a cell) that they were discovered to be abundant and widespread in many plants and animals
it also became established that they carried out functions in the cell
miRNA and Devlopment in C. elegans
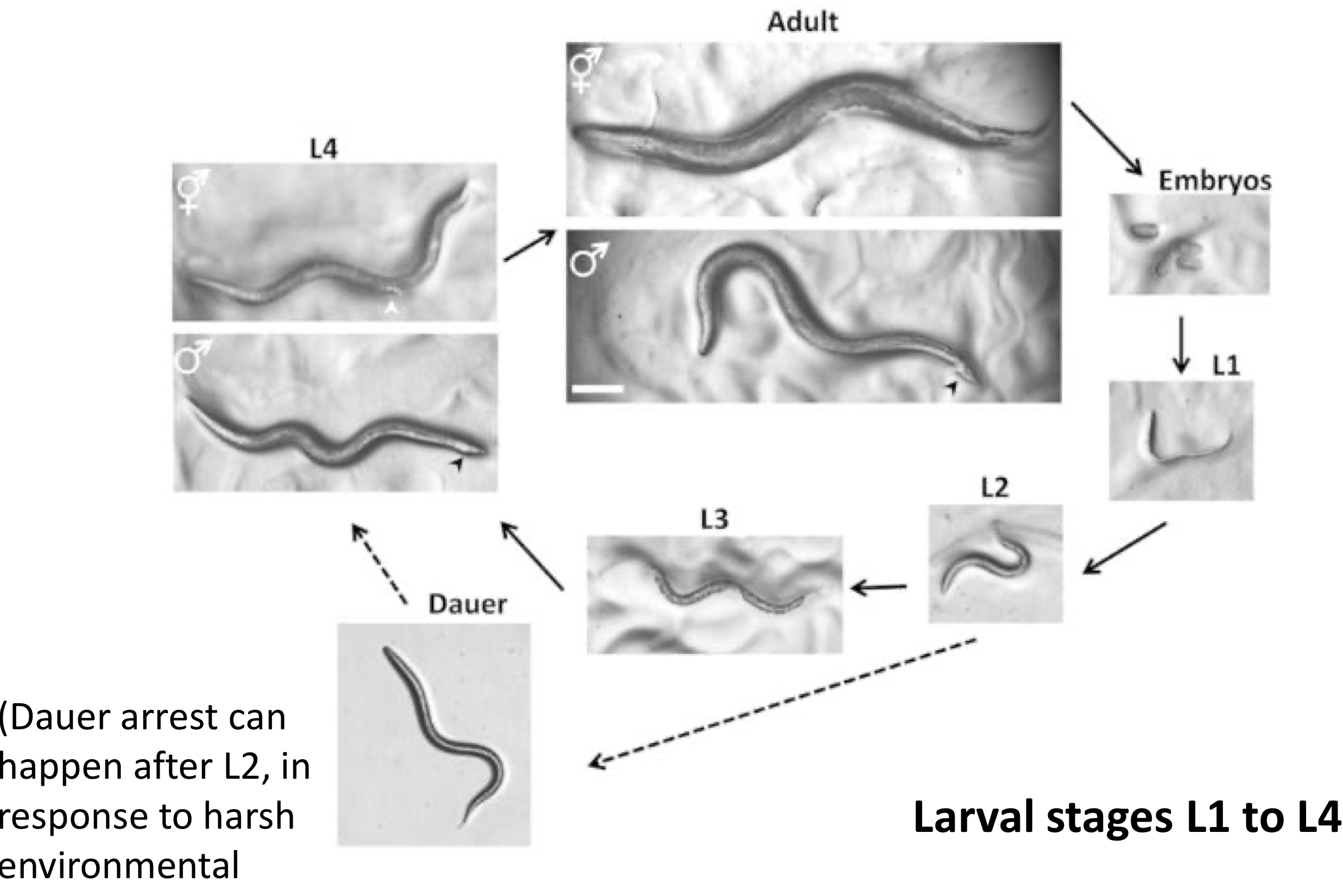
Heterochronic Mutations in C. elegans
heterochony: a developmental change in the timing or rate of events, leading to changes in size and shape
i.e a change in where and when (and how much) genes are expressed, leading to changes in a developmental program
adult lin-4 lof mutants lack many adult structures and they are unable to lay eggs on account of a failure to develop a vulva (eggs accumulate in their bodies)
lin-14 lof mutants develop certain adult features precociously at larval stages, resulting in smaller, poorly formed adults
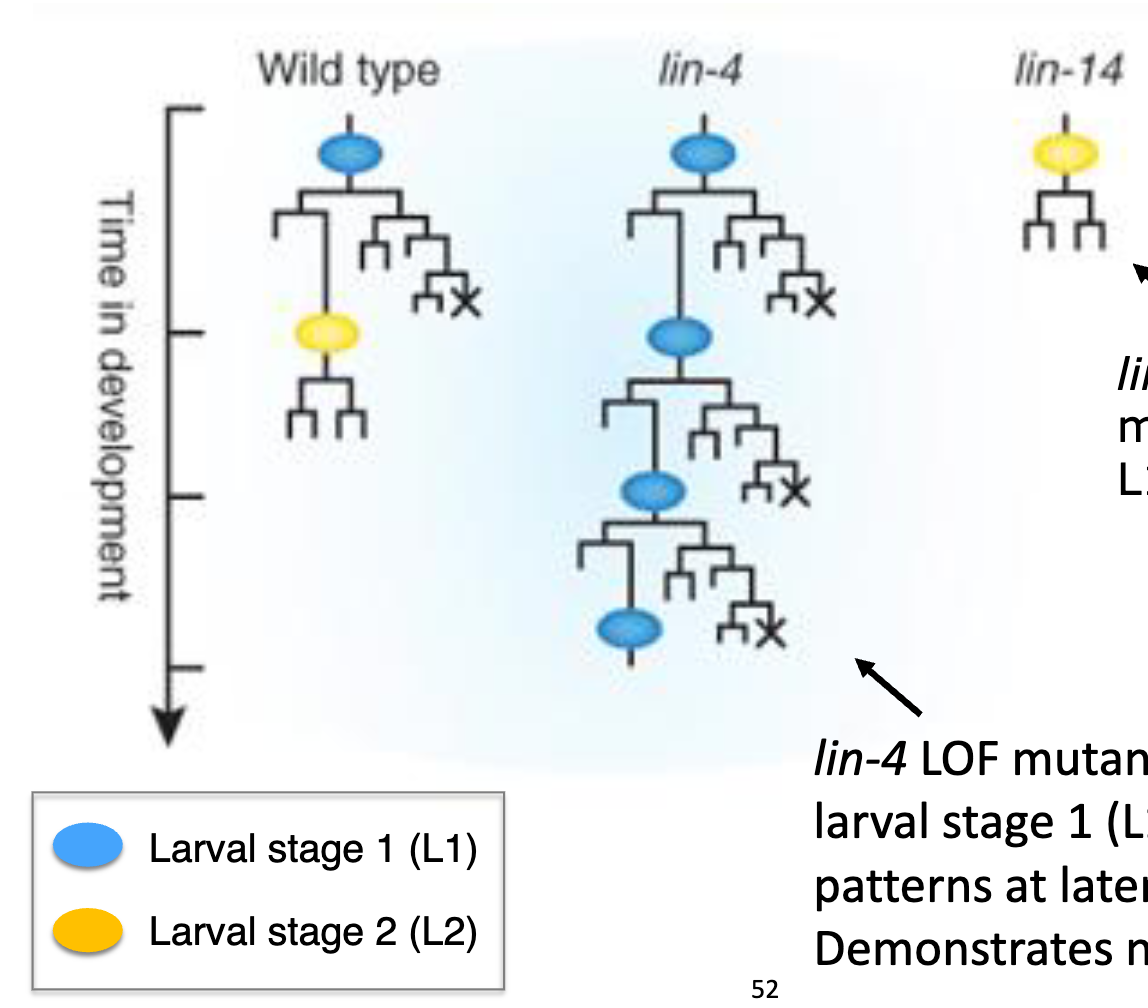
lin4 and lin-14 LOF lineages
lin4 and lin-14 LOF lineages differ from WT animals
lineages of the T cell, a lateral hypodermal cell are shown.
lin-4 LOF mutants repeat larval stage 1 (L1)-specific patterns at later stages → demonstrates neoteny (retention of juvenile physical or behavioral traits into adulthood)
lin-14 LOF mutants skip the L1 pattern
therefore, lin-4 must suppres L1, whereas lin-14 must promote L1
lin-14/ lin-4 gf/lf mutants
lin-14 (gf) mutants resemble lin-4 (lf) mutants
lin-4 (lf)/lin-14 (lf) mutants look like lin-14 (lf) mutants
therefore, lin-14 is epistatic to lin-4
lin-4 —| lin-14 → L1
lin-14 is a novel transcription factor and lin-4 acts to regulate its activity
Identifying and cloning lin-4 and lin-14 (forward genetics techniques)
choose an easily identifiable phenotype (the retention or suppression of the L1 phase)
generate mutants and identify as many of them as possible w/ the desired phenotype (they likely work in the same pathway)
basic genetic analysis to identify LOF and GOF mutants
epistatic analysis to work out a) do these genes work in a common pathway? b) if so, which one works downstream of the other
linkage analysis: crude mapping followed by fine mapping (i.e which molecular markers are closest?) and positional cloning to narrow down the location and sequence of the gene
Previous work on lin4 and lin-14
There are plenty of mutant strains of lin-14, all with mutations in a different area of the gene. However, there were only a couple of different lin-4 mutant strains.
It was very difficult to find the lin-4 gene in linkage mapping and positional cloning experiments
suggests that lin-4 must be small! It’s a hard target to hit w/ random mutations
Lee et al. Data (focus on identifying the lin-4 sequence)
Difficulties in fine mapping
closest molecular marker is 0.1 cM away (ie. recombination rate is about 1 out of every 1000 events) → need to screen many C. elegans
bli-3 is 2 cM away, and dpy-10 is 0.5cM away
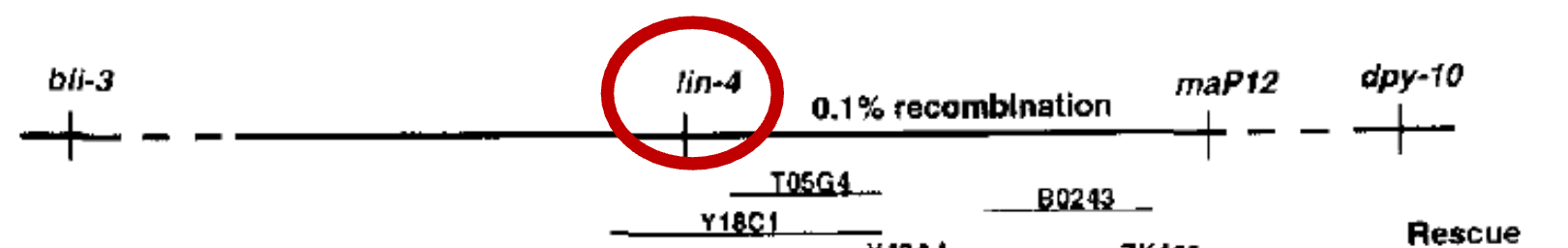
Lee et al. Data (focus on identifying the lin-4 sequence). continued
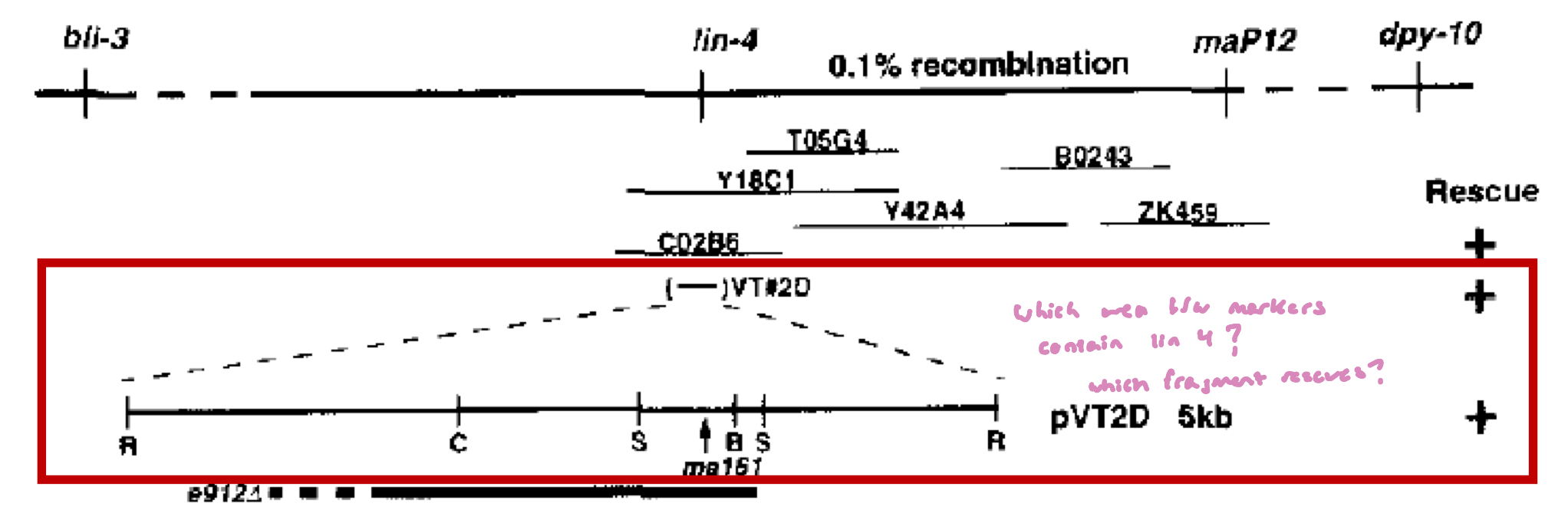
DNA fragments from this region were cloned into plasmids, which was introduced into the lin-4 (lf) mutant (the e192 strain) to see which fragment would restore the WT phenotype
the lin-4 (e192) strain has a deletion that covers most of the VT2D sequences
somewhere in that deleted region lies the functional lin-4 sequence
the VT2d clone rescued the phenotype (+ symbol)
Lee et al. Data (focus on identifying the lin-4 sequence). continued
take the VT2D fragment and cut it into smaller pieces → clone each into plasmids → introduce them into the lin-4 (lf) mutant
check if it can still resuce phenotype
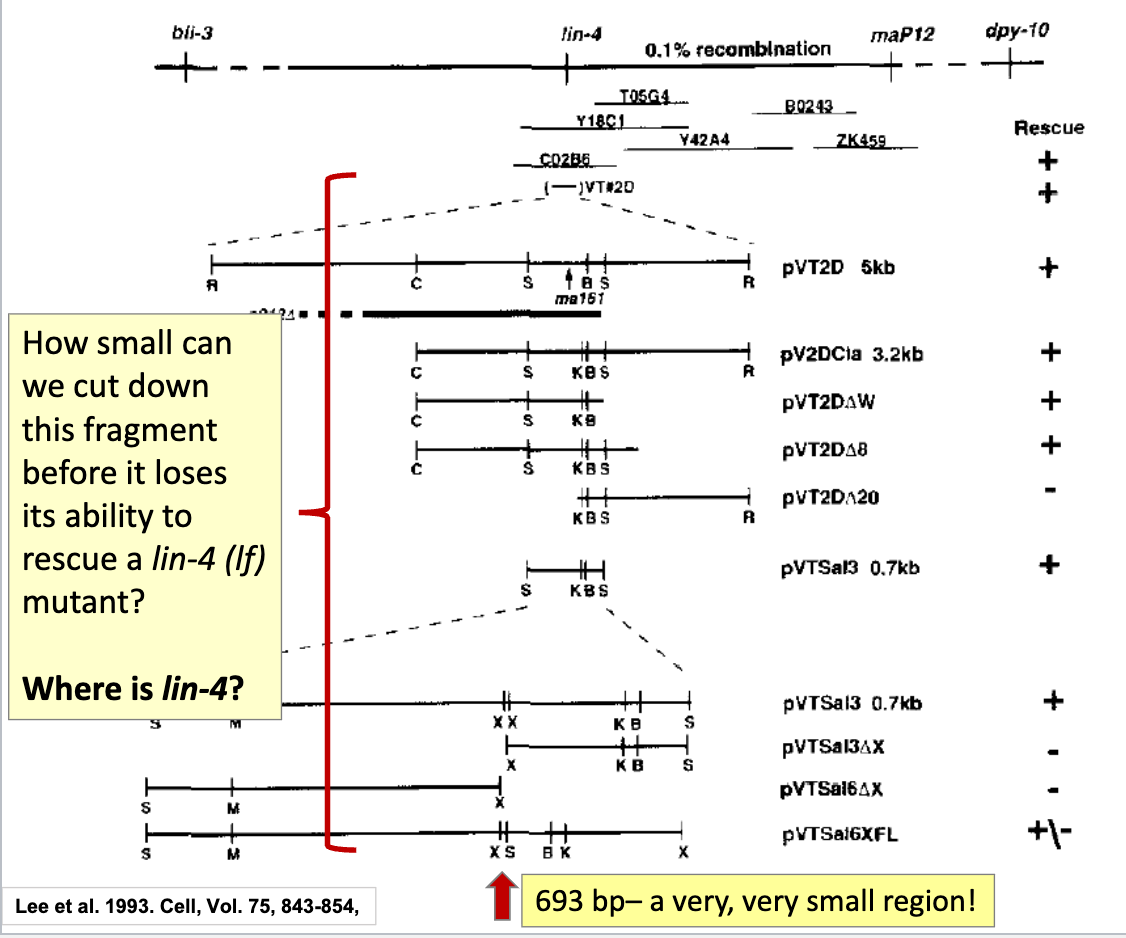
Sequence Analysis of the 693 bp region containing lin-4
lin-4 is within an intron of another unrelated gene (doesn’t contribute to development)
lin-4 is its own functional unit, even though it sits inside another gene
When they looked at RNA made from this region, they found:
A long RNA (~60–70 nt) = precursor (pre-miRNA)
A short RNA (~20–22 nt) = mature functional RNA
short RNA actually does the job
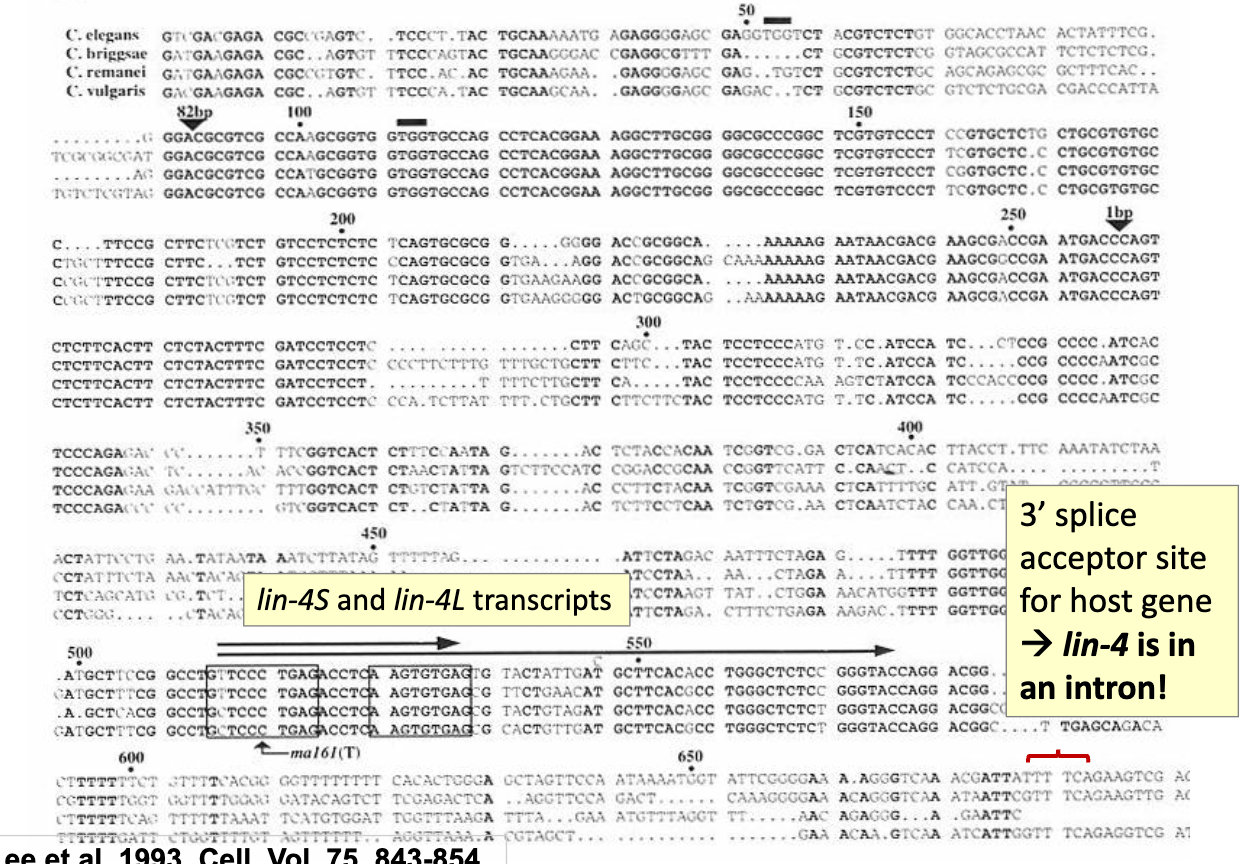
lin-4 is not likely to code for protein
VT2D sequence was used to probe a cDNA library, but none of the cDNA clones identified contained sequences that overlapped with the 692 bp region linked to lin-4 function.
lin-4 appears to lie within the intron of another gene! (Note: the deletion of this 692 bp region in the e912 LOF strain does not appear to impact any other gene function.)
No apparent ORF or start or stop codons conserved between the 4 related nematode species.
One possible ORF of some length in the C. elegans sequence, but mutations of this region did not disrupt the ability of the VTD2 fragment to rescue lin-4 function
Where is lin-4s transcript detected?
Only lin-4S transcript found at detectable levels in WT cells (*through northern blots; PCR is more sensitive)
lin-4L transcript only detected in lin-4 (lf) mutants expressing a construct containing the cloned 692 bp sequence→ likely expressing this to much higher levels than WT.
Figure:
northern blot of total RNA isolated from WT, lin-4 (lf) and rescued C. elegans
pprobe: antisense transcripts from the vector containing the cloned sequences. These would be able to anneal to the RNA transcripts produced in the cell