Chapter 10 - DNA structure and analysis
1/77
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
78 Terms
Genetic material
- information contained in genes that gets passed onto new generation
- source of variability among organisms
to serve as genetic material, molecule must be able to
- replicate
- store information n
- express information
- allow variation by mutation
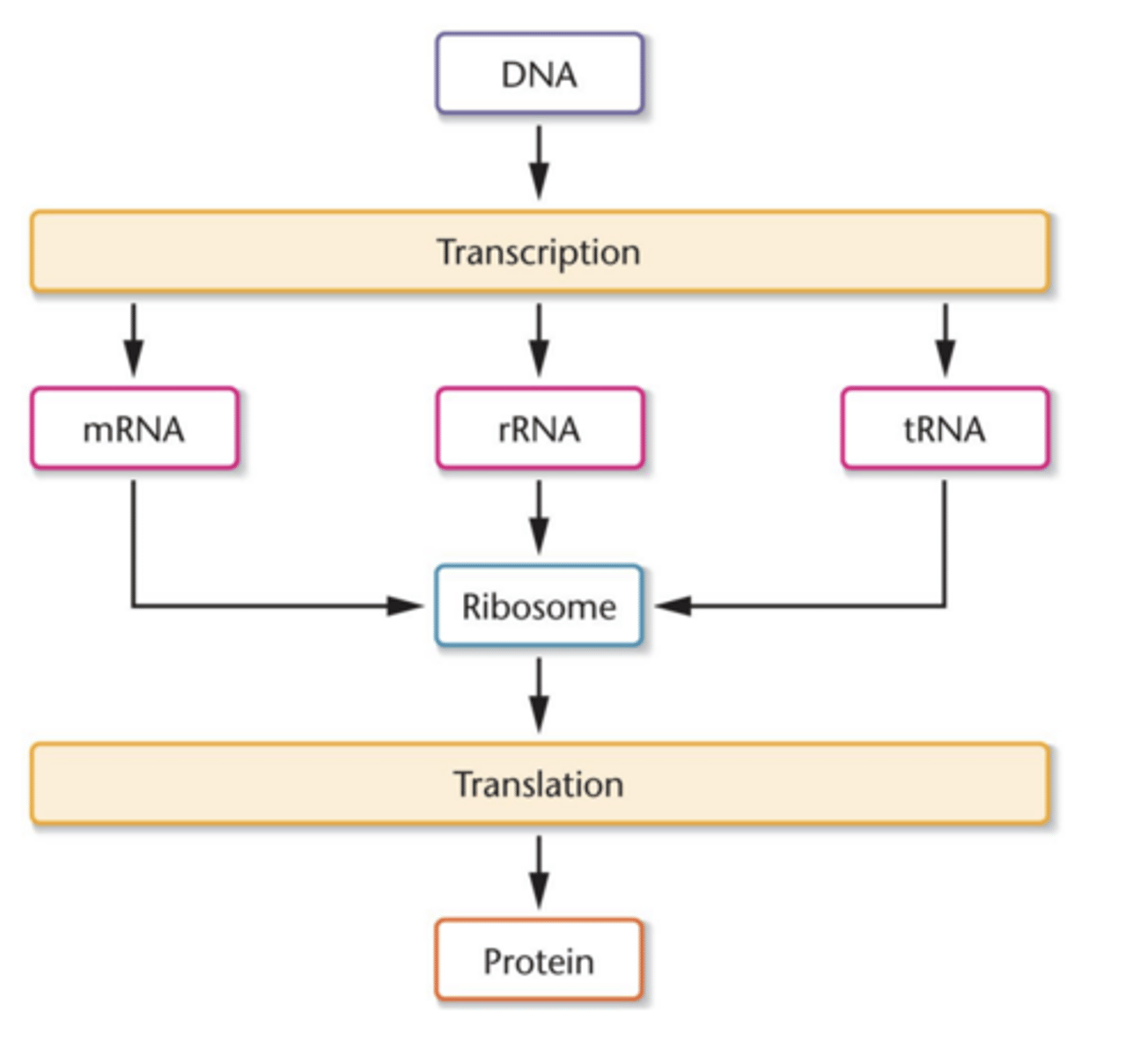
Central dogma
DNA--> RNA--> protein
- DNA makes RNA (transcription), which makes proteins (translation)
Tetranucleotide hypothesis
- DNA contains equal amounts of four nucleotides
- postulated identical groups and repeats of four components was basis for DNA structure
- lack of chemical diversity in DNA suggested it could not store extensive genetic information
- proteins favored as genetic material
Avery, MacLeod, and McCarty
- 1944 publication on chemical nature of transforming principle in bacteria
- first direct experimental proof that DNA is biomolecule responsible for heredity
Griffith
- provided foundation for Avery, MacLeod, and McCarty research
- showed avirulent strains of diplococcus pneumoniae could be transformed to virulence
- speculated transforming principle could be part of polysaccharide capsule or compound required for capsule synthesis
Transforming Principle is DNA
-1944 published results of their experiment
- DNase (deoxyribonuclease) utilized to destroy transforming activity
- demonstrated transforming principle was DNA, not protein
Hershey and Chase
- used E. coli and bacteriophage T2
- showed that DNA, not protein, is the genetic material
- used radioisotopes ^32P and ^35S
- demonstrated DNA enters bacterial cell during infection and directs viral reproduction
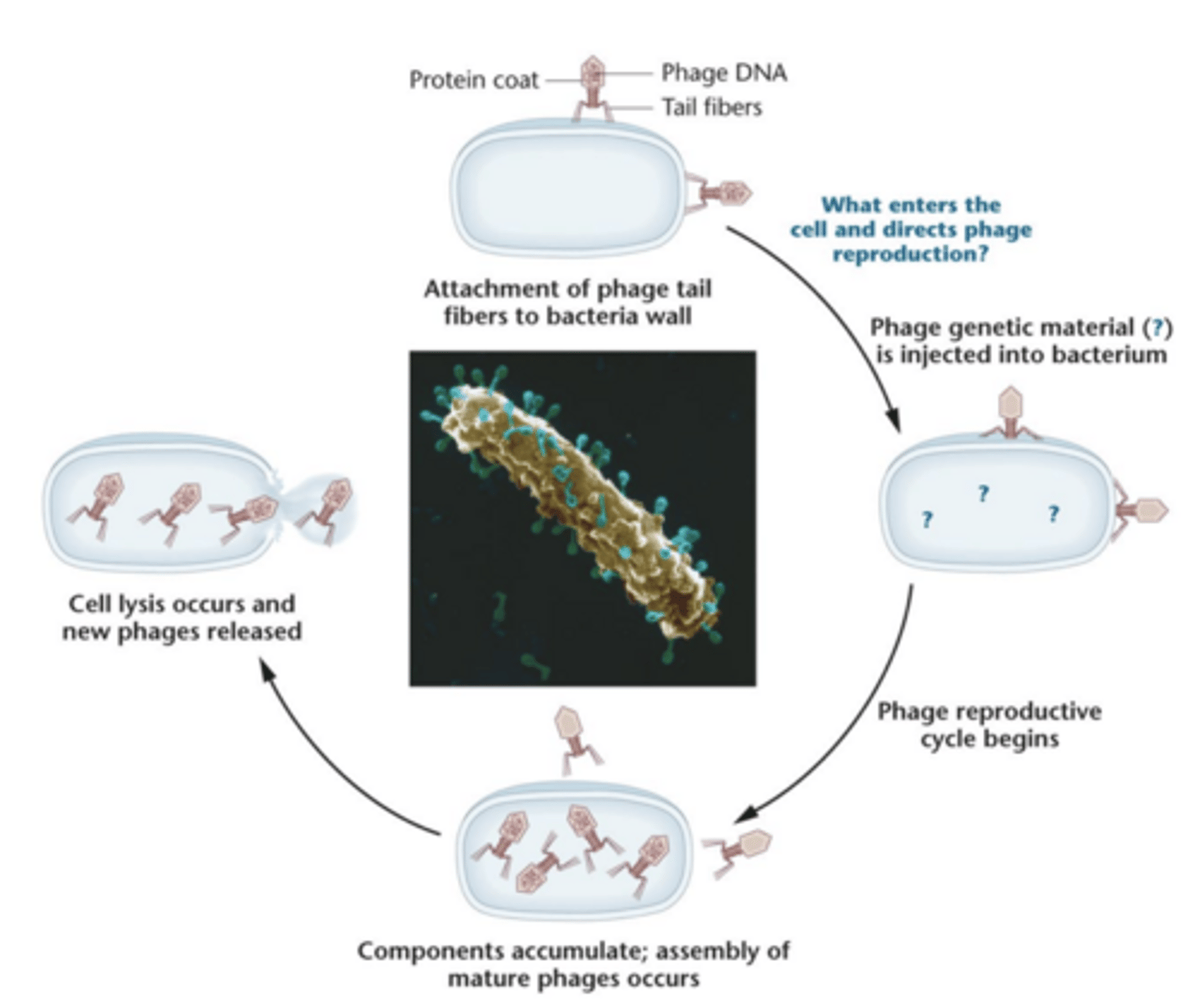
Hershey and chase experiment pt 2
Protoplasts
- E. coli is treated with lysozyme
- outer wall is removed without destroying bacterium - naked cell
transfection
- infection by only viral nucleic acid
- proves conclusively that viral DNA alone contain all necessary information for production of mature viruses
Indirect evidence: Distribution of DNA
- close correlation between gametes and diploids in amount of DNA and number of chromosome sets
- no such correlation between gametes and diploids for proteins
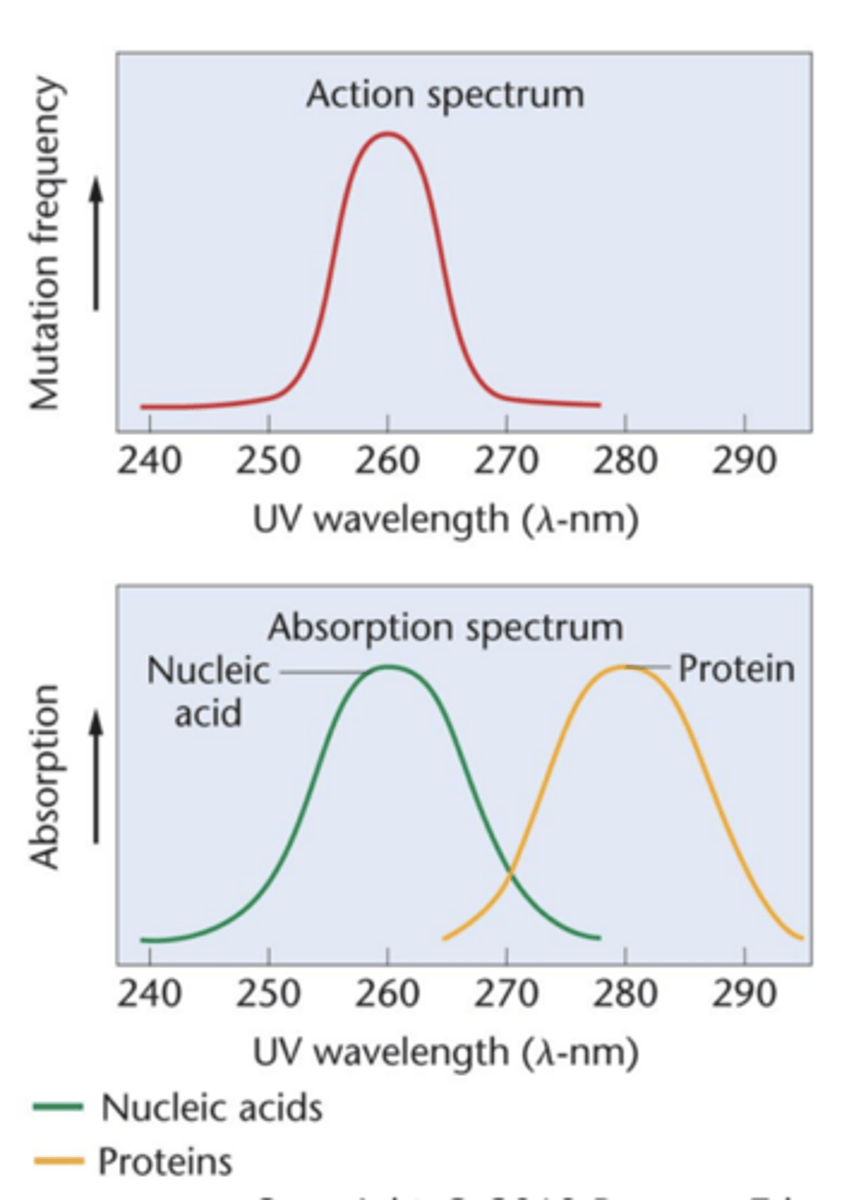
Indirect evidence: Mutagenesis
- UV light: most mutagenic at wavelength 260 nm (action spectrum)
- DNA absorbs UV at 260 nm
- Protein absorbs UV at 280 nm
--> a wavelength at which no signifiant mutagenic effects are observed
- molecule serving as genetic material expected to absorb at mutagenic wavelength
Recombinant DNA technology
- segments of eukaryotic DNA corresponding to specific genes isolated and spliced into the bacterial DNA
- complex inserted into bacterial cell and monitored
Eukaryotic DNA now functional in bacterial cell
- eukaryotic gene product in bacteria containing eukaryotic gene provides direct evidence: DNA is present and functional in bacterial cell
- Example: insulin and interferon production by bacteria
Transgenic animal example
- Human DNA microinjected into fertilized mouse egg
- DNA encoded human beta-globin gene
- Now present and expressed in mouse and transmitted to progeny
RNA as Genetic Material in Some Viruses
- some viruses have RNA core, not DNA
- TMV: Tobacco mosaic virus (1956)
--> Demonstrated RNA serves as genetic material for these viruses
RNA replicase
- Enzyme isolated from E. coli (1965, Pace and Spiegelman)
- Replication of the viral RNA is dependent on RNA replicase
Retroviruses and Reverse Transcriptase
Retroviruses
- Replicate unusually
- RNA serves as template for DNA synthesis
- Complementary synthesis of DNA by RNA-dependent DNA polymerase reverse transcriptase
reverse transcriptase
- RNA-dependent DNA polymerase enzyme
Nucleotides
- DNA is nucleic acid
- Nucleotides: building blocks of nucleic acid
- nucleotides are building blocks of DNA
Nucleotides consist of:
- nitrogenous base (two kinds)
- pentose sugar
- phosphate group
Nitrogenous bases
- two kinds of nitrogenous bases
-- purines (9 member ring)
--- adenine (A)
--- guanine (G)
-- pyrimidines (6-member ring)
---Cytosine (C)
--- Thymine (T)
--- Uracil (U)
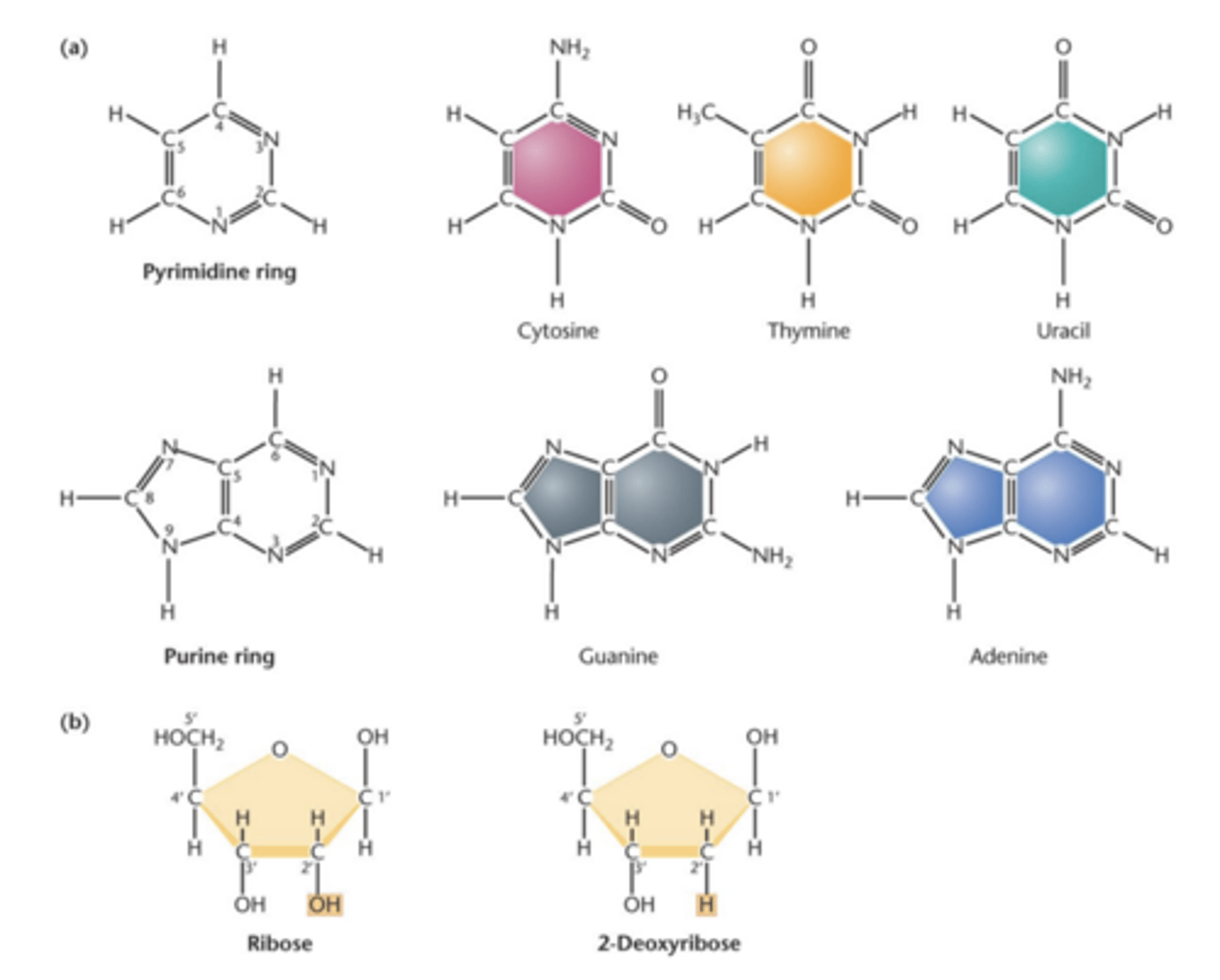
Bases of DNA and RNA
- DNA
--- based A, C, , G
--- DNA contains deoxyribose ('deoxy' (without an oxygen)
- RNA
--- bases A, C, , G
--- RNA contains ribose sugar
- only DNA contains T
- only RNA contains U
Nucleoside
- contains nitrogenous base and pentose sugar
- molecule is composed of purine or pyrimidine base and ribose or deoxyribose sugar
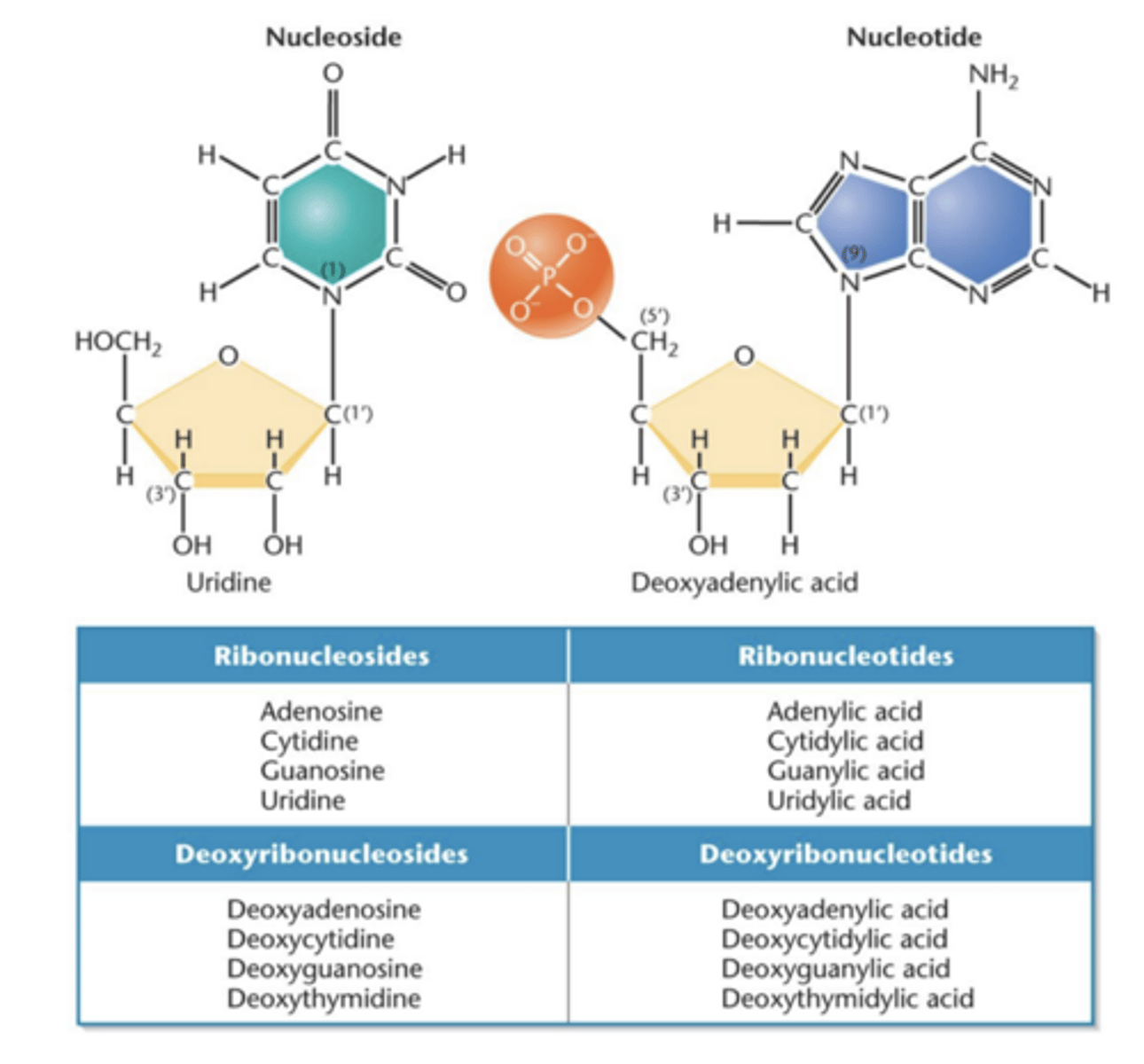
Nucleotide
- nucleoside with phosphate group added
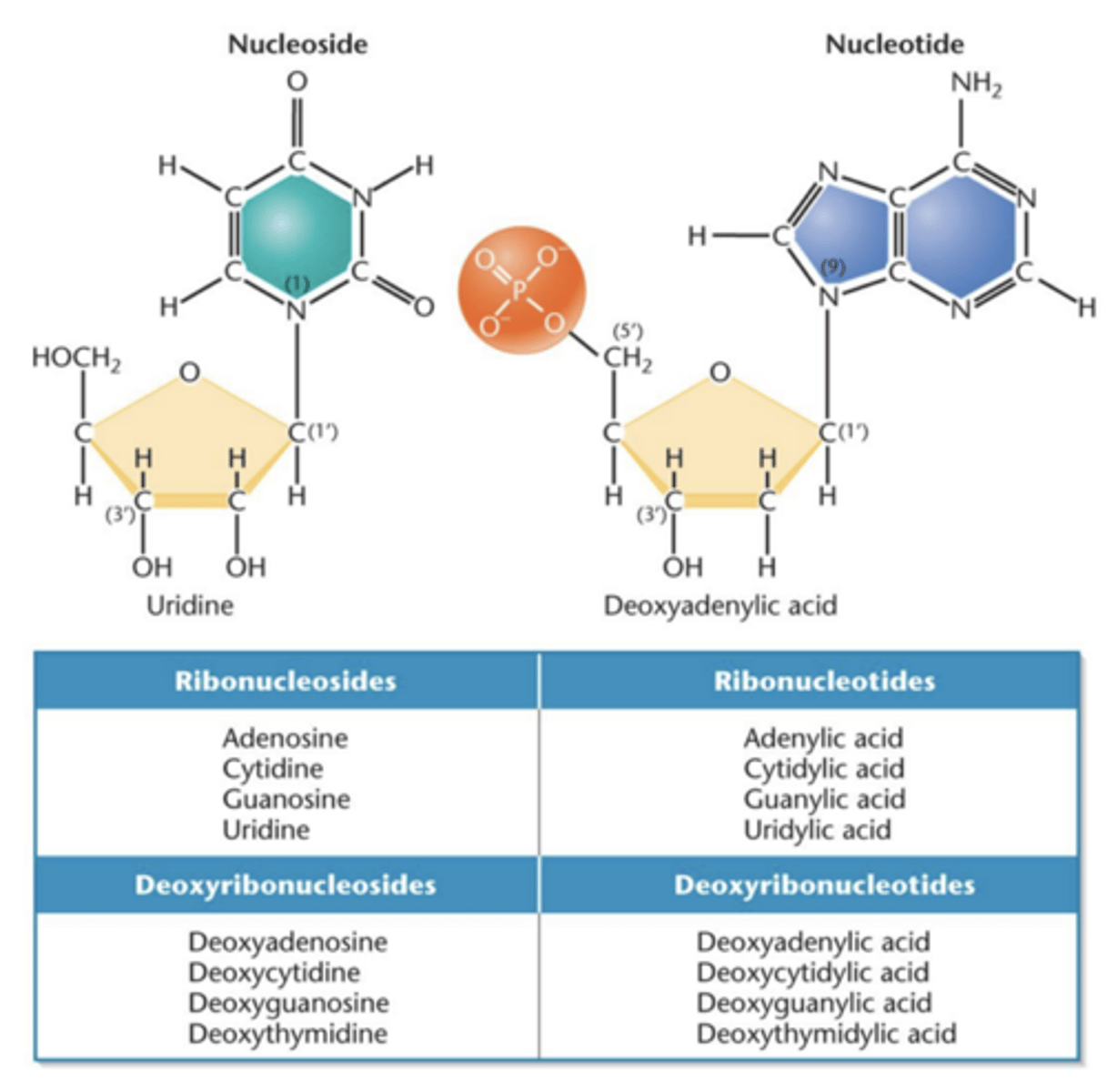
Nucleoside monophosphate (NMP)
- a nucleotide
Nucleoside diphosphate (NDP)
- nucleotide with addition of two phosphate groups
nucleoside triphosphate (NTP)
nucleotide with addition of three phosphate groups
triphosphate
serve as precursor molecule during nucleic acid synthesis
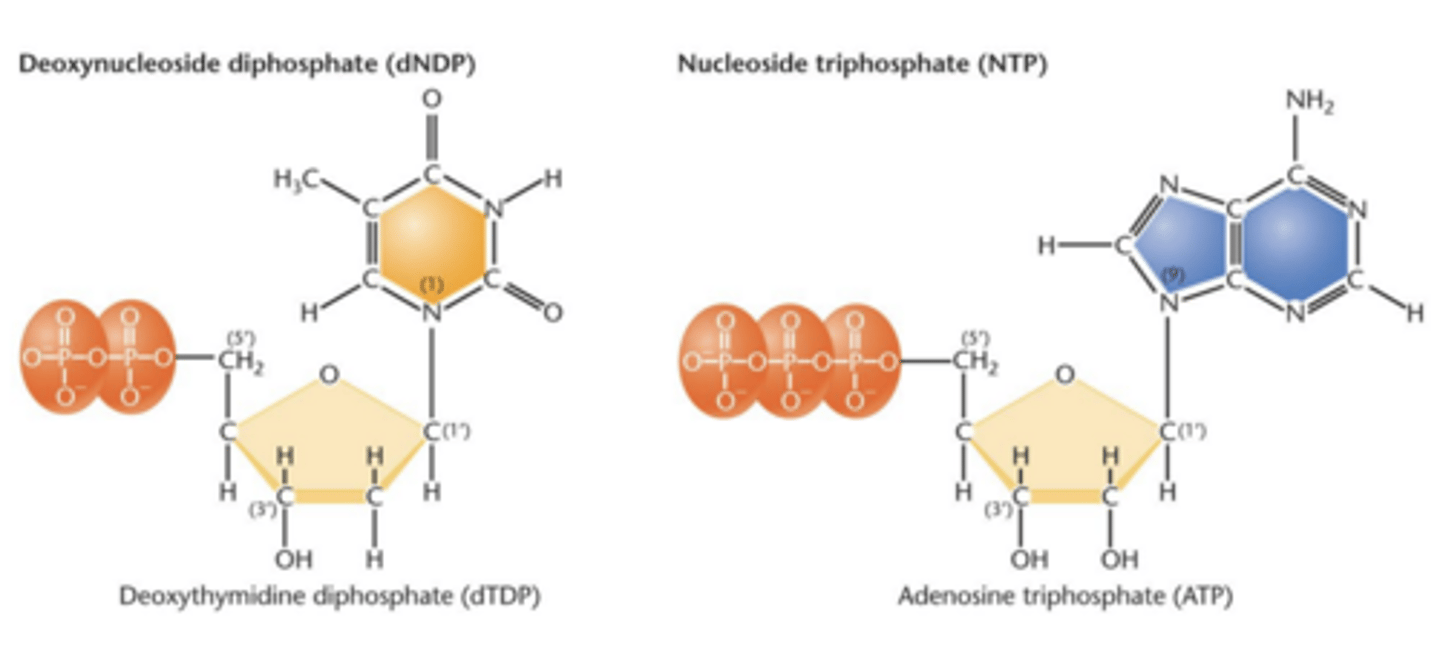
ATP and GTP
adenosine triphosphate and guanine triphosphate
- large amount of energy involved in adding/removing terminal phosphate group
phosphodiester bond
nucleotides are linked by phosphodiester bonds between phosphate group at C-5' position and OH group on C-3' position
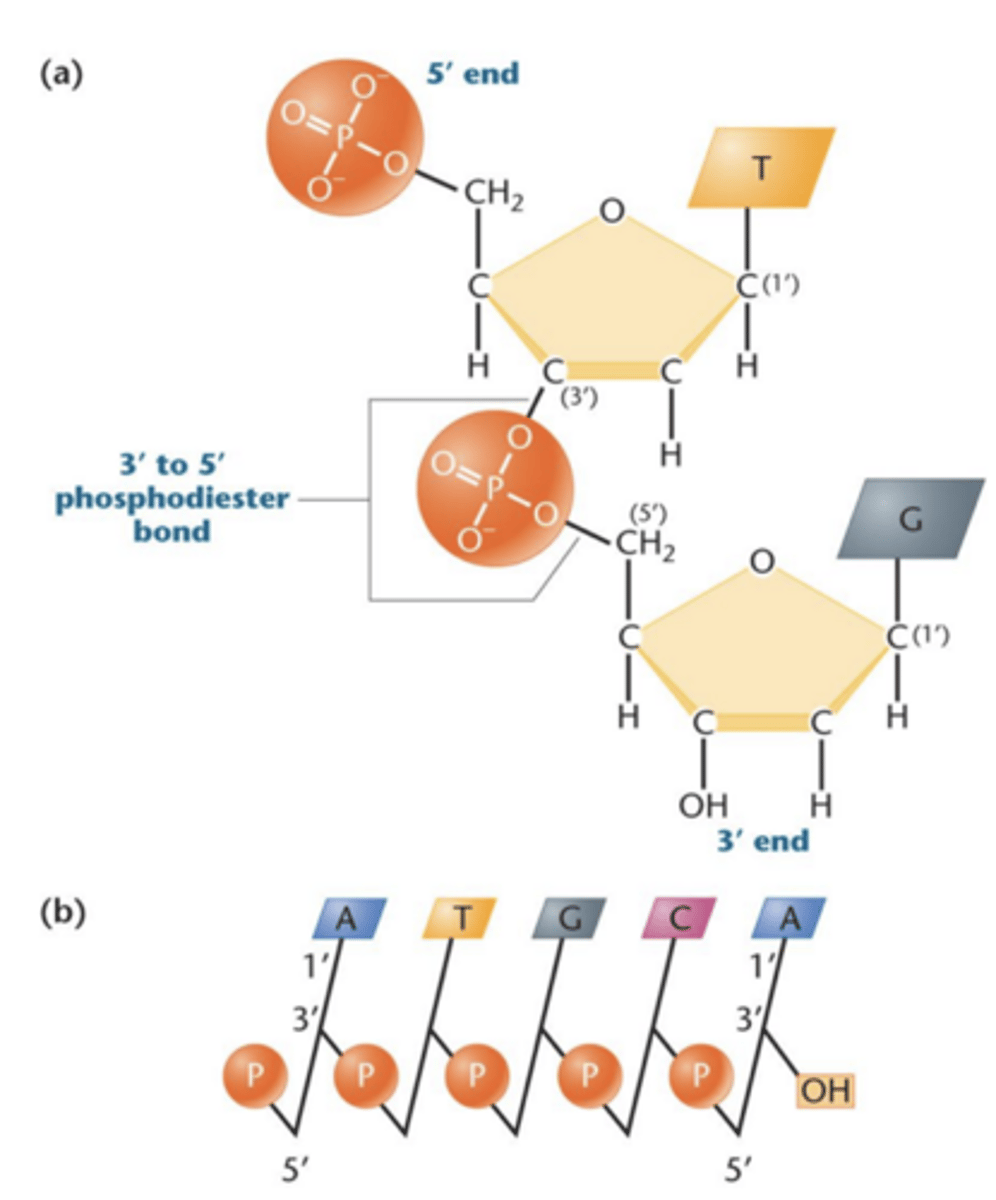
Structure of DNA
Watson and Crick, 1953
- proposed structure of DNA as a double helix
proposed as
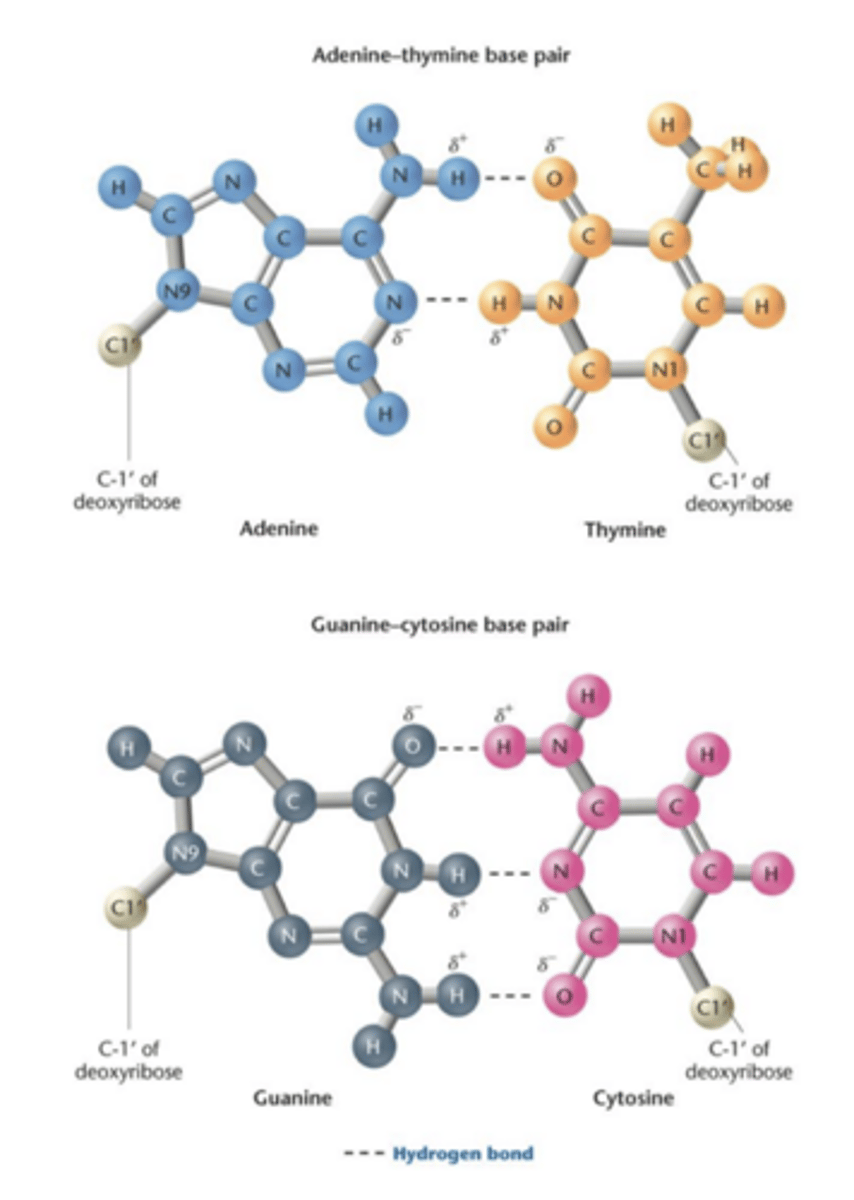
Chargaff (1949-1953)
- proposed base composition
- amount of A is proportional to T
- amount of C is proportional to G
- Percentage of C+G does not equal percentage of A+T
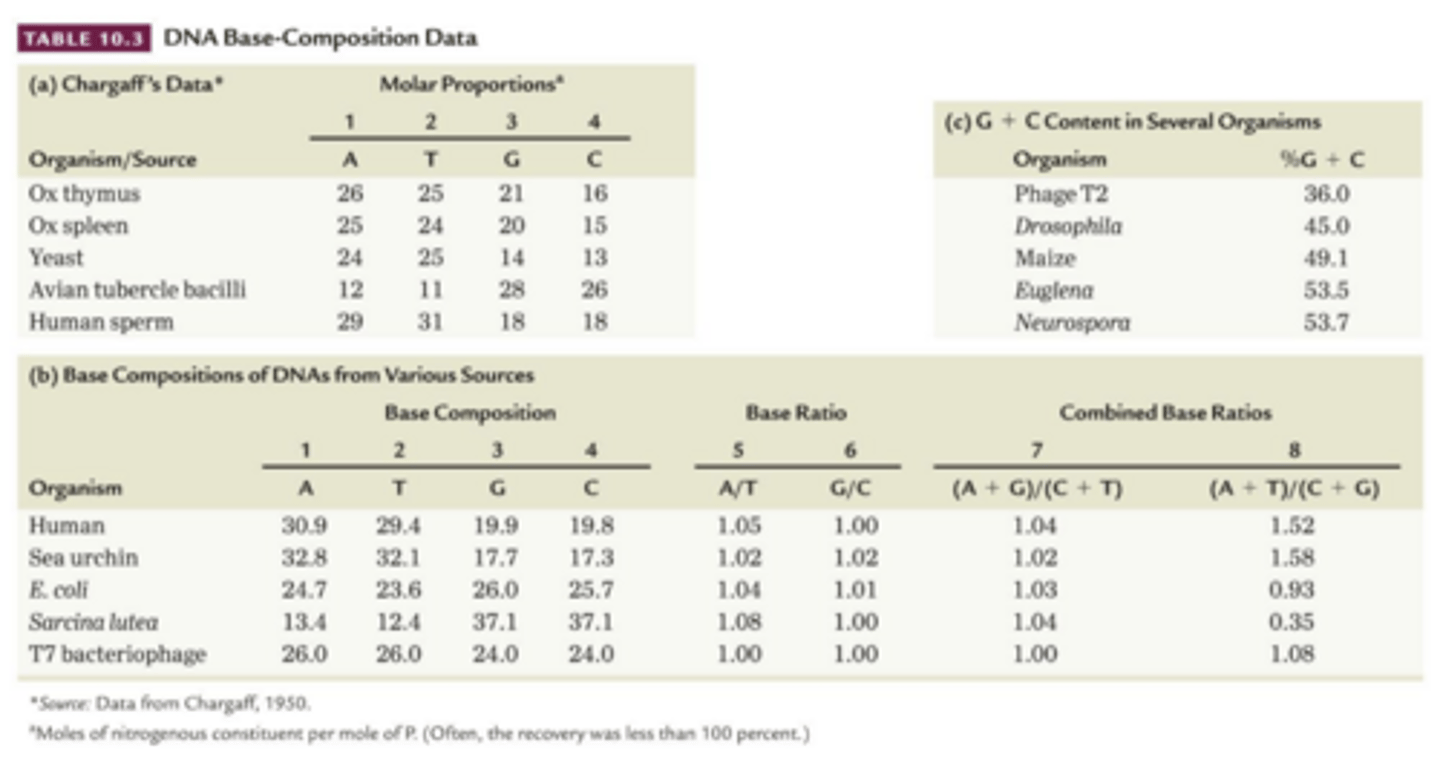
X-Ray Diffraction
base composition analysis (Chargaff) and X-ray diffraction provided crucial data to Watson and Crick
- studies by Rosalind Franklin showed DNA had a 3.4 angstrom periodicity, characteristic of helical structure
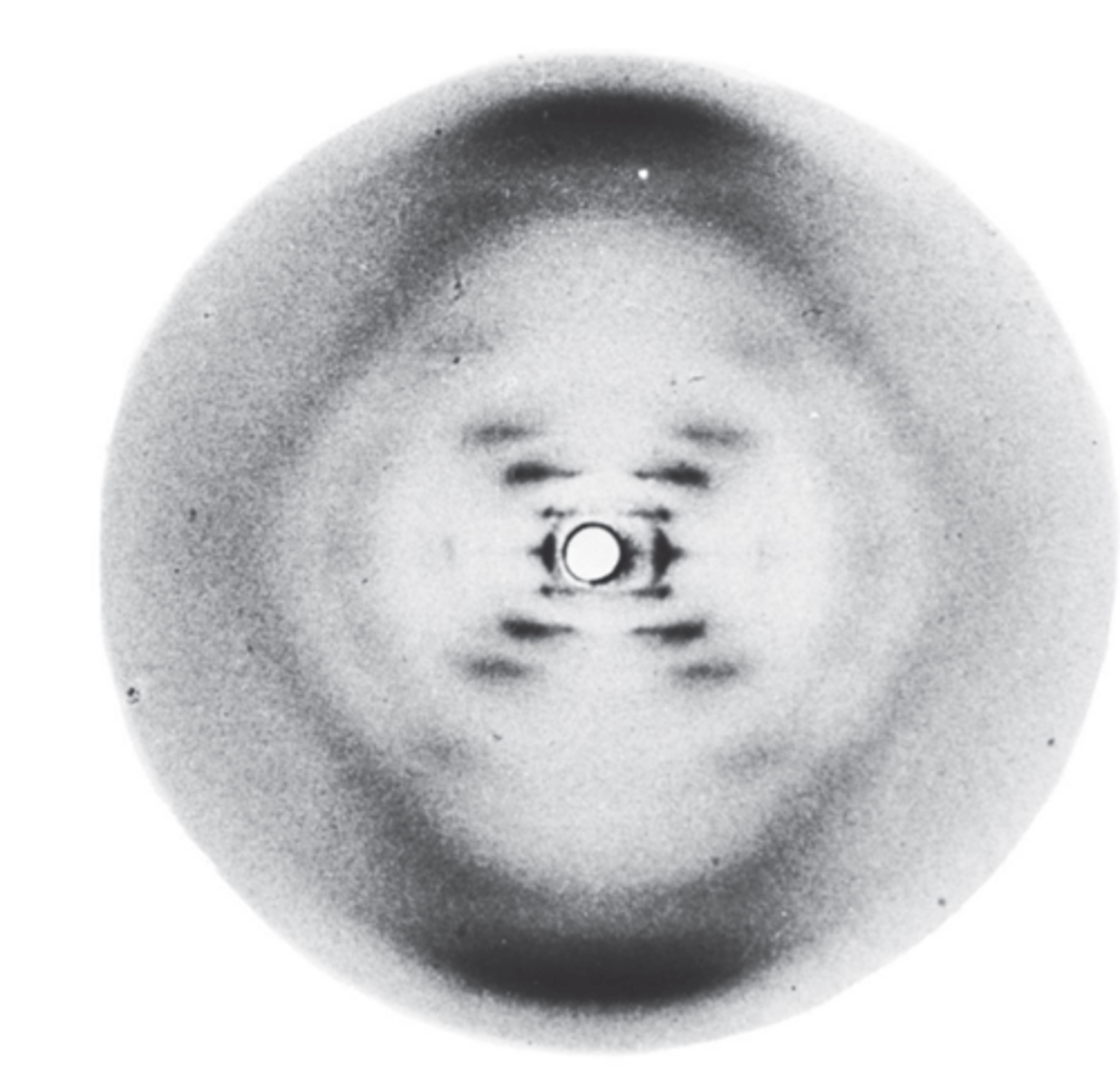
Model of structure
- Double helix
- two anti-parallel strands connected by base pairing
- stacked nitrogenous bases
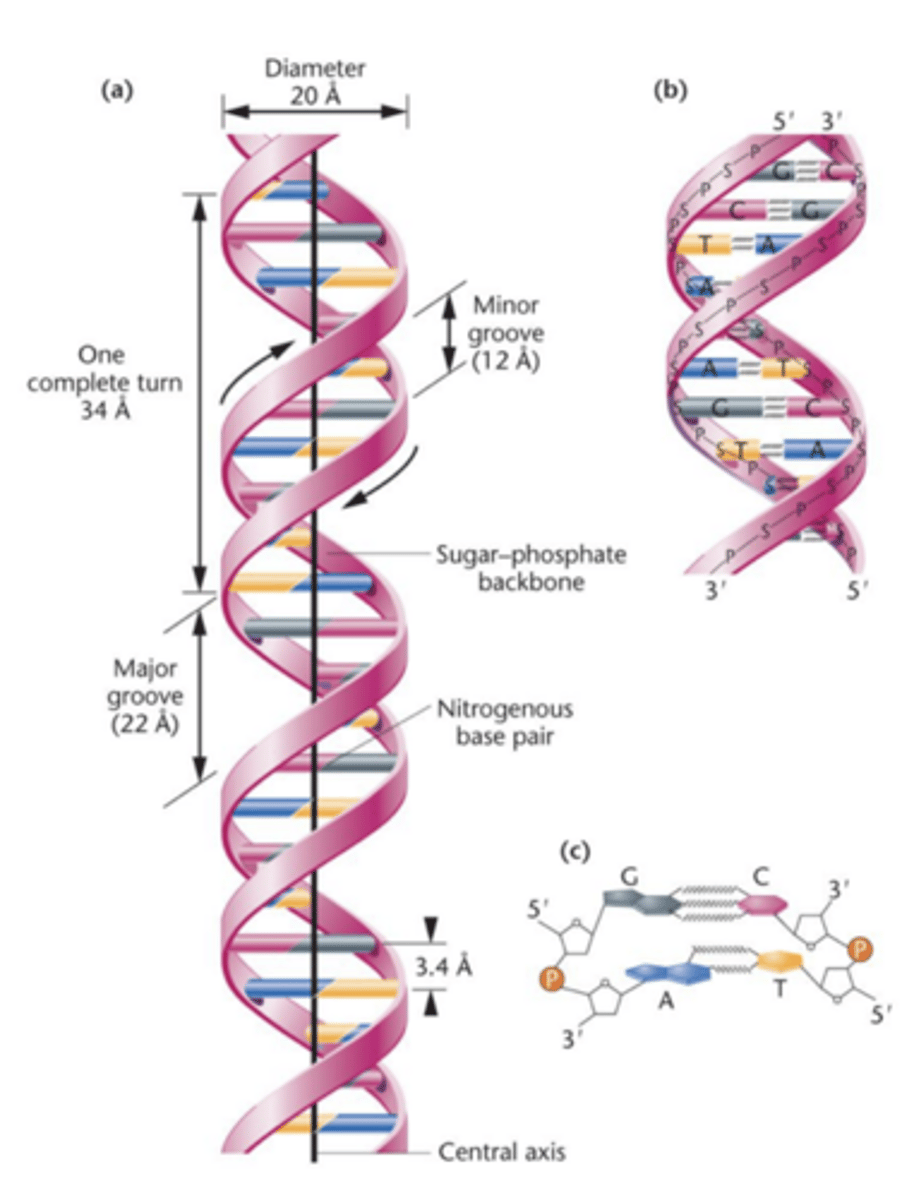
Base-pairing - hydrogen bonds
chemical affinity produces hydrogen bonds in pair of bases
- A-T and G-C base pairing provides complementarity of 2 strands and chemical stability to the helix
- A-T: double bond
- G-C: triple bond
Semiconservative model
storage of genetic information in sequence of bases
- mutations or genetic changes that could result in alteration of bases
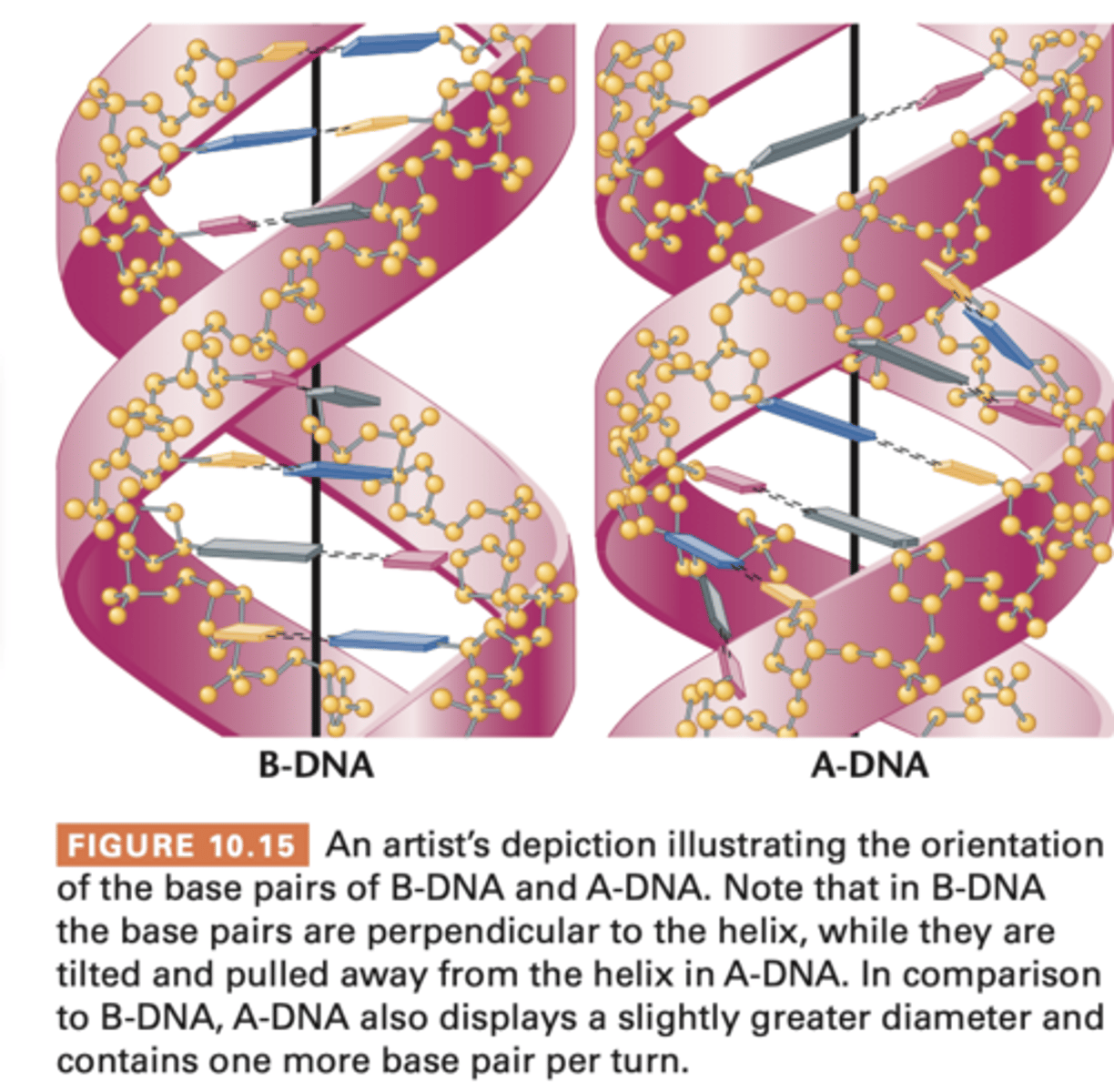
Alternative forms of DNA
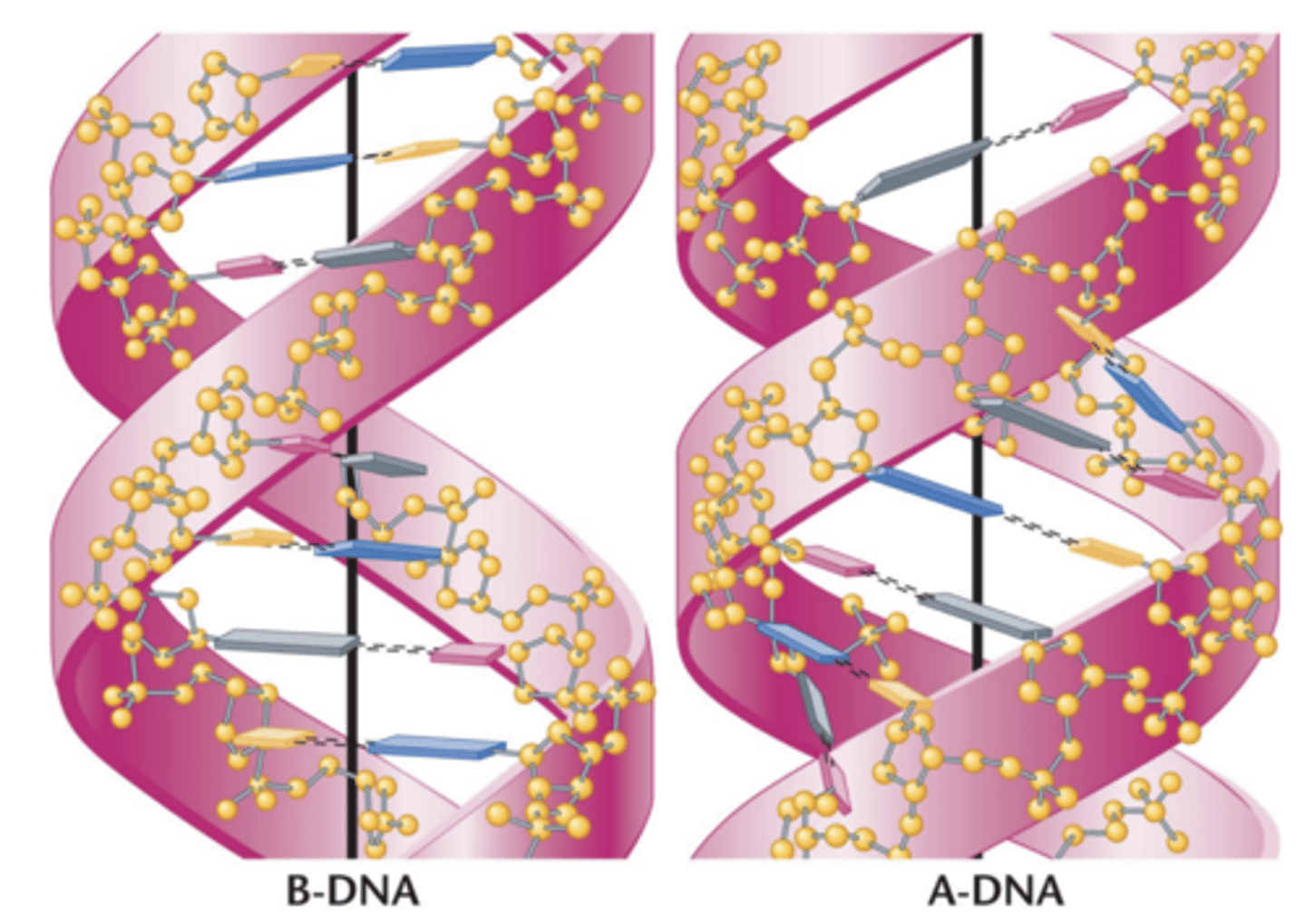
Watson-crick DNA model of B-DNA
- seen under aqueous, low-salt conditions
alternative forms of DNA
- different conformations of DNA observed under different conditions of isolation
Under different conditions of isolation, different conformations of DNA are seen
A-DNA
-- slightly more compact than B-DNA
-- prevalent under high-salt or dehydration conditions
C-DNA, D-DNA, E-DNA, and P-DNA
-- right-handed forms of DNA
-- less compact than B-DNA
D-DNA and E-DNA
-- lack guanine
Z-DNA
-left handed double helix
structure of RNA
-sugar ribose replaces deoxyribose of DNA
- nitrogenous base uncial replaces thymine of DNA
- most are single-stranded (ss)
-- exception: animal viruses have double-stranded helices
Three classes of RNA
three classes of cellular RNAs (function during gene expression)
- messenger RNA (mRNA)
- ribosomal RNA (rRNA)
- transfer RNA (tRNA)
originate as complementary copeies of one of two DNA strands during transcription
Three major classes of RNA
rRNAs: ribosomal RNAs
- structural components of ribosomes for protein synthesis
mRNAs: messenger RNAs
- template for protein synthesis
- carry genetic information from gene to ribosome
tRNAs: transfer RNAs
- carry amino acids for protein synthesis
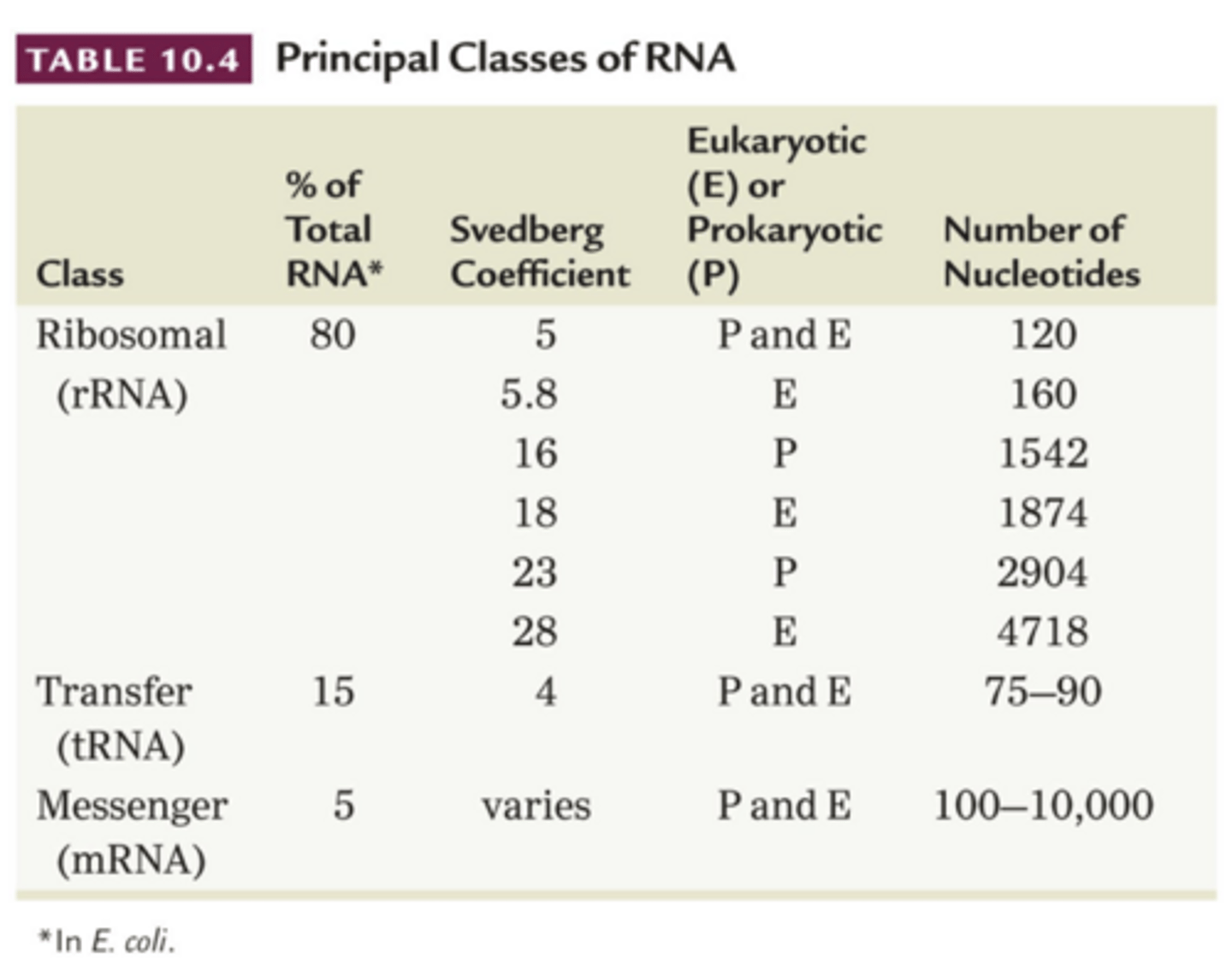
Unique RNAs in Eukaryotes
Unique RNAs
telomerase RNA and RNA primers
- involved in DNA replication at chromosome ends
SnRNA: small nuclear RNA
- process in mRNAs
Antisense RNA, microRNA, siRNA
- involved in gene regulation
Analytical techniques useful during DNA and RNA investigation
- absorption of UV light
- denaturation and renaturation of nucleic acids
- molecular hybridization
- FISH: Fluorescent in situ hybridization
- Reassociation kinetics
- electrophoresis of nucleic acids
Absorption of UV light
nucleic acids
- absorb UV light strongly at 254-260 nm due to interaction between UV light and ring systems of the bases
- UV light used in localization, isolation, and characterization
- use of UV critical to isolation of nucleic acids following separation
Denaturation and renaturation of nucleic acids
DNA can denature due to heat or stress
hyperchromic shift
- increase in UV absorption of heated DNA in solution
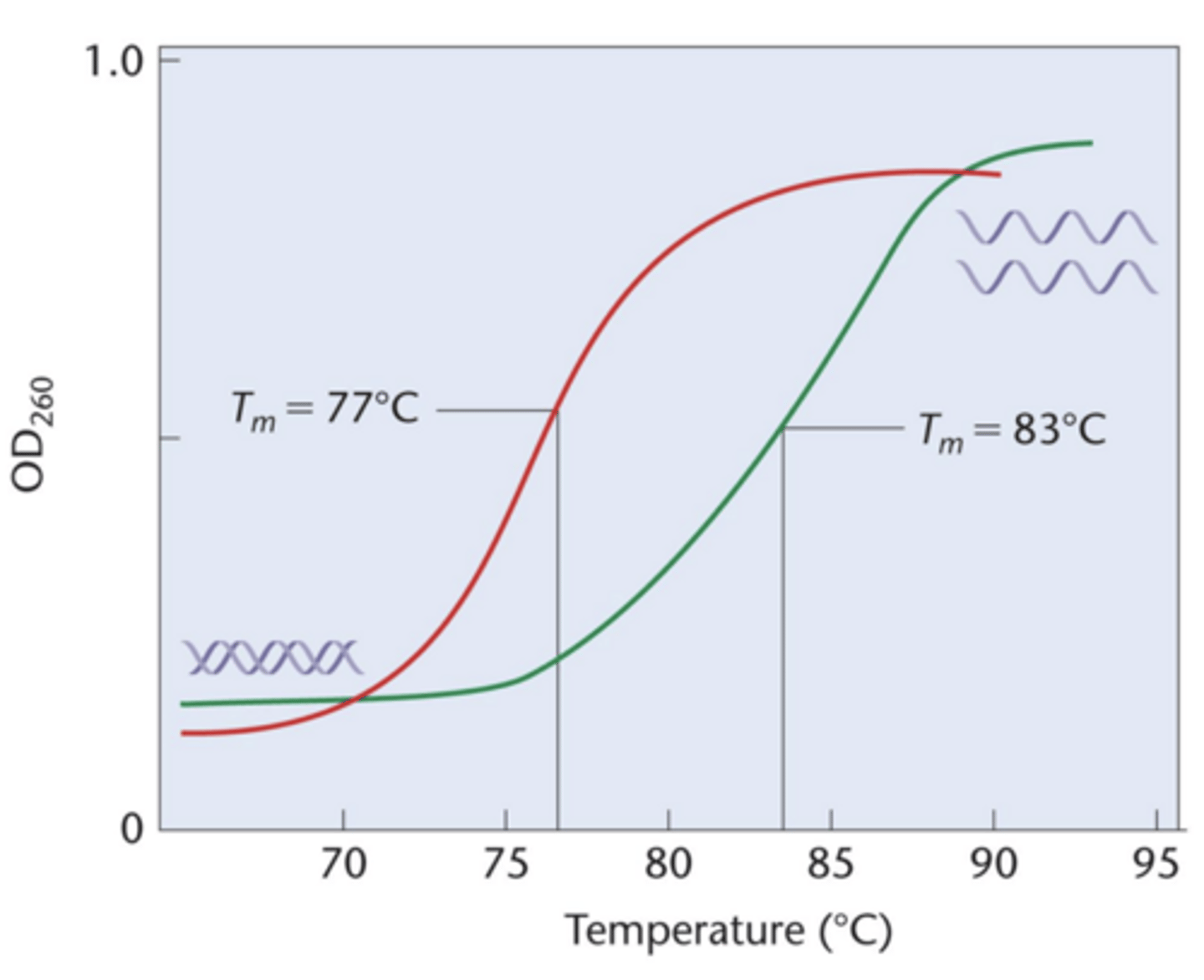
- denaturation used to determine melting temperature (Tm)
- Graph of absorption vs temperature gives melting profile of DNA
Molecular hybridization
- denaturation and renaturation of nucleic acids are the basis for molecular hybridization
- example: single strand of nucleic acids combine duplex structures, yet are not from the same source
-- if DNA is isolated from two distinct sources, double-stranded hybrids will form
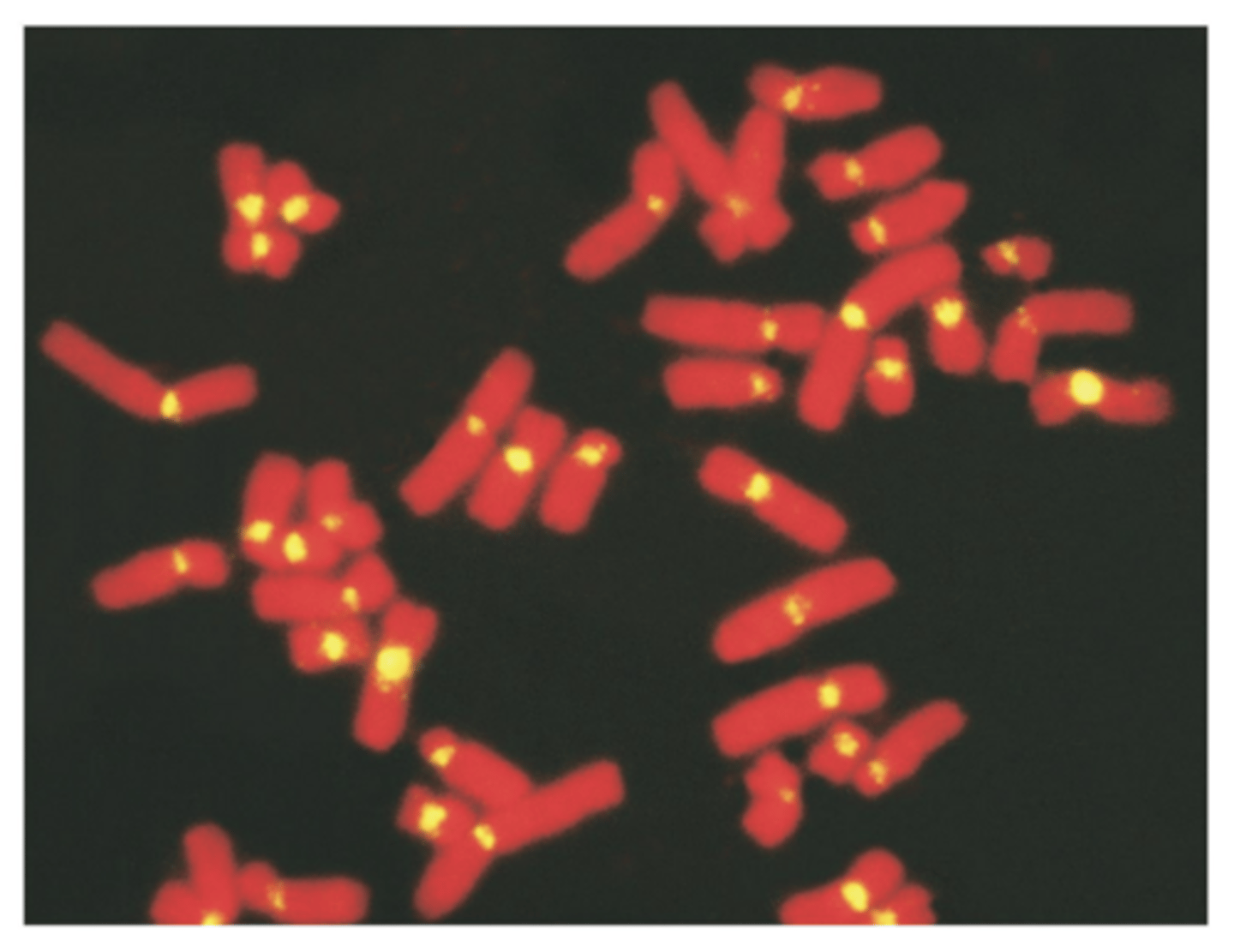
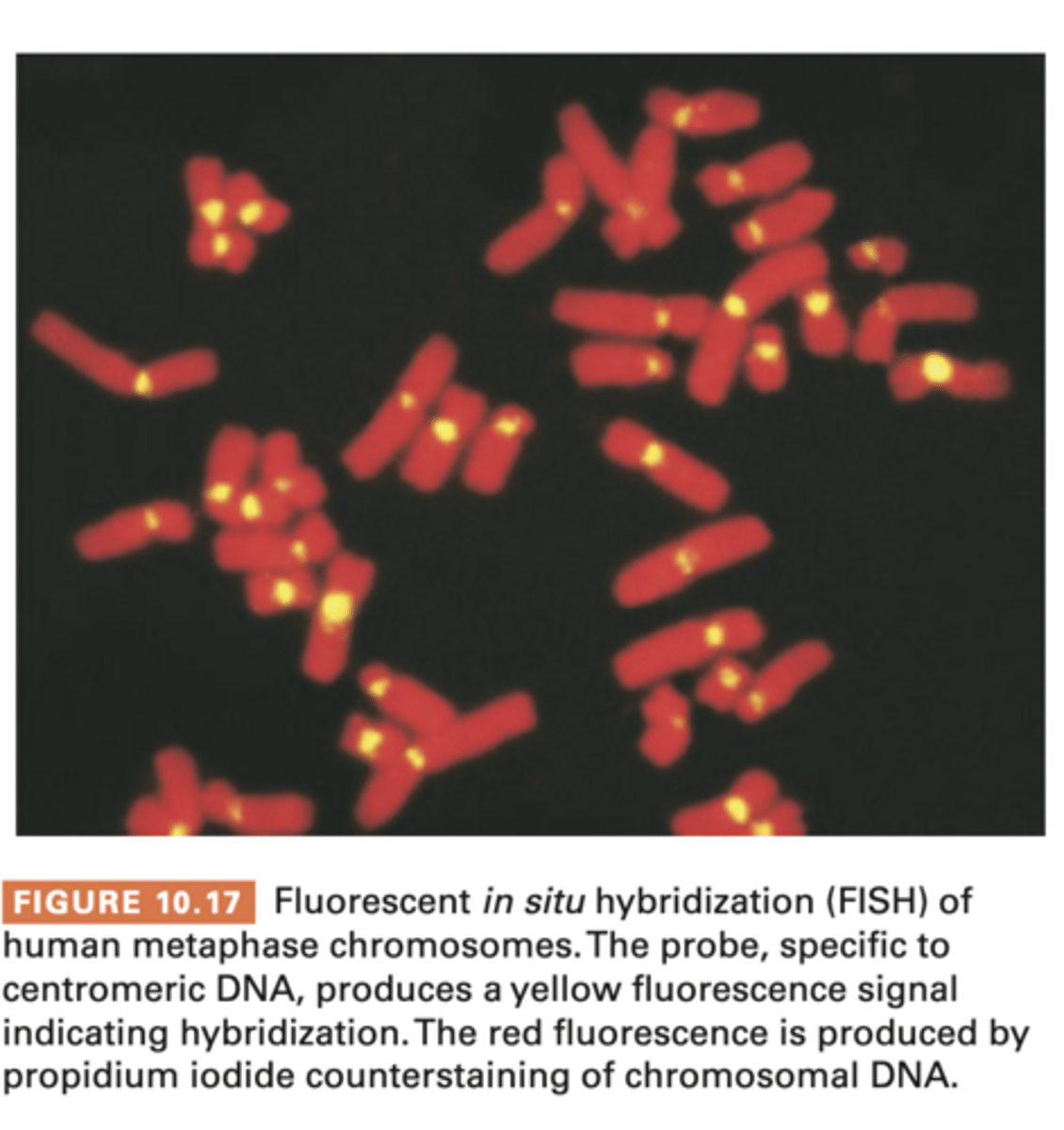
FISH: Fluorescent in situ hybridization
- uses fluorescent probes to monitor hybridization
- mitotic cells fixed to slide and subjected to hybridization
- ssDNA is added and hybridization is monitored
- probes are nucleic acids that will hybridize only with specific chromosomal areas
Reassociation kinetics
- analyzes rate of reassociation of complementary single DNA strands
- DNA fragmented into small defragmented/dissociated pieces
- reassociation of ssDNA provides information on size and genomic complexity of organism
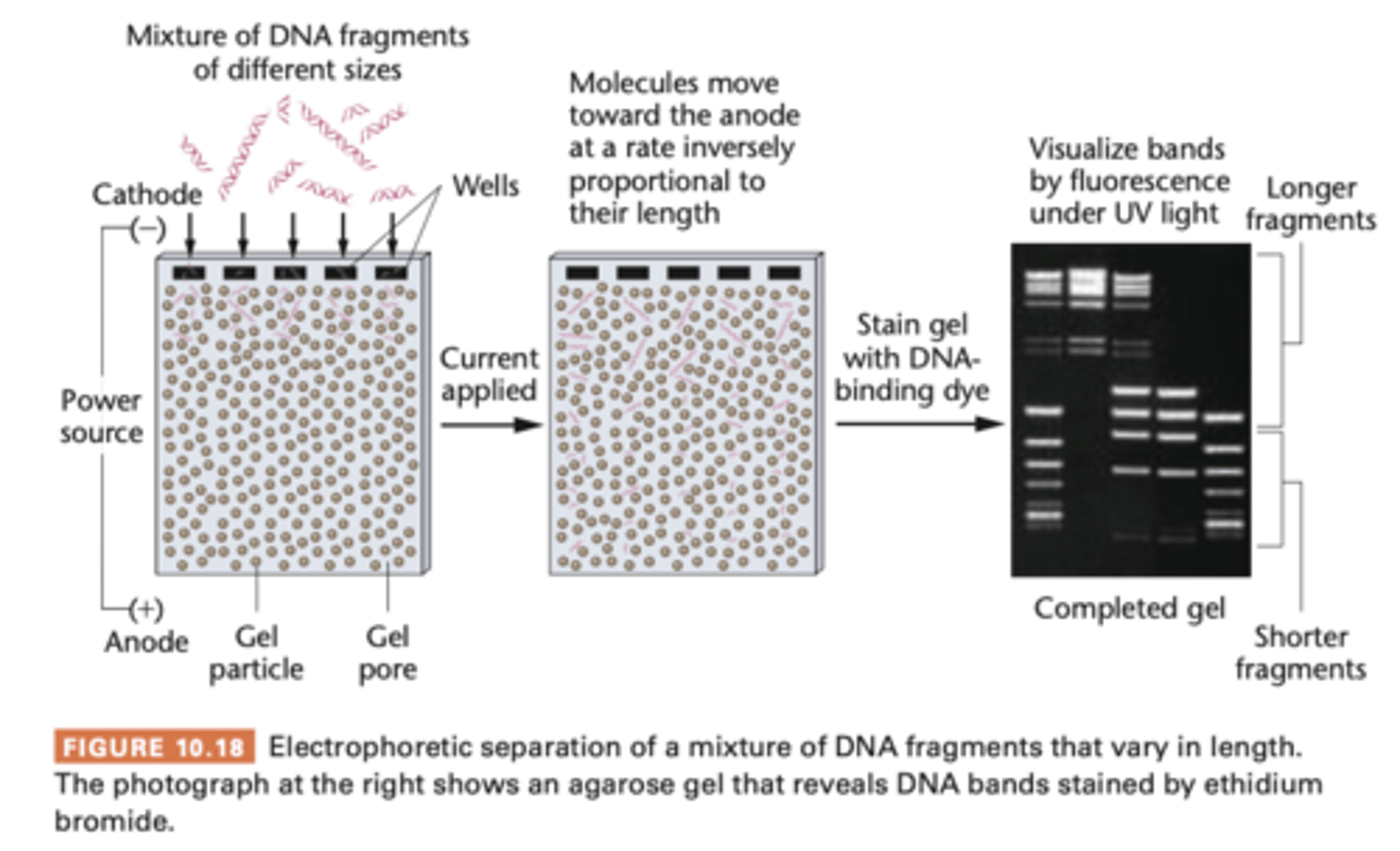
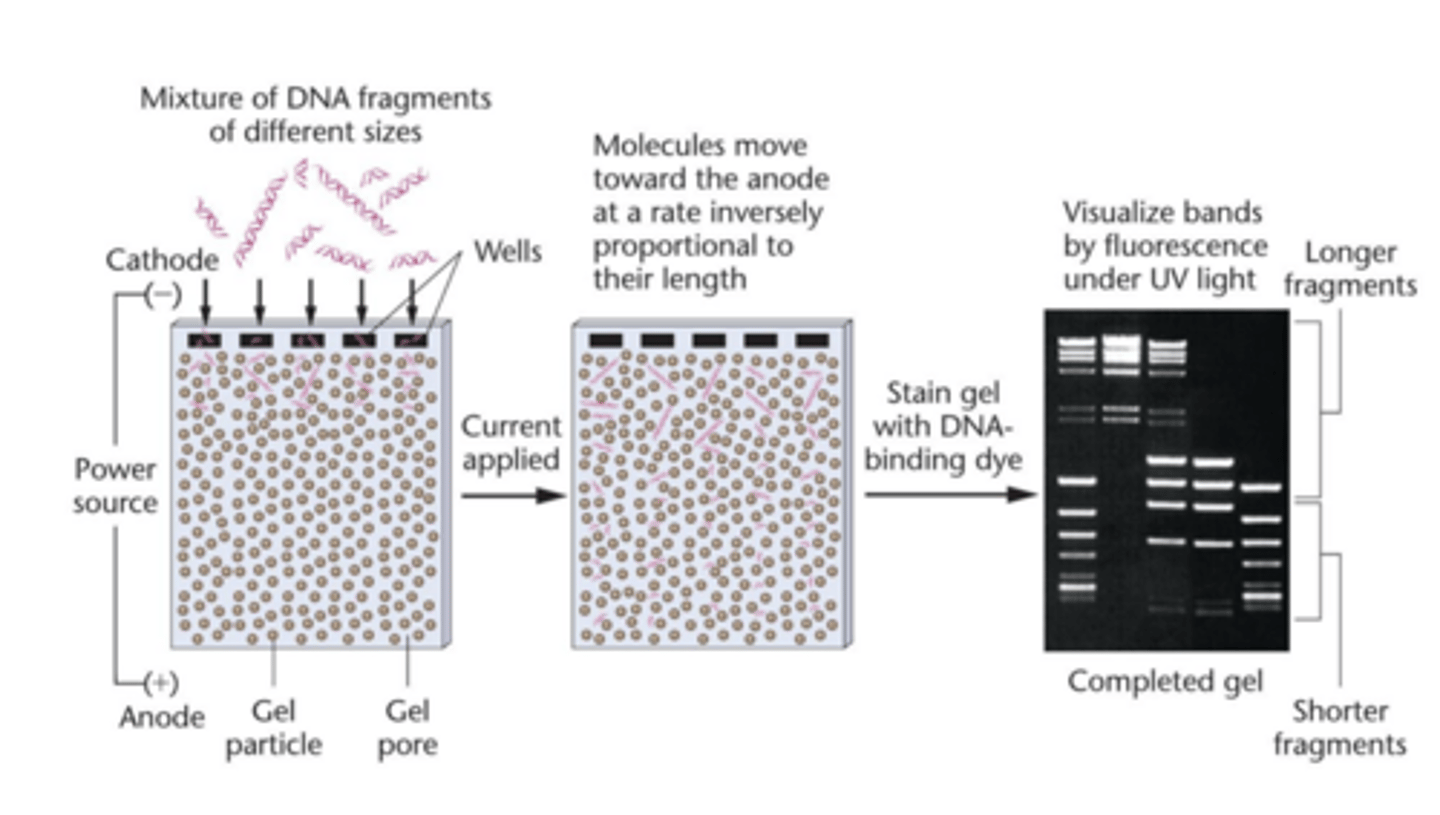
Electrophoresis
nucleic acid electrophoresis
- separates DNA and RNA fragments by size
- smaller fragments migrate through gel at faster rate than larger fragments
agarose gel
- porous matrix restricts migration of larger molecule more than it restrict smaller ones
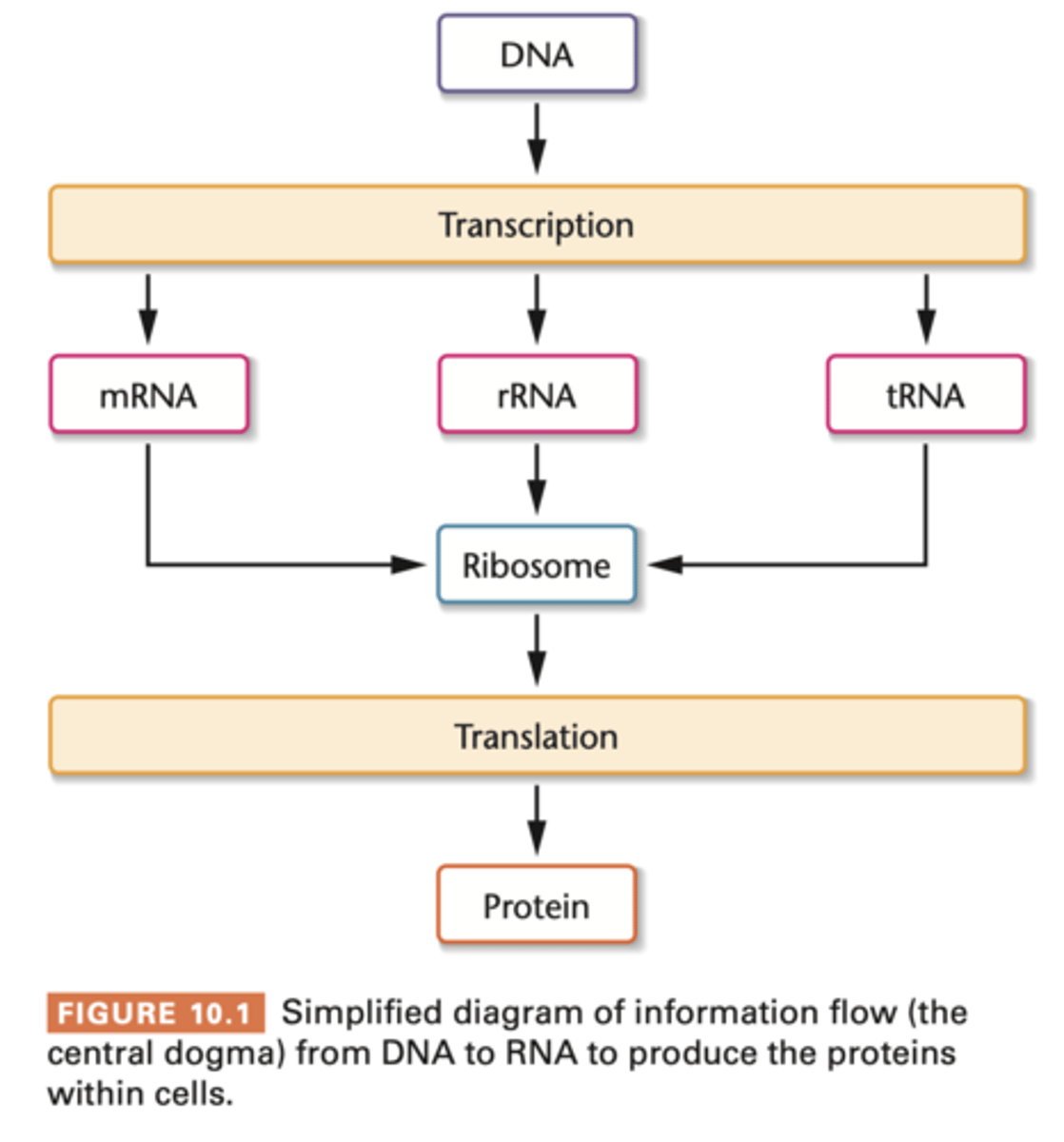
Information flow
Until 1944, observations favored protein as
the genetic material
- proteins were known to be diverse and abundant in cells, and most researched topics
- nuclein: acidic substance, now known to contain DNA, from isolating cell nucleic, and derived an acidic substance, now known to have DNA
Levene observation
DNA contained approximately equal amounts of four similar molecules called nucleotides
Was wrong about identical bases repeated over and over = basis for tetra nucleotide hypothesis
Evidence favoring DNA as the genetic material was first obtained during the
study of bacteria and bacteriophages
Transformation: Early Studies
Worked with strains of bacterium Diplococcus pneumoniae
- some virulent (with capsule), infectious, strains that cause pneumonia in vertebrates and some avirulent (noninfectious strains) - don't cause illness
Nonencapsulated bacteria - readily engulfed and destroyed by phagocytic cells in host animal's circulatory system, virulent bacteria with coat, don't get engulfed and they multiply and cause pneumonia
encapsulated bacteria make smooth, shiny surfaced colonies (S), when grown on an agar plate; non encapsulated strains make rough colonies (R). therefore, easily distinguished
each strain is type of serotypes, differ in the precise chemical structure of polysaccharide of thick, slimy capsule (identified by immunological techniques, label w roman numberals) ie: types I and II are most common causing pneumonia
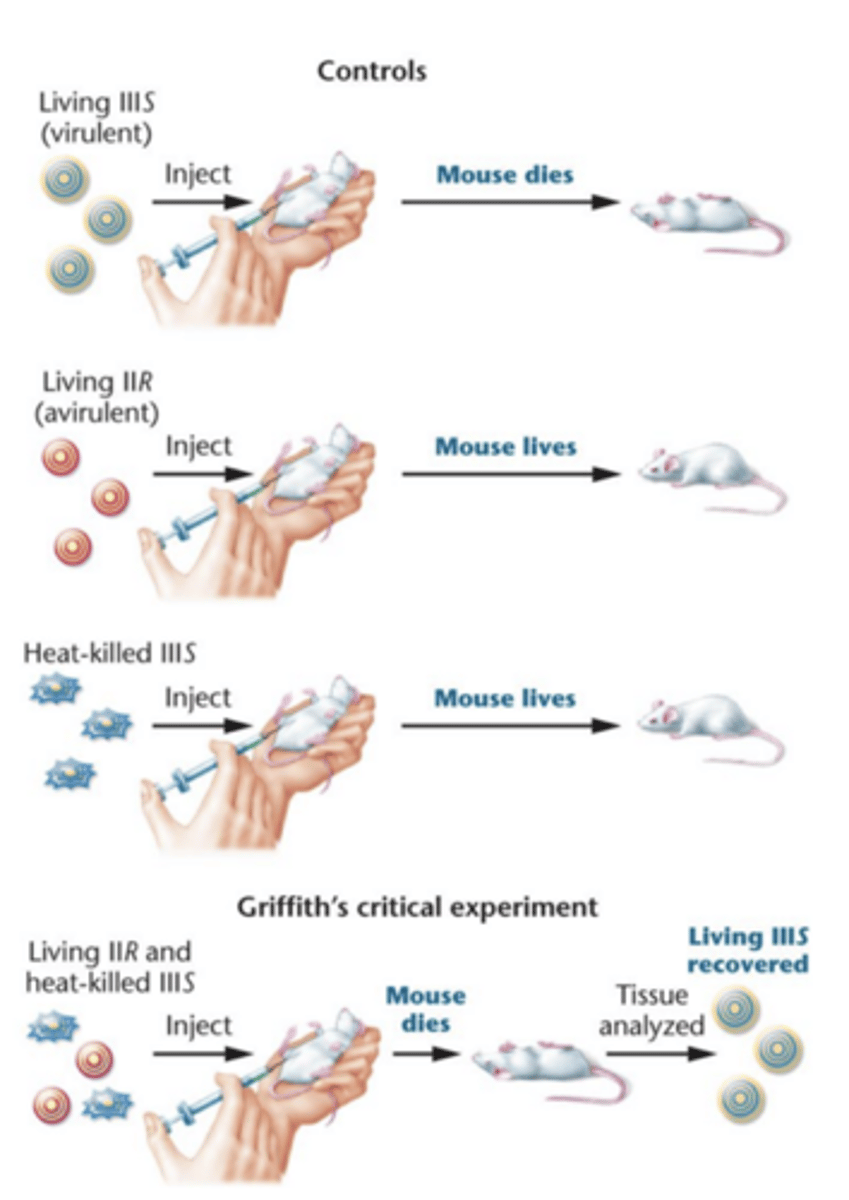
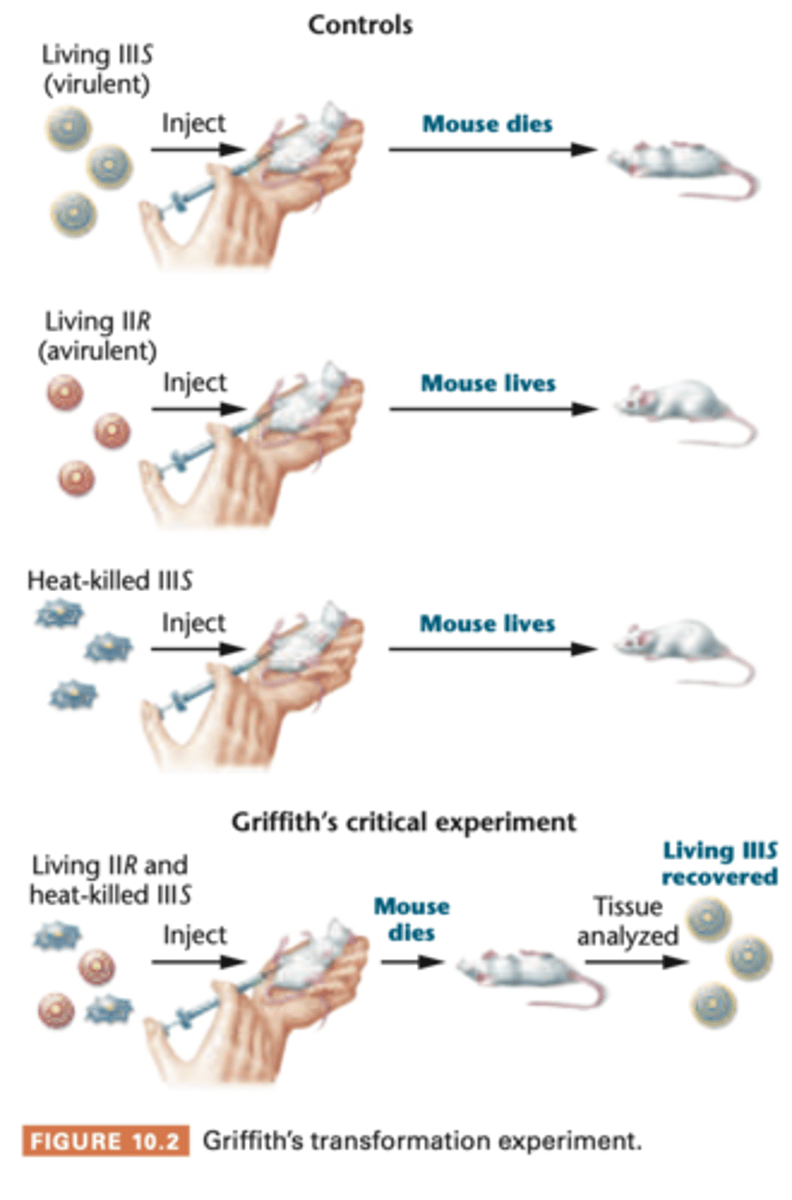
Griffith experiment
in this study, used IIR and IIIS
injection into mice of living IIR (avirulent) cells combind with heat-killed IIIS (virulent). Alone, they did nothing to mice, but together the mice were killed
-blood test showed large number of living type IIIS (virulent) bacteria
IIIS bacteria were identical to IIIS stain from heat-killed cell perp had been made --> must hav interaction
transformation --> transforming principle might be some part of polysaccharide capsule or compound required for capsule synthesis, although alone did not cause pneumonia
other studies showed, injection to mice was not necessary for transformation bc made it in vitro
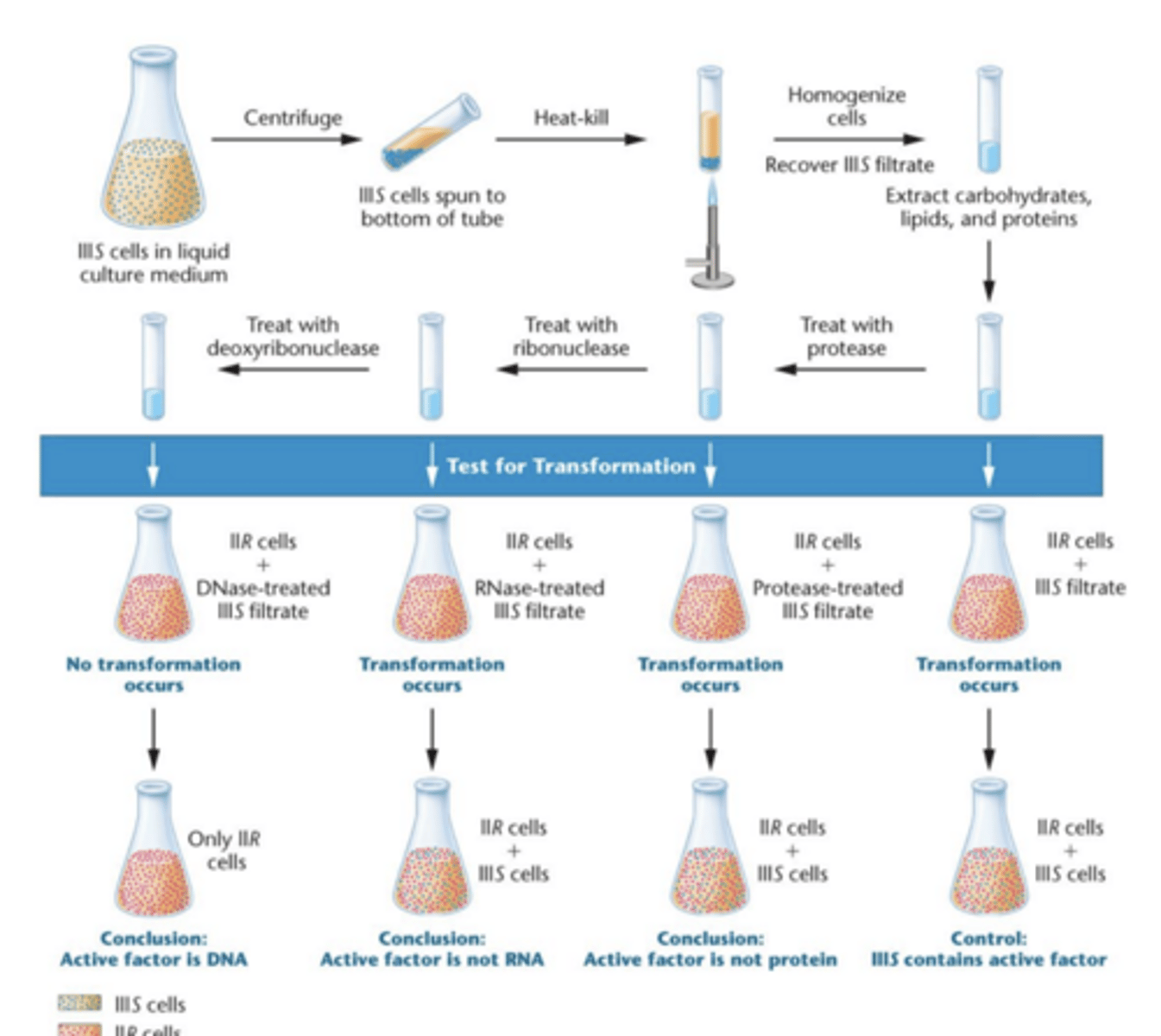
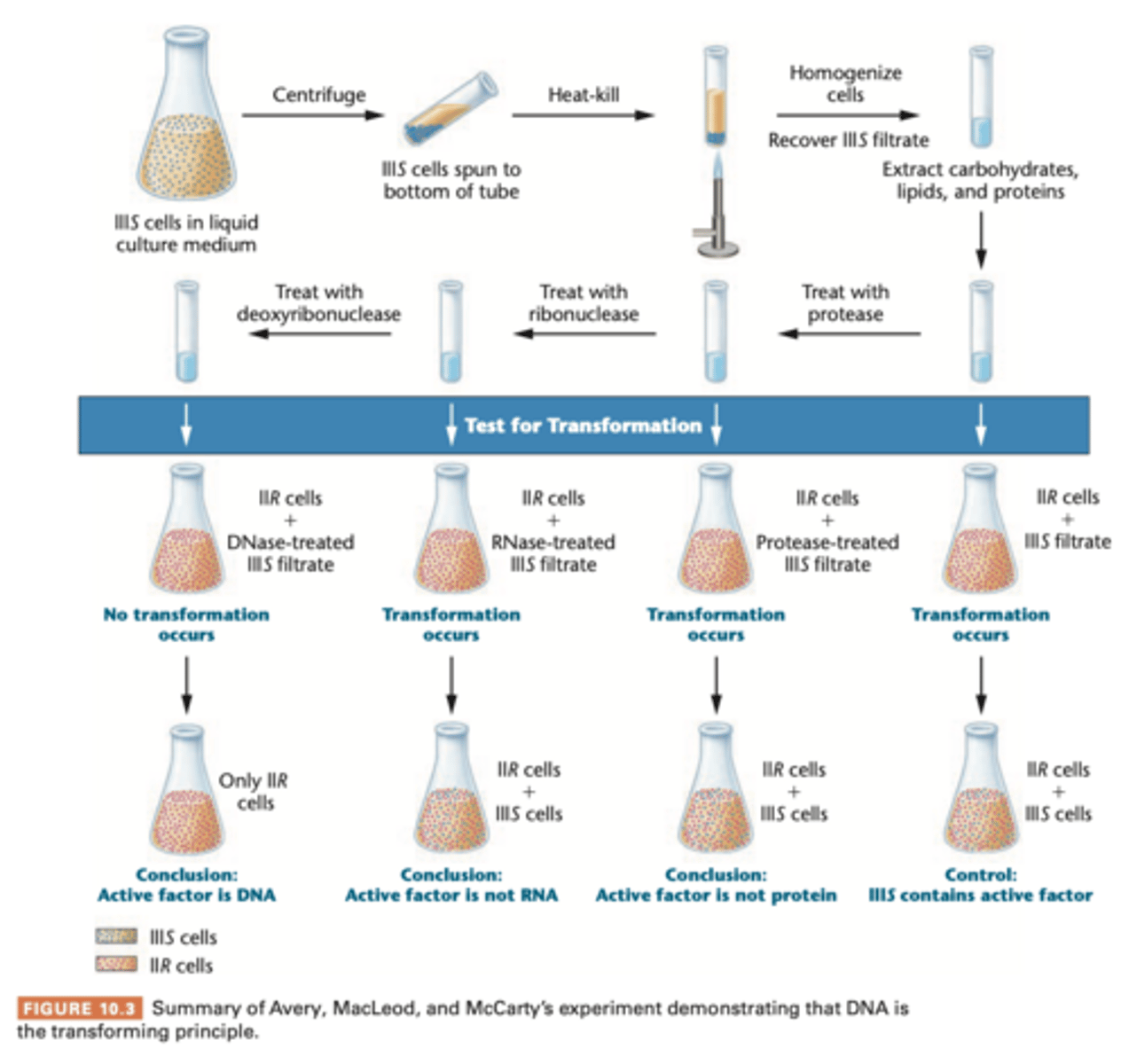
Transformation: Avery Macleod and McCarty experiment
- They used liquid cultures of IIIS virulent
- extractions with detergent deoxycholate (DOC)
- extracted proteins from the IIR avirulent cells and it still transformed == DNA
- analyzed fibrous mass first analyzed for nitrogen:phosphorous ratio, coincide with ratio of 'sodium desoxyribonuucleate" (AKA DNA back then)
to further verify, treated with proteolytic enzyme trypsin and chymotrypsin and then with RNA-digesting enzyme (ribonuclease RNase) which further destroyed activity of proteins and RNA
transforming activity remained, chemical testing confirmed DNA
final test = use samples of DNA digesting enzyme deoxyribonuclease (DNase), which isolated from dog and rabbit = destroyed the transforming activity
& once transformaron occurs, the capsular polysaccharide is made in successive generations, it is therefore heritable and affects genetic material
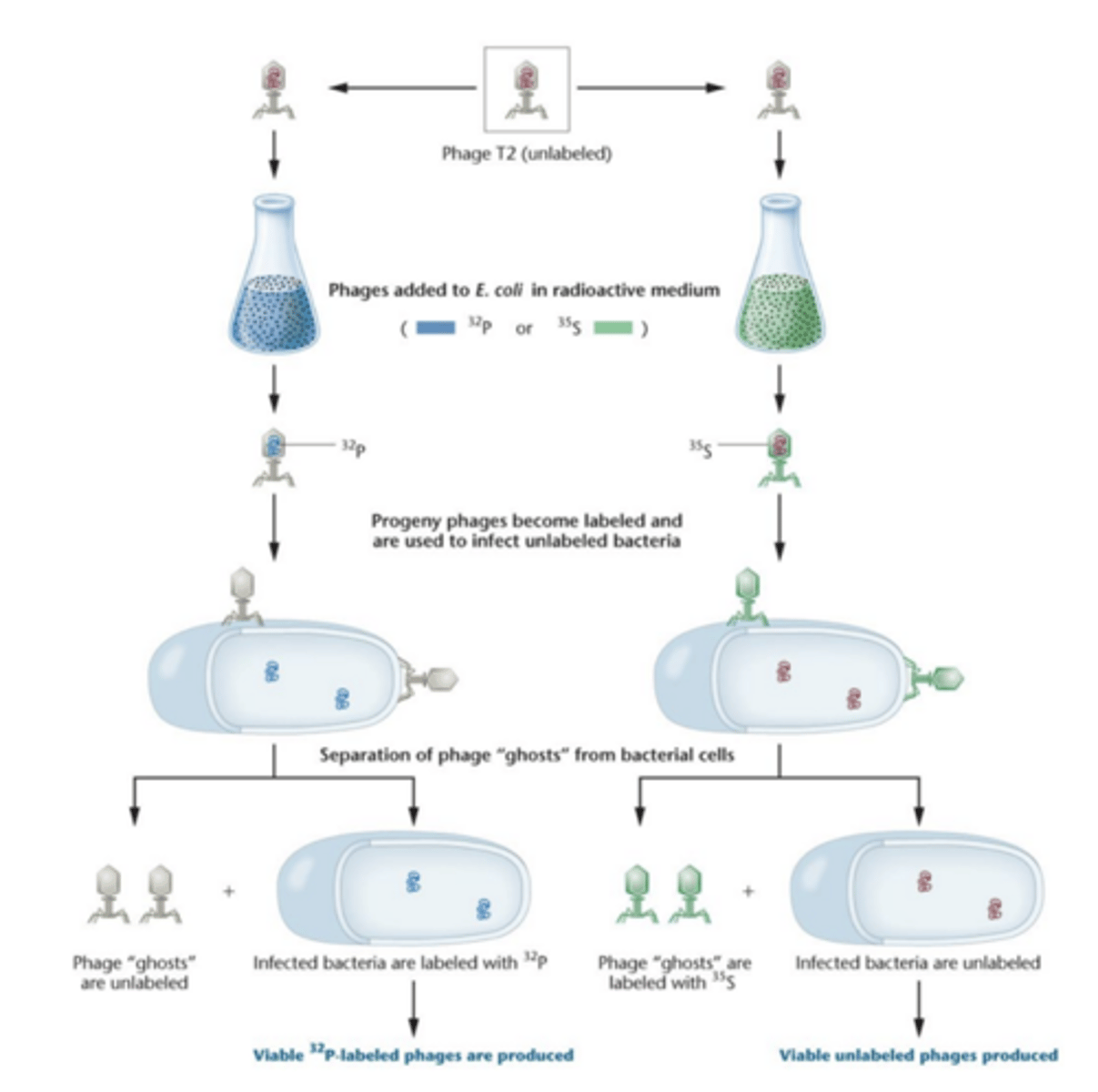
The Hershey-Chase Experiment- t bacteriophage
- study with bacteriophage T2 (protein core surrounding core of DNA)
- electron micrograph show the phage's external structure is made of icosahedral heat plus a tail, phage adsorbs to bacterial cell, and some component of the phage enters cell causing viral reproduction, causes bacterial cell lysis - lytic cycle
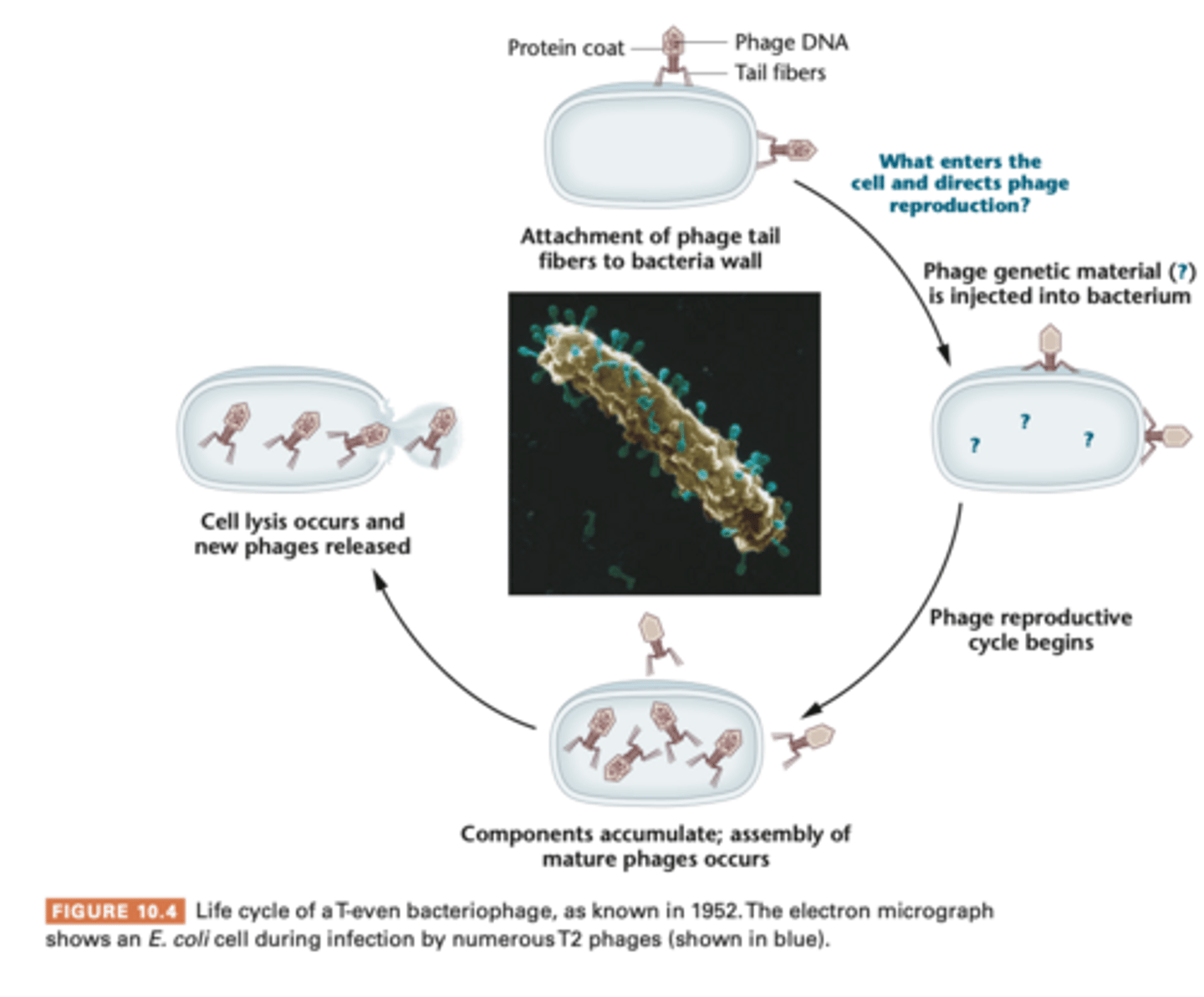
Hershey-Chase Experiment
- new some molecular component of the phage - DNA or protein- entered bacterial cell for viral reproduction.. which one?
-used ratioisotopes ^32P and 35^S to follow the molecular components of phages during infection
--- DNA contains phosphorus not sulfur = 32P label DNA; proteins have sulfur not phosphorus= ^35S labels protein.
-grown in presence of 32P or 35S, then they'll have either radioactive labeled DNA core or radioactive protein coat, respectively.
- centrifugation separates the higher phage particles from heavier bacterial cells
-almost all 32P-labeled DNA had been transferred into bacterial cell following adsorption and 35S stayed outside. then they were lysed and progeny contained only 32P
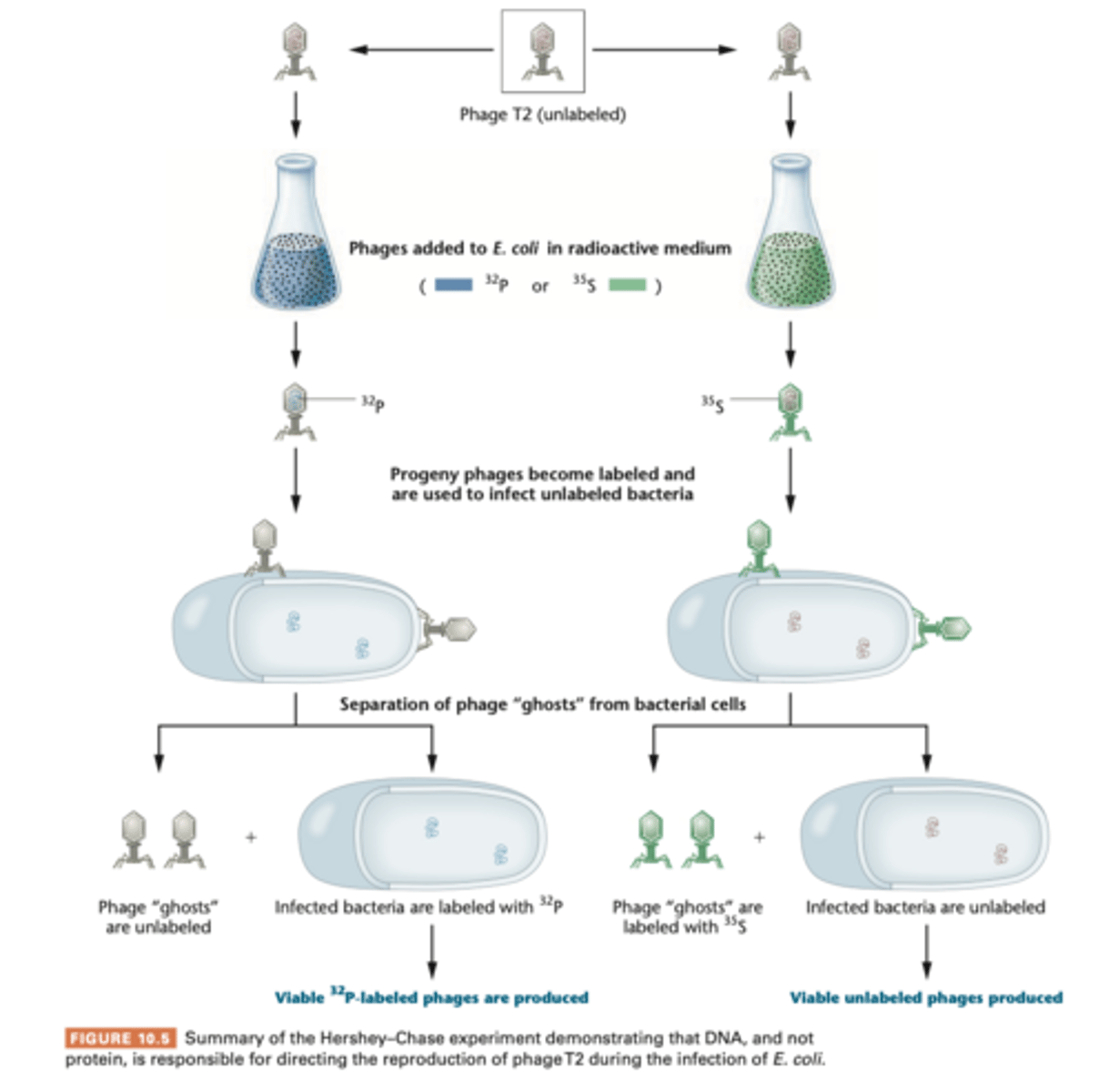
Transfection experiments
protoplasts (or spheroplasts) : e.coli treated with enzyme lysozyme, the outer wall of the cell can be removed without destroying bacteria => they're naked w cell membrane only
-Spizizen and Fraser reported that using protoplasts, initiate phage reproduction with DNA from T2 phages = don't need cell wall wen using protoplasts
- Guthrie and Sinsheimer
-- Purified DNA from bacteriophage o/X174, single stranded circular DNA molecule - resulted in this production = transfection (by only the viral nucleic acid)
Euk indirect evidence - distribution of DNA
- location of DNA v proteins
-- DNA is localized and protein all over the cell
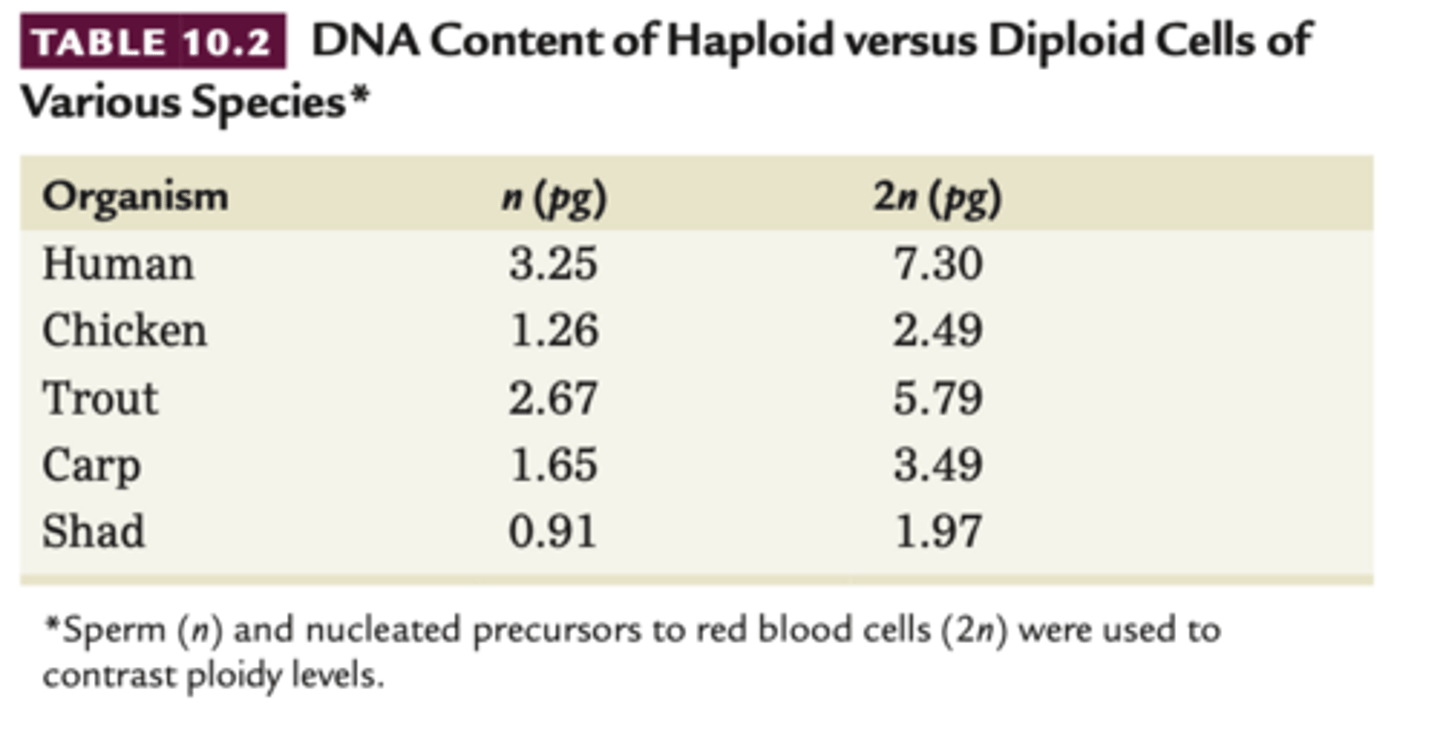
- since know chromosomes carry gen info, looked at correlation of haploid and diploid data -- correlate with DNA not proteins
indirect evidence: mutagenesis
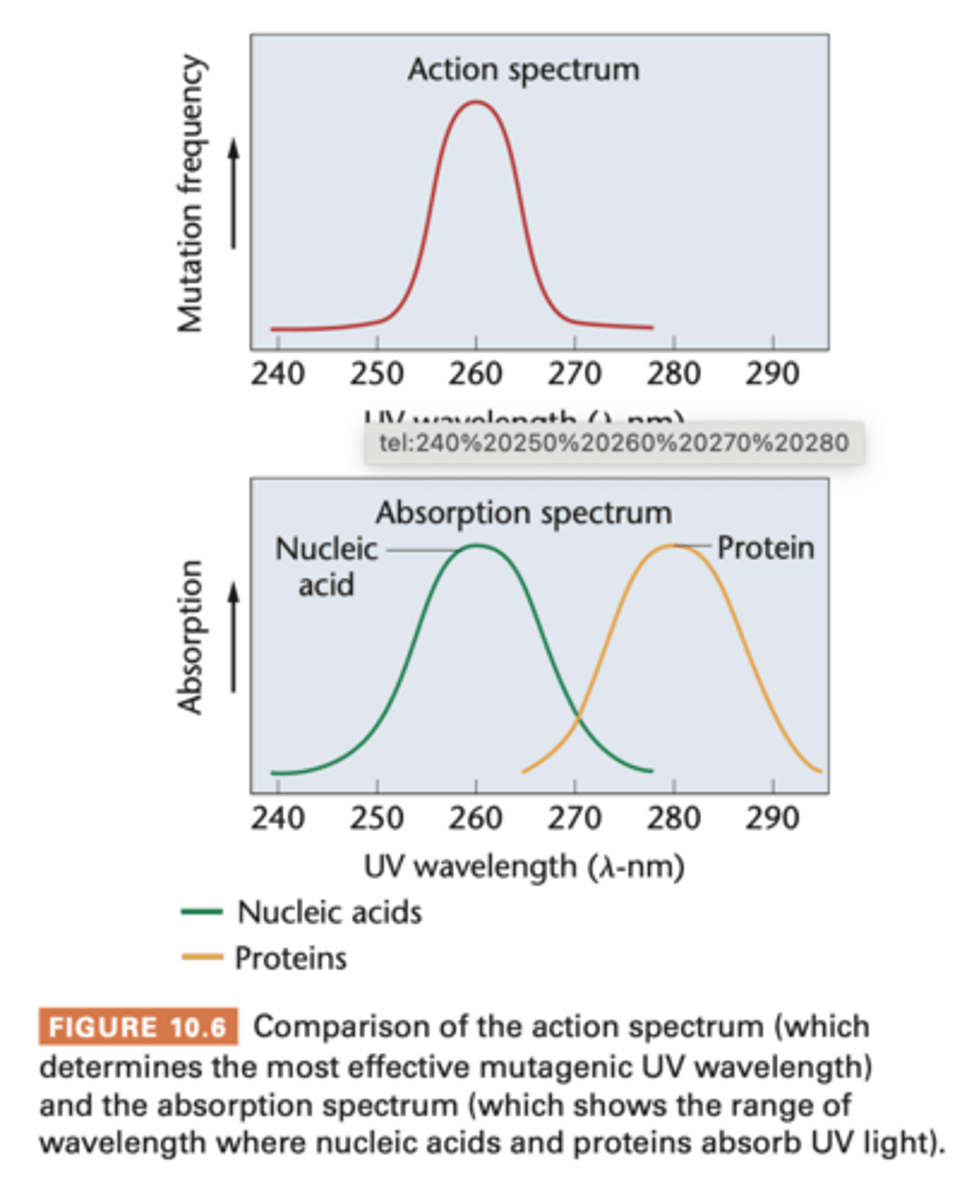
- UV light is capable of inducing mutations in gen material, use yeast and more time IV light = more mutations --> connection btw wavelength and number of mutations = action spectrum
--- of UV light as mutagenic agent is obtained
- absorption spectrum -> compared to absorption spectrum of any molecule suspected to be the genetic material
(molecule serving as the genetic material is expected to absorb at the wavelength(s) found to be mutagenic
- UV light most mutagenic at wavelength 260 nm and both DNA and RNA absorb UV strongly at 260 nm
- protein absorbs 280 nm, no significant mutagenic effects are observed
Direct evidence: recombinant DNA studies
- involved splicing together DNA sequences from different organism, has combined DNA sequence encoding the human hormone insulin w bacterial DNA sequences
- replicate and progeny carry DNA and produce human insulin
DNA= heritable info for genes
genomics
- can provide full set of DNA sequences of organisms, basis of another example
- mid 1970s DNA as gen material was accepted
RNA serves as the genetic material in
some viruses
tobacco mosaic virus (TMV)
1956 it was the purified RNA was spread on tobacco leaves, the characteristic lesions caused viral infection => RNA is the genetic material of this virus
Pace and Spiegelman showed that RNA from the phage QB can be isolated and replicated in vitro
replication depends on RNA replicase, isolated from host e.coli cells following normal infection. When added in vitro, infection occurs
retroviruses
replicate in unusual way
- RNA template for synthesis of complementary DNA
process reverse transcription - under the direction of an RNA-dependent DNA polymerase enzyme called reverse transcriptase
- get incorporated into genome of host cell and DNA trascribed, copies of original viral RNA are produced
HIV & AIDS,
RNA tumor viruses
Nucleotides
- a nitrogenous base
- pentose sugar
- phosphate group
2 nitrogenous bases: 9 member double ring purine (guanine, adenine) and 6 member single ring pyramidines (cytosine, thymine, and uracil)
sugars:
RNA has ribose
DNA has deoxyribose (absence of hydroxyl grp - 2-deoxyribose)
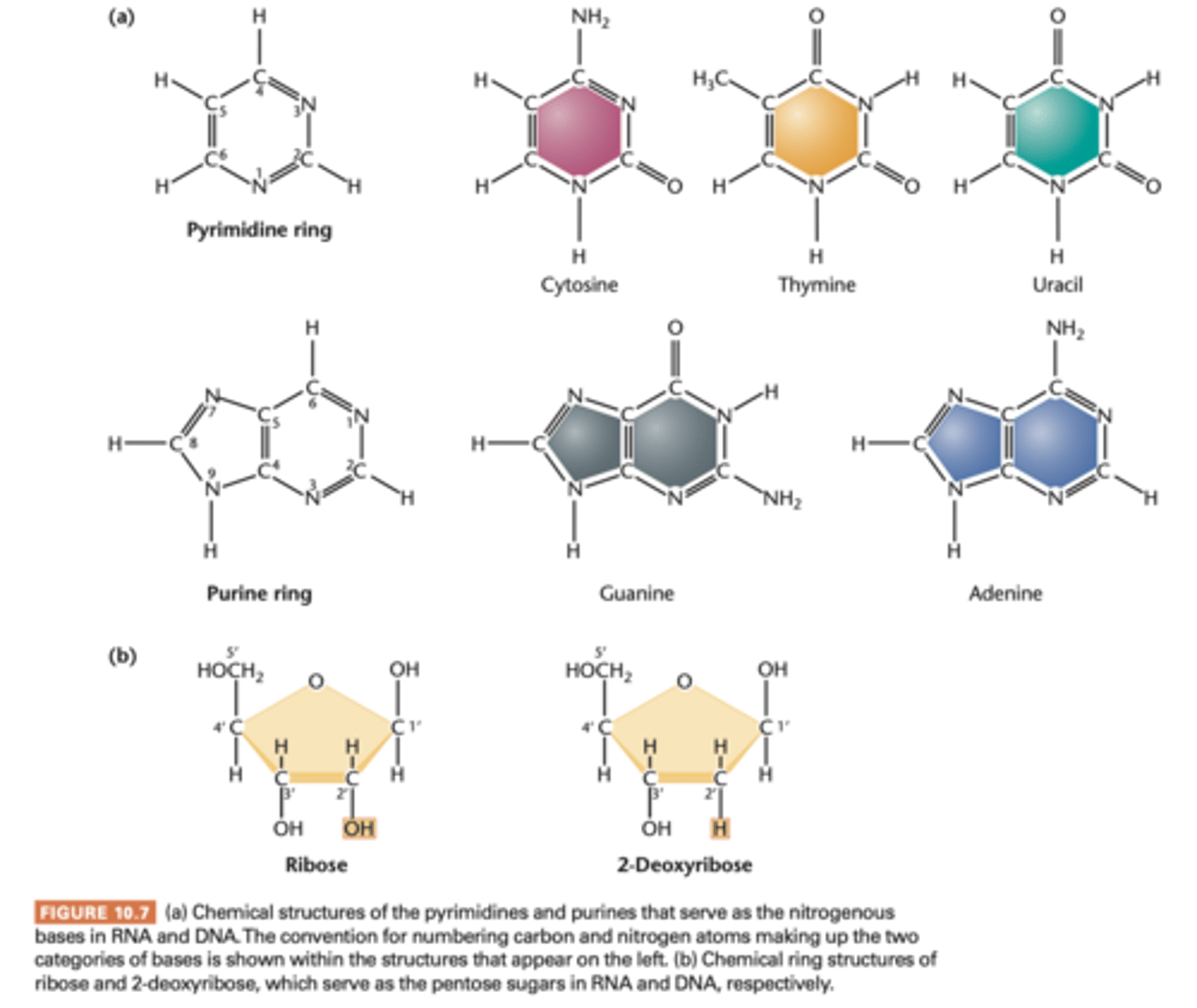
nucleoside
if a molecule is made of a purine or pyrimidine base and a ribose or deoxyribose sugar, chemcial unit is called this.
- if u add a phosphate group, it is called a nucleotide
- named according to nitrogenous bases
purine: N-9 arm is covalently bonded to sugar
pyrimidine: N-1 atom bond to sugar
DNA, phosphate gap may be bonded to C-2', C-3', or C-5' atom of sugarr
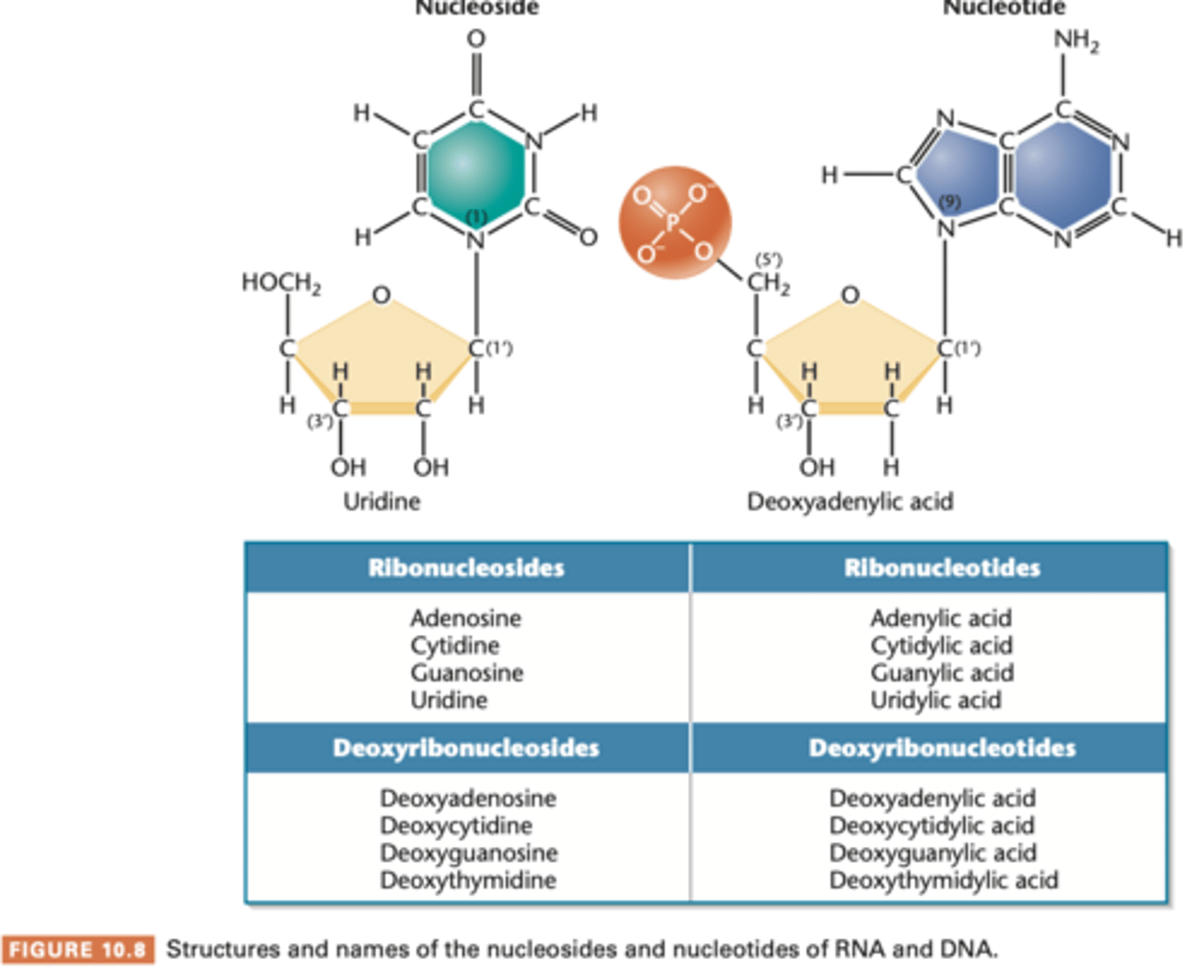
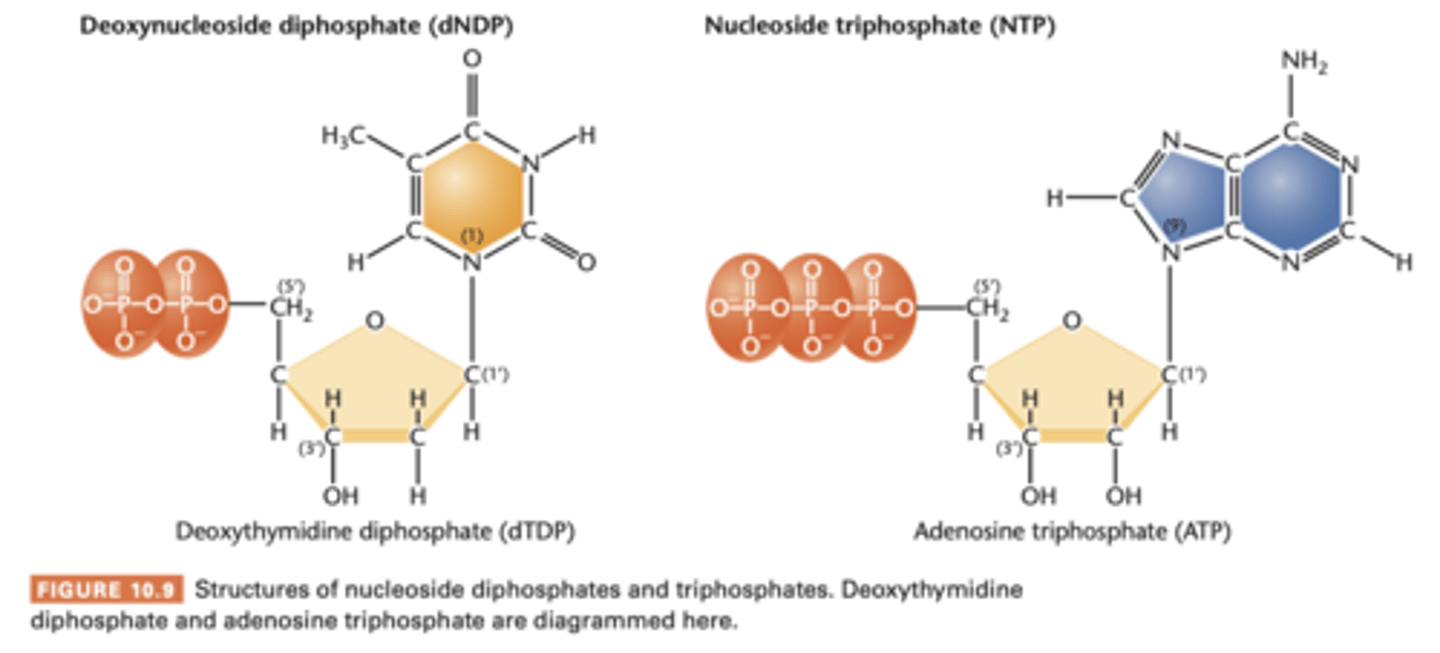
nucleoside diphosphate and triphosphate
nucleosides = nucleoside monophosphate (NMP)
add one or two phosphate groups = nucleoside disphosphates (NDPs) and triphosphate (NTPs) respectively.
NTP is precursor molecule during nuclei acid synthesis within the cell
Adenosine triphosphate (ATP) and guanine triphosphate (GTP)
- cell biogenetics because of the large amount of energy involved in adding or removing the terminal phosphate group
- hydrolysis is accompanied by the release of large amount of energy
- involved in genetic events
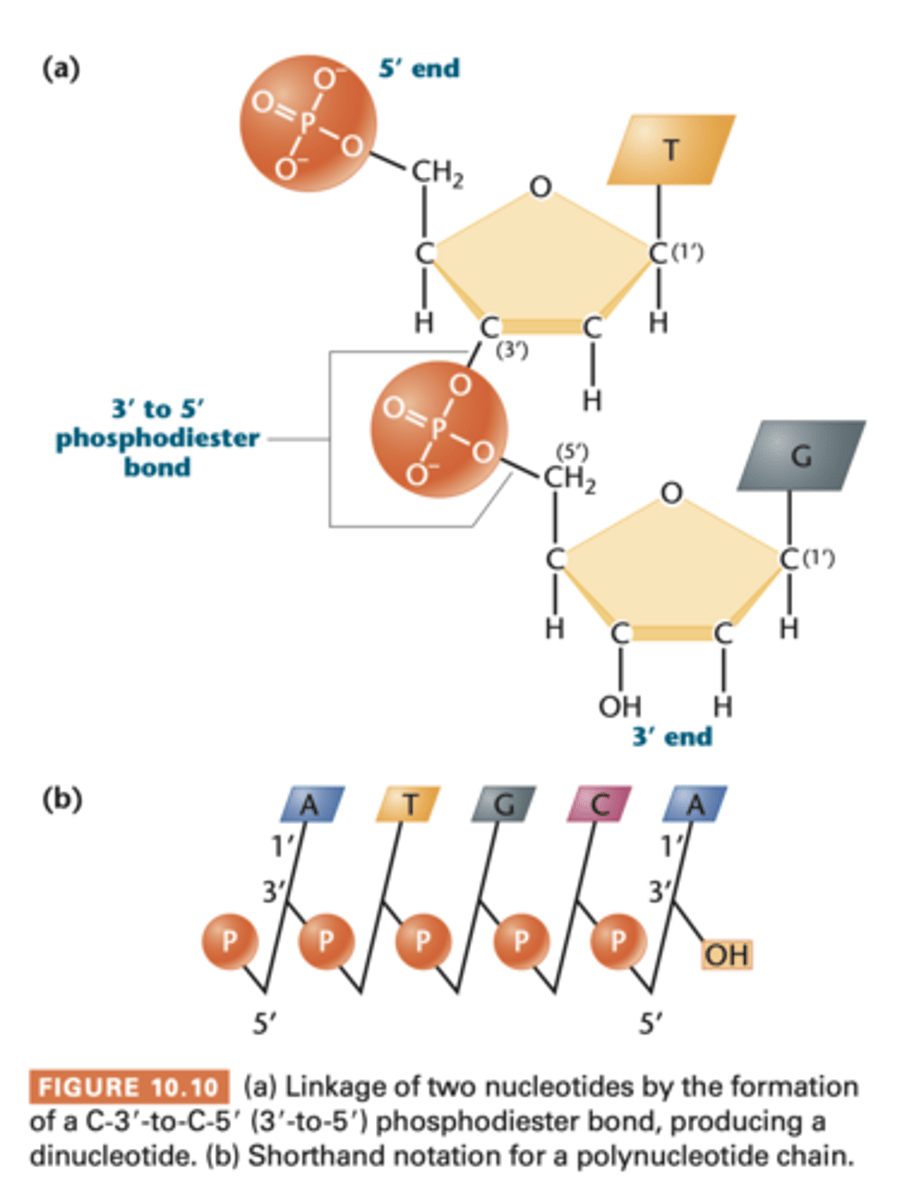
phosphodiester bond t
linkage between 2 mononucleotides consist of a phosphate group linked to 2 sugars
- phosphoric acid has been joined to two alcohols (hydroxyl groups on the 2 sugars) by ester linkage on both sides
- found in RNA too
--> has C-5' end and C-3' end
--> 2 joined nucleotides form dinucleotide; 3=trinucleotide... oligonucleotides (30)... polynucleotides
in image b). vertical lines are pentose sugar, nitrogenous base is at top in C-1'
diagonal line with P in middle it is attached to C-3' position of one sugar and the c-5' of neighboring sugar; represents phosphodiester bond
long polynucleotides chains account for large molecular weight of DNA, explains most important property- storage of vast quantities of genetic info
1000 nucleotides 4^1000 different combos
The structure of DNA holds the key to
understanding its function
Watson and Crick 1953
proposed that the structure of DNA is in the form of double helix. their model was described in a short paper included:
1. base composition analysis and hydrolyzed samples of DNA
2. X-ray diffraction studies of DNA
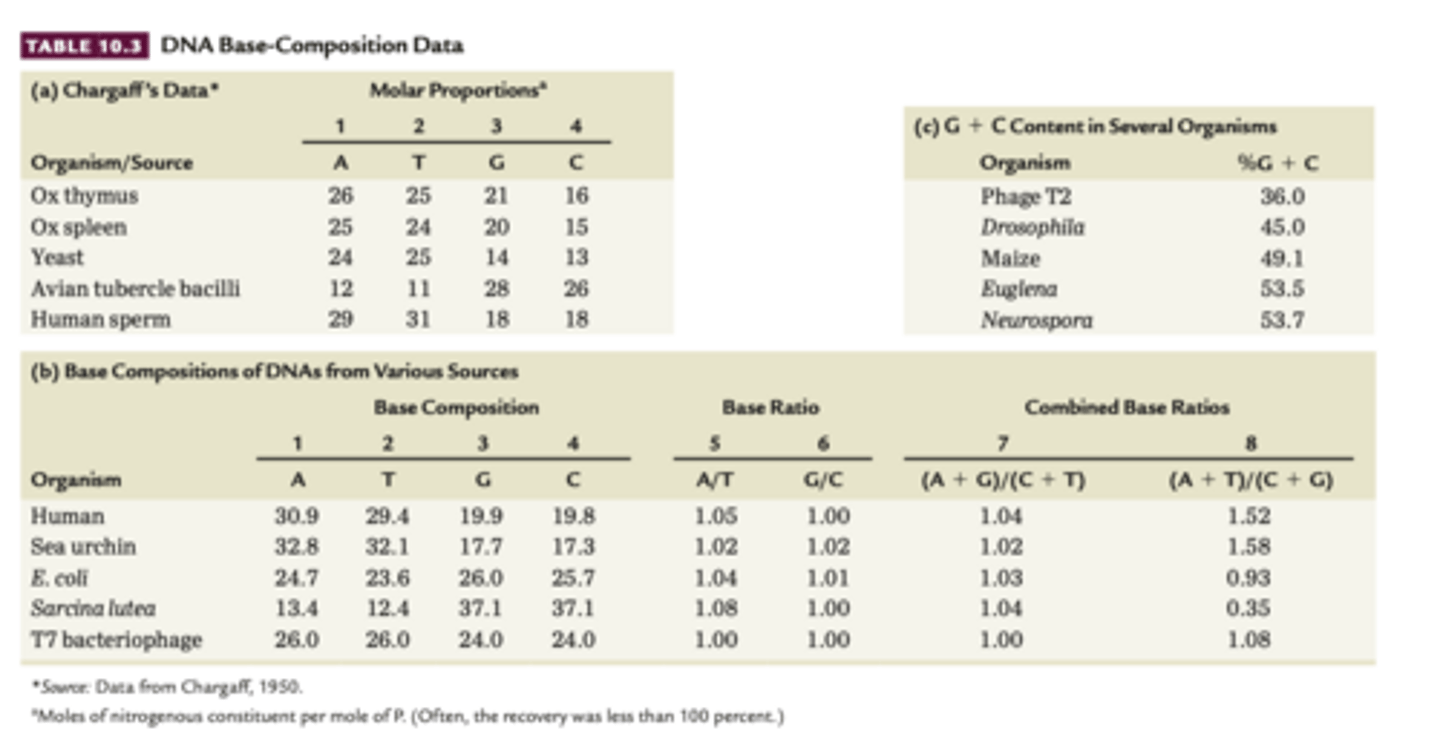
Base-composition studies
1949-1953 Chargaff used chromatographic methods to separate the 4 bases in DNA samples from various organisms
-quantitative methods, on table u can see
1. the amount to adenine residues is proportional to the amount of thymine residues In DNA (1,2,5) and G proportional to C (3,4,6)
2. sum of purines equals the sum of pyrimidines (7).
3. Percentage of (G+C) does not necessarily equal the percentage of(A+T), ratio varies greatly among organisms
- definite pattern of base composition in DNA molecules -- refute tetra nucleotide hypothesis (4 bases are present in equal amounts)
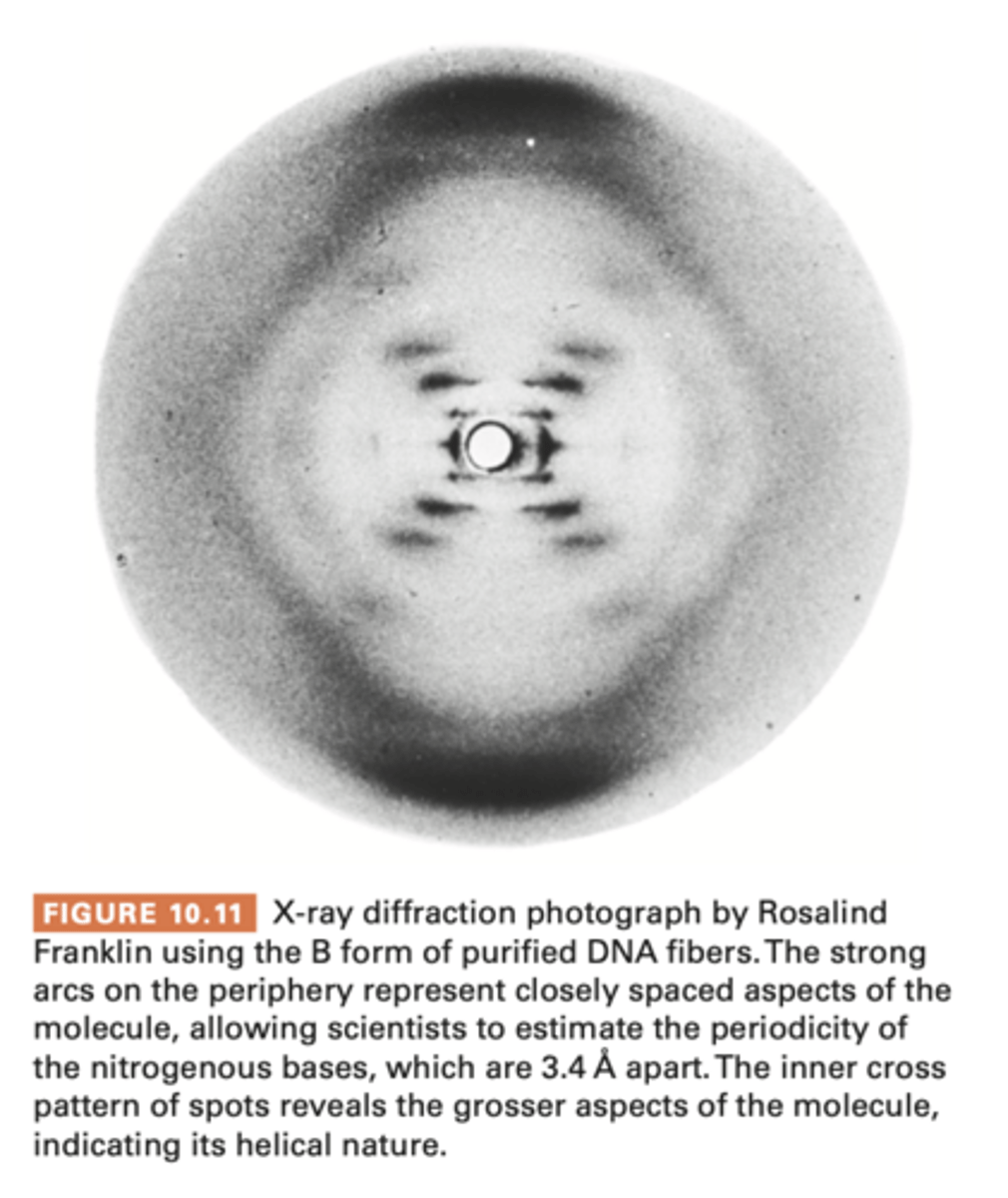
X-ray Diffraction analysis
Fibers of DNA molecule are subjected to X-ray bombardment, x rays scatter in pattern that depends on molecules atomic structure
is captured as spots on photographic film and analyzed for clues to overall shape of regularities within molecule
- used to study protein structure, attempted in DNA
1950-1953- rosalind franklin obtained improved x ray data from purified samples of DNA - suggesting it was a type of helix
did not propose a definitive model - Pauling had analyzed the work of Asbury and others and incorrectly proposed that DNA was a triple helix
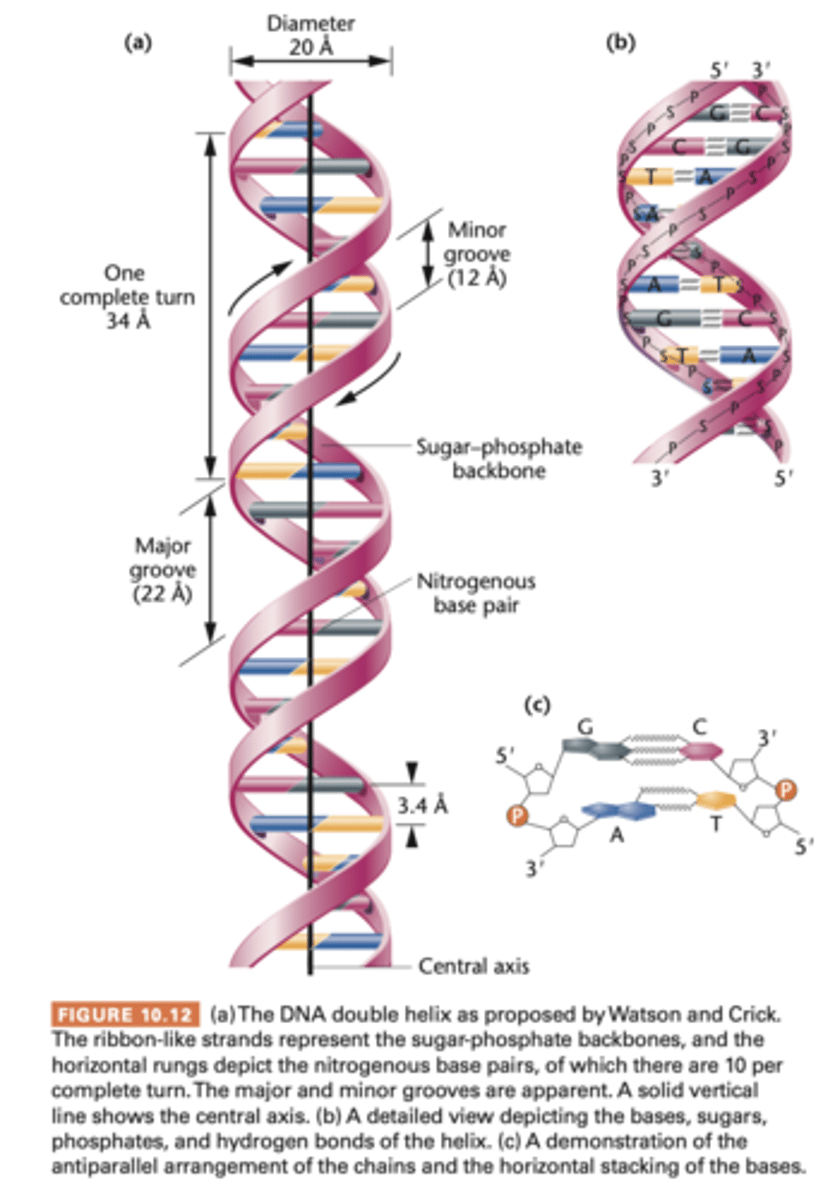
The Watson-Crick model
analysis of DNA structure in 1953, by building models based on above-mentioned parameters, arrive at the double helical form of DNA
following major features:
1. 2 long polynucleotide chains are coiled around a central axis, forming a right handed double helix
2. the 2 chains are antiparallel, their C-5' to C-3' orientations run in opposite directions
3. bases of both chains are flat structures lying perpendicular to axis; are 'stacked' on one another, on inside of double helix
4. the nitrogenous bases of opposite chains are paired as the result of the formation of hydrogen bonds; in DNA only A-T and C-G pairs occur
5. Each complete turn of helix is 3.4nm long; thus, each turn of the helix is the length of a series of 10 base pairs
6. a larger major groove alternating with a smaller minor groove winds along the elnegth of the molecule
7. the double helix has a diameter of 2.0nm
AT CG complementarity, results from chemical affinity that makes H bonds
-nitrogenous hydrophobic on inside
- sugar phosphate backbones on outside hydrophilic
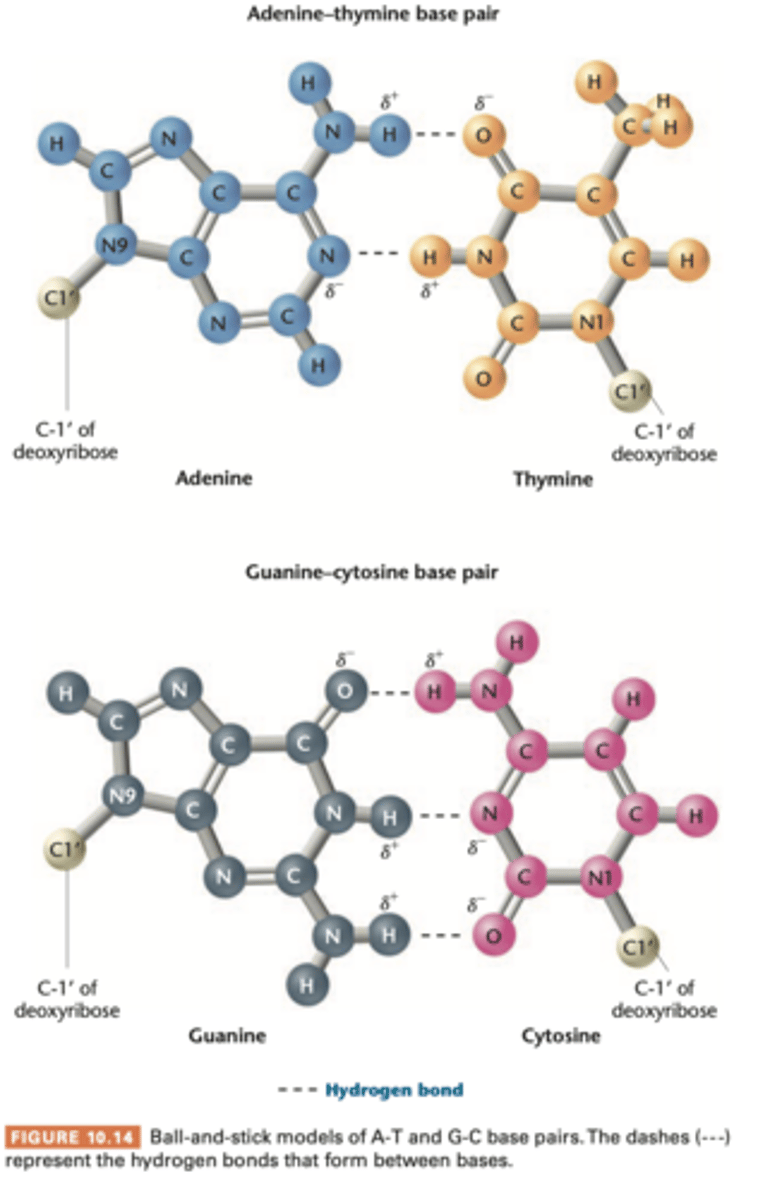
hydrogen bond
very weak electrostatic attraction between a covalently bonded hydrogen atom and an atom with an unshared electron pair
assumed a partial positive charge while the unshared electron pair assumes a partial negative charge
opposite charges are responsible for the weak chemical attraction that is the basis of h bonds
a and t have 2
g and c have 3
When Watson and Crick performed analysis, they had to forms of DNA
A-DNA and B-DNA
based on RF studies on the B form, under aqueous, low salt conditions and believed to be the biologically significant conformation
single crystal x ray analysis
diffract x rays at about 1A intervals (v 5) therefore every atom is visible and greater in structural detail
saw:
A-DNA is more compact, 9 bp in each complete turn of the helix, 23A in diameter (compared to B-DNA), right-handed (also), bases are tilted and placed laterally in relation to axis of helix therefore different appearance of major and minor grooves, thus doubtful A-DNA happens in vivo
C-DNA found under greater dehydration conditions than those observed in isolation of A and B DNA
D and E DNA occur in helices lacking G
P DNA, if DNA is artificially stretched,
Z-DNA left handed helix, think regions exist in chromosomes of living organisms (still unknown)
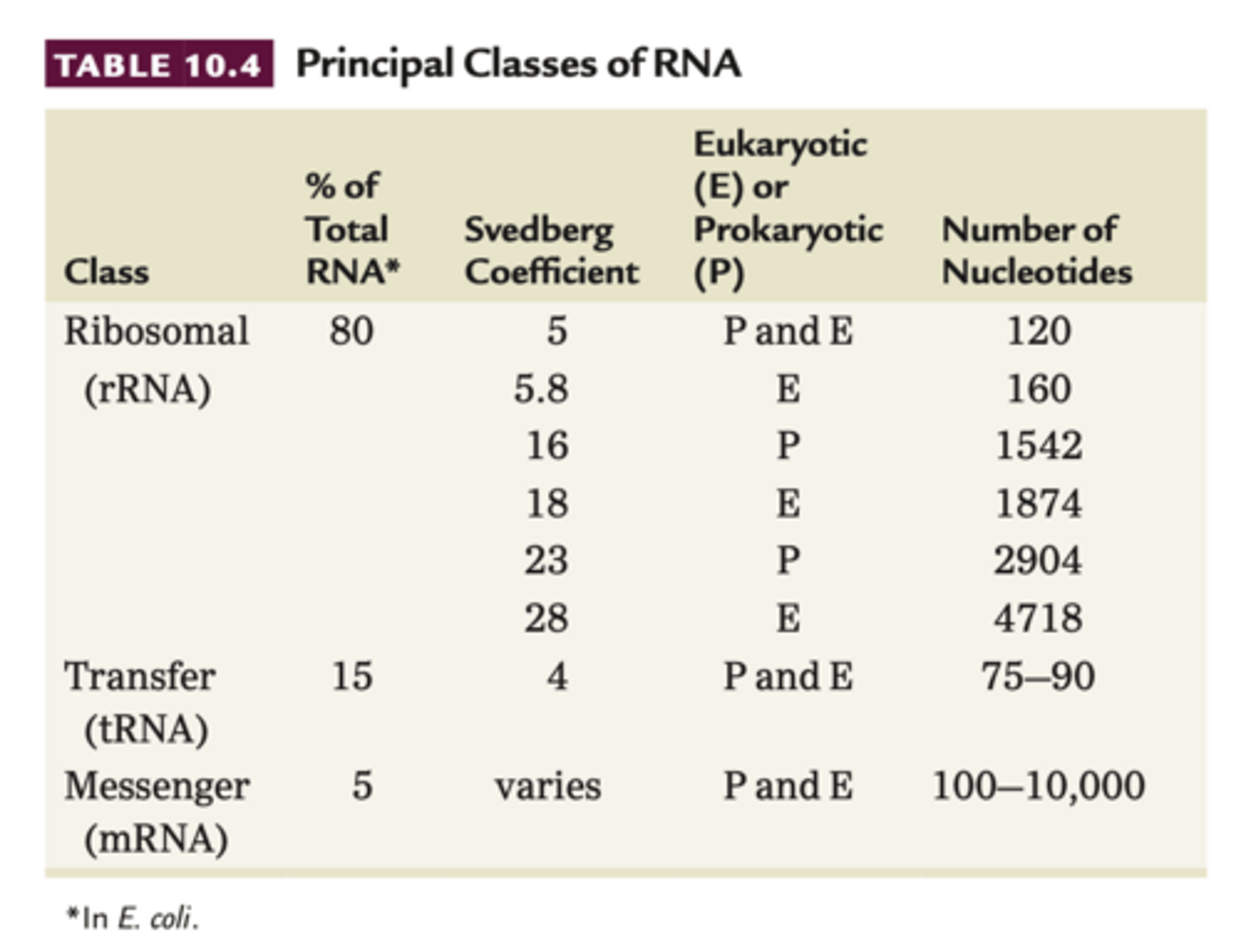
RNA
RNA can fold back on itself to make double stranded regions of complementary bp
- some animal viruses w RNA have double stranded helices (can regulate gene expression in eukaryotic)
3 major RNA
- rRNA
-mRNA
- tRNA
Svedberg coefficient (S)
- sedimentation behavior depends on molecules density, mass and shape, called this
- high value = high molecular weight
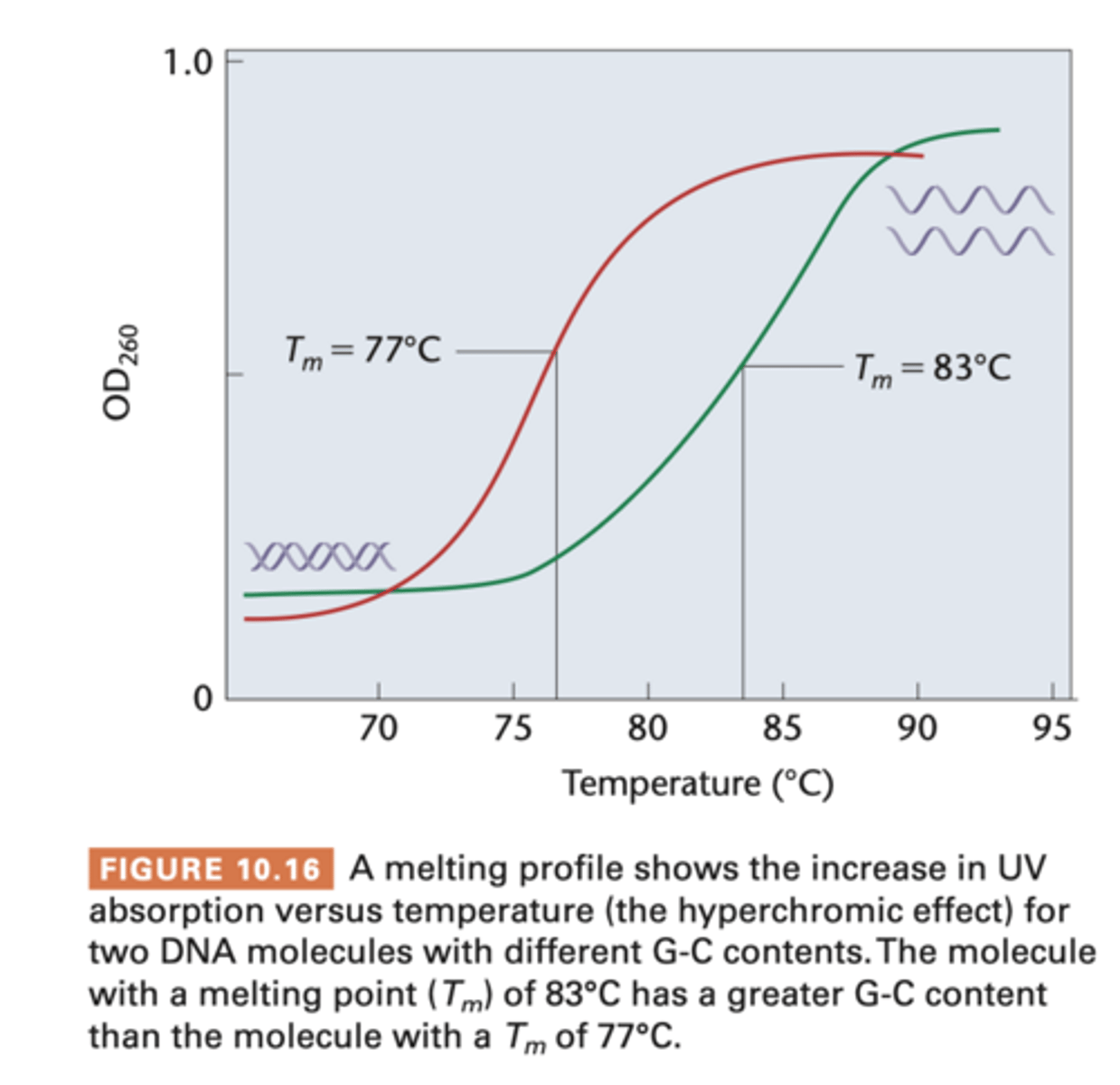
If DNA is subjected to heat (ie hyperchromic shift)
double helix is denatured and unwinds
- during unwinding, viscosity of DNA decreases and UV absorption increases (AKA hyperchromic shift)
- on graph, midpoint of each curve is called melting temperature (Tm) where 50% strands are unwound - higher Tm has higher % GC bp than AT, since CG have 3H vs 2H
molecular hybridization
denaturation/renaturation
duplexes can be made btw DNA strands, even from different organisms and btw DNA and RNA
probes are used to ID complementary sequences
in situ molecular hybridization
- use DNA in chromomal prep as target for hybrid formation
- mitotic cells are fixed to slides then hybridization conditions
- single strand DNA/RNA (probe) added and hybridization monitored
- added pt can be radioactive or have fluorescent label
w fluorescence = FISH
- to identify chromosomal locations housing specific genetic info, valuable addition to techniques
electrophoresis
can separate different sized fragments of DNA and RNA chains
- DNA migrate under influence of electric field
- on semisolid gel, in solution that conducts electricity (agarose gel)