lala added ch 16
1/150
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
151 Terms
Negative Numbers ( -100, -1000)
Positions Upstream of the start site
diffeential gene regulation
Cells of different tissues express different sets of genes
Why does transcription initiation in eukaryotes require numberous proteins and allosteric changes?
Assembly of the transcription complex requires multiple proteins (activators, coactivators, RNA Pol II, GTFs) and allosteric changes that allow them to properly bind each other and DNA, ensuring efficient transcription initiation
transcription initiation complex
exists to assemble proteins at the promoter to position RNA polymerase and unwind DNA
Enhancer
DNA sequence that increases transcription when bound by activators.
Where are enhancers found?
Upstream, downstream, or within introns-- or far from gene
"Action at a Distance"
Enhancers affect transcription from far away via DNA looping
"Upstream"
Region before the transcription start site.
"Downstream"
Region after the transcription start site.
+1
transcription start site
Cell Differentiation
Cells becoming specialized for structure & function; occurs during embryonic development and can continue into adulthood
positive feedback loop
Some Transient Signal turns on expression of protein A, making transcription factor activate its own gene. Once its on it continues to be produces even after initial signal is gone. This memory allows cell to remember its identity and maintain differentiated state over time
stem cell
unspecialized cell that can develop into a specialized cell under the right conditions
RNA Polymerase
transcription enzyme
All genes...
have a series of enhancer sequences with specific regulatory proteins
Similarities of prokaryotic and eukaryotic gene regulation
Both regulate gene expression with DNA-Binding proteins & respond to environmental signals
Prokayotic gene regulation
operons, transcription and translation are couples, simpler regulation
Eukaryotic gene regulation
Complex regulation, changes in chromatin structure, transcription occurs in nucleus, many levels of control
RNA Pol I
transcribes rRNA
RNA Poly II
transcribes mRNA and some snRNA
RNA Poly III
transcribes tRNA and 55 rRNa
general transcription factors (GTFs)
Correctly position polymerase at/around start site and help intiate transcription. analogous to Bacteria Sigma Factor
TFIID/TBP
bind to TATA box using TBP, bending DNA and allowing other proteins access
TFIIH
releases Pol II by phosphorylation of the C-Terminal tail of RNA Pol II. Leads to start of transcription
How do activators stimulate transcription through direct association
Activators directly bind RNA Pol II/GTFs to increase transcription
How do repressors inhibit transcroption through direct association
By binding/interfering with transcription machinery
How do activators and repressors regulate transcription by chromatin modification
Activators: reruit chromatin-modifying enzymes to open DNA
Repressors: Promote Chromatin condensation, making DNA less accessible to transcription machinery
How do HATs affect chromatin structure
add acetyl groups to histones that loosen the chromatin --> increasing transcription
How do HDACs affect chromatin structure?
remove acetyl groups, tightening chromatin --> decreasing transcription
How does the glucocorticoid receptor regulate genes
A ligand-activated transcroption factor that binds to glucocortid hormones, enters nucleus, and regulates gene ecpression by binding specific DNA sequences
Why is a skin cell different from a muscle cell?
They're diff because they express different genes, allowing them to produce different proteins and giving each cell a unique structure & function
If DNA is a molecule with Partial and Full charges
then it can form non-covalent bonds w/ proteins with polar and charged amino acids
What is the significance of the phosphorylated tail of RNA Pol II
- start of elongation
- recruits RNA processing proteins (capping, splicing, polyadenylations)
- Coordinates initiation --> elongation transition, integrating transcription with RNA maturation
How do cells control gene expression in a temporal manner?
By controlling the expression of regulatory factors allowing genes to be ecpressed only when needed
Still learning (15)
You've started learning these terms. Keep it up!
Nuclear Import
1. Protein contains a nuclear
localization signal (NLS) (basic)
2. Binds to a specific receptor protein
(importin) via the NLS
3. Importin associates with nuclear
pore
4. GTP-mediated import
5. Complex released inside nucleus
Nuclear Export
1. Protein contains nuclear export signal
(NES) (leucine-rich)
2. Protein binds to specific receptor
protein (CRM-1) via the NES
3. CRM-1 associates with nuclear pore
4. GTP-mediated export
5. Complex is released outside nucleus
6. Protein may be associated with RNA.
What are sorting signals?
Amino acid sequence that specifies where a protein is sent.
What are sorting signals made of?
Specific, short amino acid sequences (5-60 residues) within proteins that act as molecular "zip codes," determining their proper destination within or outside the cells
What happens when a signal sequence is swapped btwn proteins that function in a different subcellular compartment?
the protein will be trafficked to the wrong location. Proteins cannot function in the wrong environment --> leads to degradation or incorrect cellular function
What do sorting signals do? Where are they found?
Sorting signals are at the N-Terminus, hydrophobic and part of the primary structure of the protein. Used to sort proteins, insert protein into organelle/membrane
Where do chloroplast and mitochondrial proteins originate from?
Nuclear Genome & Organelle's Genome. endosymbiosis
Where are ribsosomes made?
Nucleolus
What are free ribosomes
Riobosomes in cytosol making proteins for cytosol, nucleus, mitochondria, peroxisomes
What are bound ribosomes?
Riobosomes attached to rough ER making secreted, membrane, or lyossomal proteins
Do ribosomes start as free or bound?
Free
What makes a ribosome become bound?
Signal sequence and SRP (signal recognition particle) targeting to ER
What happens to ribosomes after translation?
They detach and become free again
Difference bewtween free and bound ribosomes?
Depends on protein being made
The N-Terminus is
nonpolar, facing the cytosol
Proteins are sorted to locations beyond the ER within
transport vesicles
Secetory Pathway (exocytosis
-for export
- deliveries lipid/protein to plasma membrane
Endocytic Pathway
- for import
How do vesicles form?
Budding driven by assembly of a protein coat
How is vesicle formation controlled?
Different membranes have their own protein coats
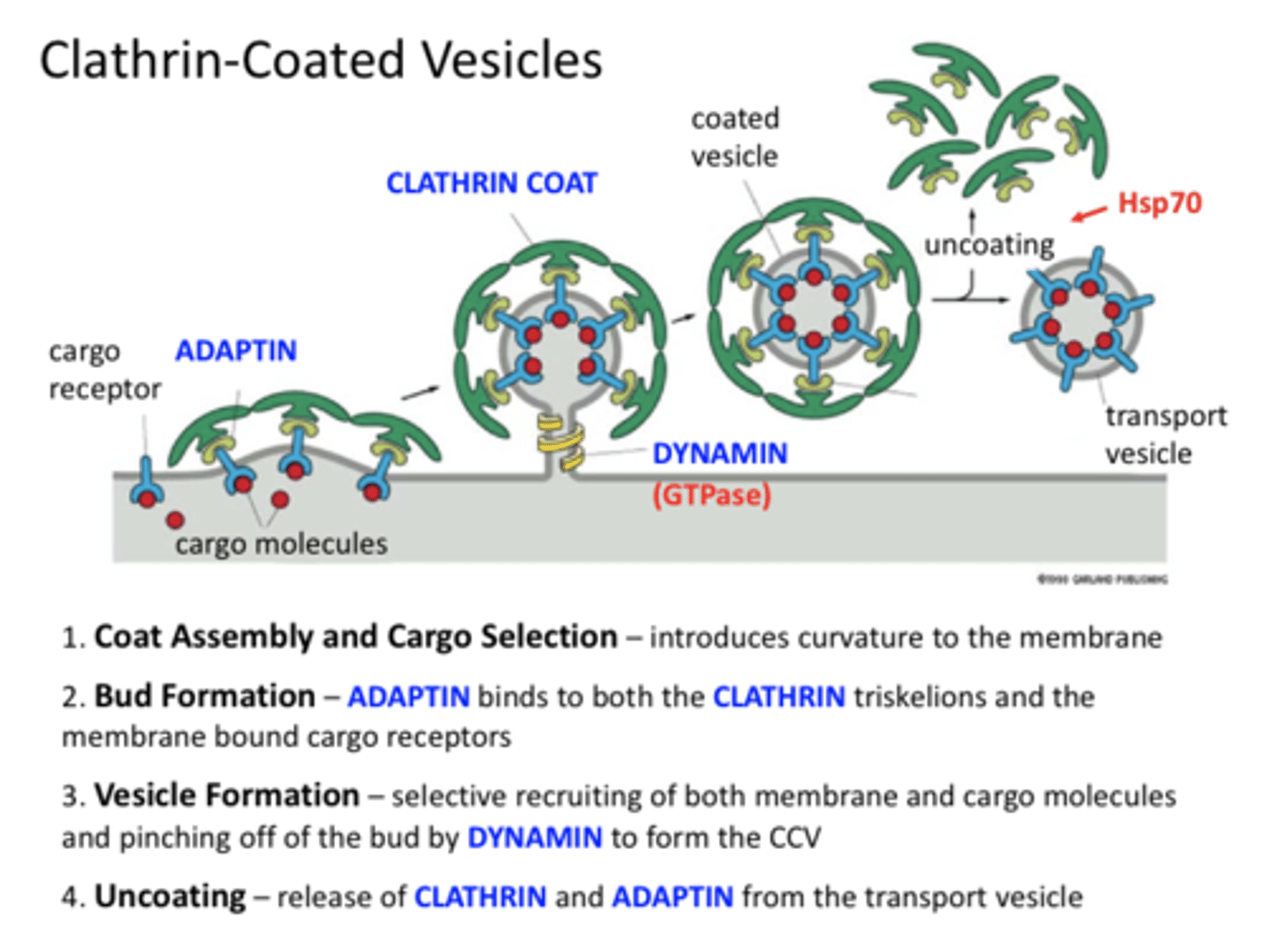
Clathrin
protein that coats the plasma membrane's inward-facing surface and assists in forming specialized structures, like coated pits, for phagocytosis
coated pit becomes
a coated vesicle
Adaptin
binds clathrin and cargo receptors (an adaptor protein)
Endocytosis at Plasma Membrane
1. Binding of cargo to cargo receptors
2. Lateral movement into a coated pit
3. Binding of adaptin to receptor
4. Binding of CLATHRIN to adaptin
5. BUDDING of coated pit
6. Vesicle is pinched off by dynamin
(GTP)
7. Uncoating
8. Onward to target
Enables endocytosis to occur
What needs to happen in budding/fusion?
• Each vesicle must take ONLY those
proteins destined for target.
• Each vesicle must fuse with
CORRECT membrane.
• Lipid MAKE-UP of target must be
conserved.
• AMOUNT of lipid in target must be
conserved.
The cytosolic face
remains facing the cytosol
How does a vesicle know where to go?
Specific protein in its membrane (aka v-Snare)
Target membrane is the receptor (T-Snare)
How does a vesicle reach its destination?
Uses micotubules as tracks w/ motor proteins using ATP hydrolysis (conformational change)
oligasaccharides
short chain of sugar residues/ often lipid linked to protein
Glycosylation
enable cell to cell communication using sugars on the cell suface
Protein folding in the cell is facilitated by
chaperon proteins
Chaperon proteins
- retain misfolded proteins in ER then degrades in a lysosome
Proteins move through golgi via
vesicles
cisternae
flattened stacked membrane folds that make up the golgi apparatus
Cisface
receiving side of golgi apparatus; recieves vesicles
Transface
shipping side of golgi apparatus; buds & sends off vesicles
Golgi cisternae...
further modify proteins by adding or removing sugars to the oligosaccharides
Constitutive pathway (think constant)
- operates continually
- supplies lipids, membrane proteins, and soluble proteins to cell membrane
Regulated pathway-- when the time is right..
- operates in SPECIALIZED SECRETORY CELLS (e.g glands)
-secretory proteins are stored in specialized secretory vesicles (hormones, digestive enzymes, etc)
pinocytosis
- cellular drinking
- generalized
Endocytic pathway
form plasma membrane to a lysosome
-> pinocytosis
->phagocytosis
Phagocytosis
- "cell eating"
- large
-specific cells do dis
Endocytosis
continual intake of vesicles and recycling of surface receptors
-> non receptor mediated
-> receptor-mediated (e.g protein coating)
Endosome
destination of vesicles
function of the endosome
- Allows cargo to come apart from receptor originally bound to
- receptor is recycled and sorted back to cell surface
-control for endocytosis
why does cargo come off receptor?
The endosome is slightly acidic.
One function of the golgi
- control for exocytosis
High cholesterol is from
a mutation in LDL receptor
Receptors can be
1. recycled to the same memrbane
2. degraded in the lysosome
3. moved to another suface with ligand
Lysosomes
Sites of intracellular digestion of extracellular compounds
How do enzymes get to the lysosome?
theyre made by ribosomes on the ER surface, sorted to the golgi, then the endosome, then finally the lysosome
Why do enzymes in the lysosome not destroy the ER
The environment of the lysosome has a pH of 5, snapping the acidic hydrolytic enzymes into an active conformation —> enzymes are active at a low pH but a lysosome is acidic.
Why is the interior of the lysosome acidic
proton pumps in the membrane of the lysosome; it is an ATP driven pump
What is a phospholipid
It is a lipid that has a phosphate group attached to the glycerol and only two fatty acid chains.
What is the phospholipid head composed of?
Glycerol, phosphate, and a chemical group (choline, ethanolamine, serine and inositol). hydrophobic and hydrophilic regions make phospholipids amphipathic
What keeps the closed compartments made of phospholipids
Phospholipids naturally form bilayers and sealed compartments in water in ball formations. Thermodynamically this is the most favorable conformation of phospholipids
What part(s) of the membrane is fluid?
The phospholipid bilayer
What motion occurs in the membrane?
-Lateral diffusion
-rotation
-flip-flopping (rare, fixed by flippases)
Why is the membrane fluid?
Fatty-acid tails move laterally, where fluidity originates
How does the saturation-level of fatty acid tails affect membrane fluidity
"Saturated" --> max # of Hydrogens, no kinks in tail. Better packing therefore the lipid is less fluid
"Unsaturated" --> less than the max # of H which causes kinks due to double bonding; kinks mean less packing which means lipid is more fluid
How does the presence of cholesterol affect membrane fluidity?
High temps --> reduces fluidity by restricting phospholipid movement
Low temps --> increases fluidity by preventing tight packing & freezing.
What is cholesterol? Where is it located?
Cholesterol is embedded in the membrane (20% of lipid) and decreases permeability by restricting lipid movement which prevents crystallization
Located in the top half of phospholipid tails
Why is fluidity important?
1) membrane permeability
2) protein functioning (e.g protein-protein interactions)
3) redistribution of lipids/proteins (i.e synthesis, fusion, cell division)
How are membrane proteins associated with membranes?
- Transmembrane (integral)
- Membrane associated (integral)
- Lipid linked (integral)
- Protein-protein (peripheral)