BioChem Exam 4
1/220
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
221 Terms
what is the general central dogma of biology? is it accurate?
DNA →RNA → Protein via transcription and translation
not entirely accurate, there are some exceptions - still a useful principle but not really “central dogma” anymore
what are the basic descriptors of a strand of DNA?
made by two complementary strands, the two strands are anti-parallel, and it has a right-handed helix structure
the backbone of a strand of DNA is made of _______ and ________
sugar and phosphate
the two strands of DNA interact with eachother via __________
H-bonds
a base + a ribose sugar = ___________
nucleoside
a base + a deoxyribose sugar = _______
deoxynucleoside
a nucleoside is a ______ and a ________
base and a (deoxy) ribose sugar
name the four RNA nucleosides
adenosine, guanosine, cytidine, uridine
name the four DNA nucleosides
deoxyadenosine, deoxyguanosine, deoxycytidine, (deoxy)thymidine
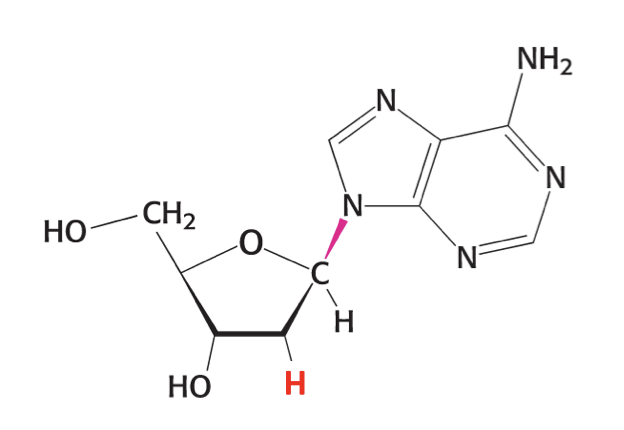
what is this
deoxyadenosine
in purines, which nitrogen is attached to C_ of the sugar. what about pyrimidines?
N9 for purines, N1 for pyrimidines, attach to C1’ of sugar
the bond between the N1-C1’ in pyrimidines and the N9-C1’ in purines is known as ________
β-glycosidic bond (linkage)
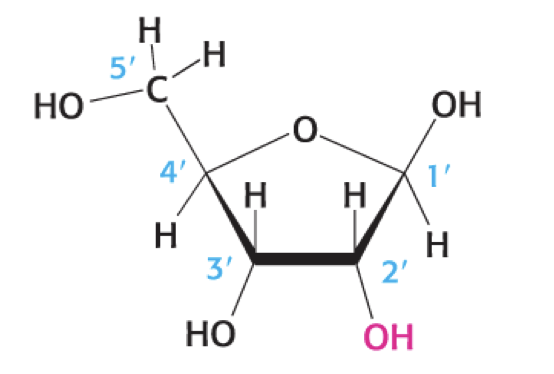
what is this
ribose sugar
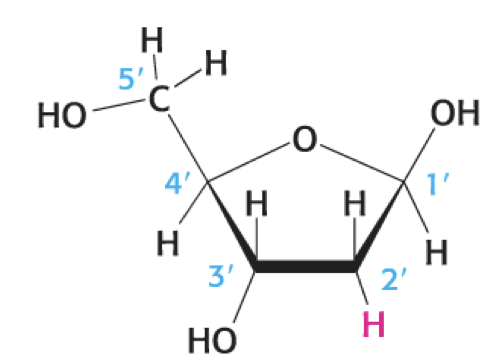
what is this
deoxyribose sugar
the sugars in both RNA and DNA are called _______ (general term)
pentose sugars
how are the carbons on a pentose sugar labeled?
1’ - 5’, starting with the carbon bonded to the OH group that is on the ring
what is the difference between a ribose sugar and a deoxyribose sugar
ribose has an OH at the 2’C, while deoxyribose has an H
what is a nucleotide?
nucleoside + phosphate
a nucleotide is a _____ + _______
nucleoside + phosphate(s)
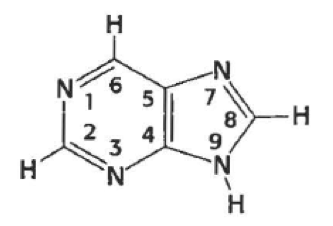
what is this (generally)
purine
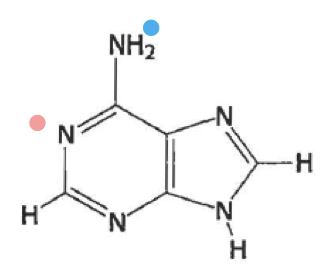
what is this
adenine
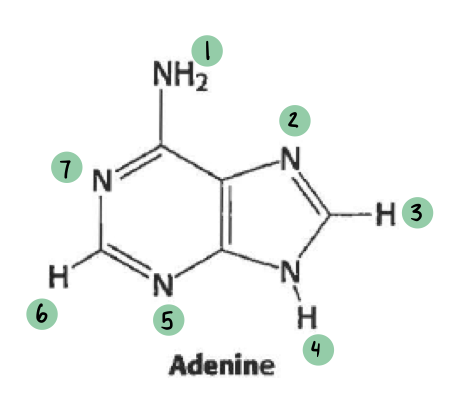
which of these have a 𝛿- and which have a 𝛿+ (not all will have a charge)
𝛿- ~ 7
𝛿+ ~ 1
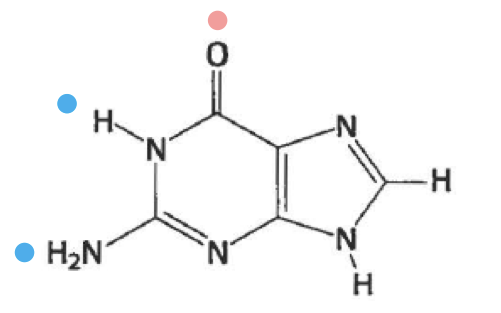
what is this
guanine
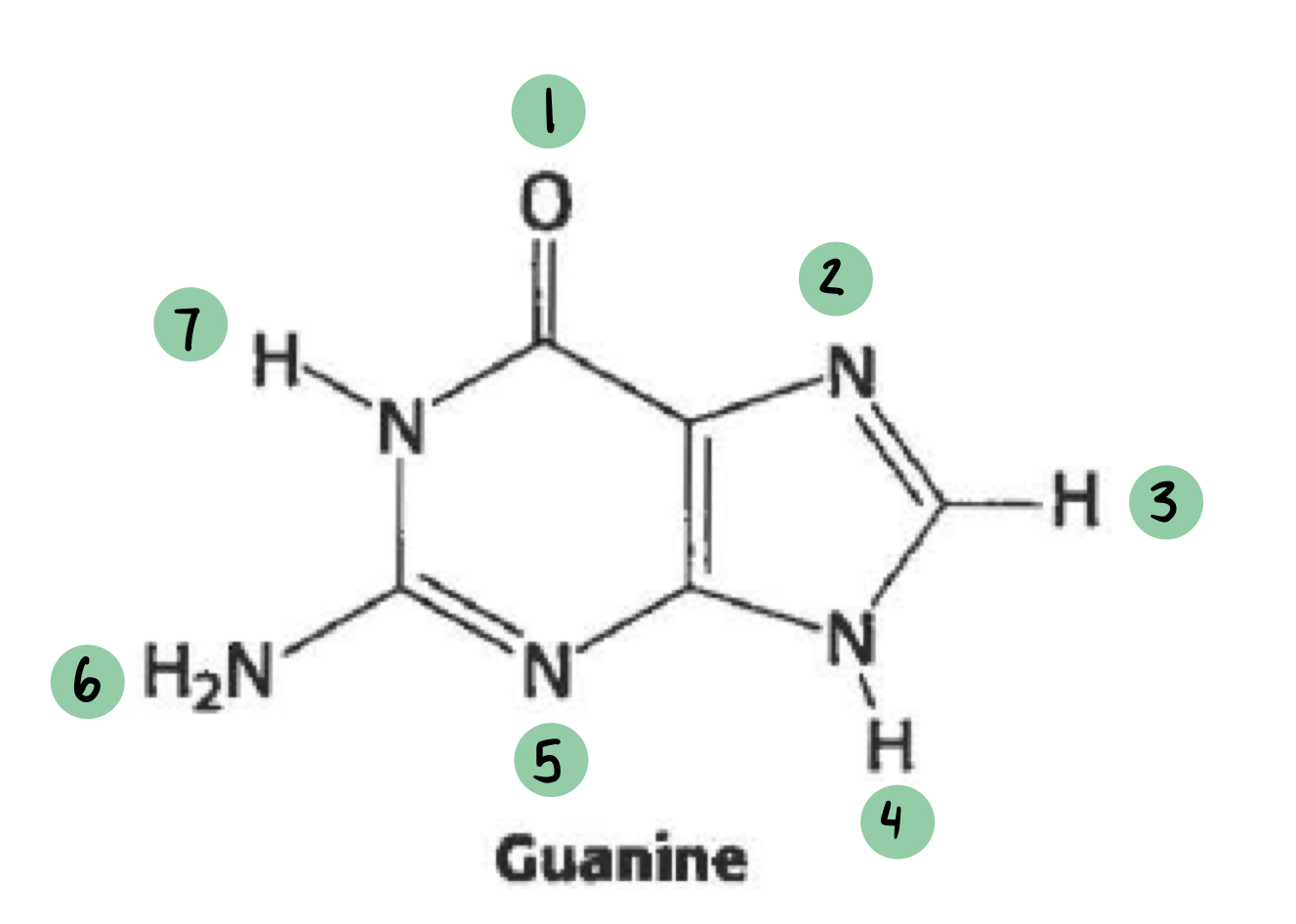
which of these have a 𝛿- and which have a 𝛿+ (not all will have a charge)
𝛿- ~ 1
𝛿+ ~ 6, 7
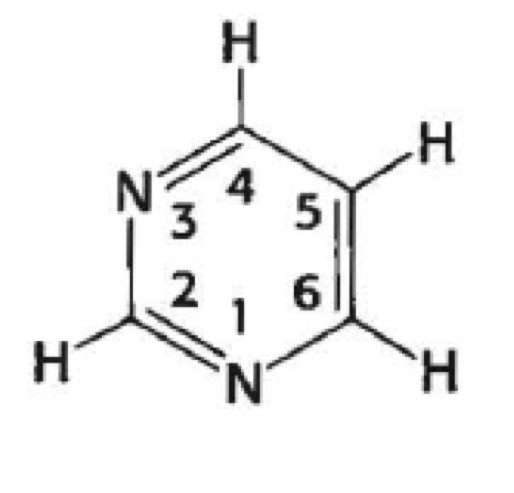
what is this (generally)
pyrimidine
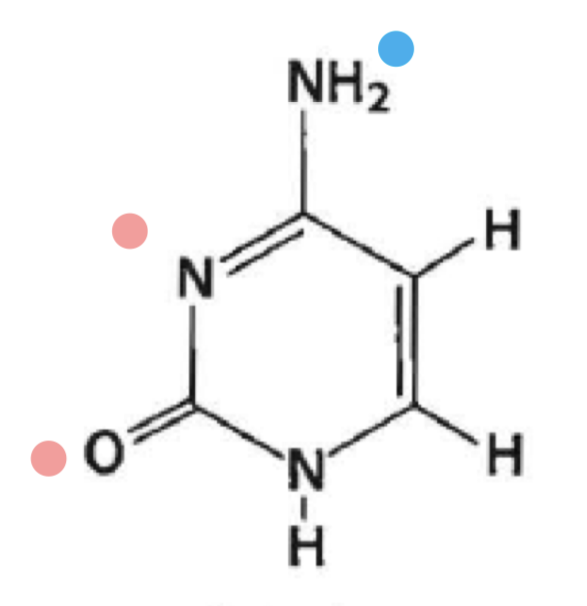
what is this?
cytosine
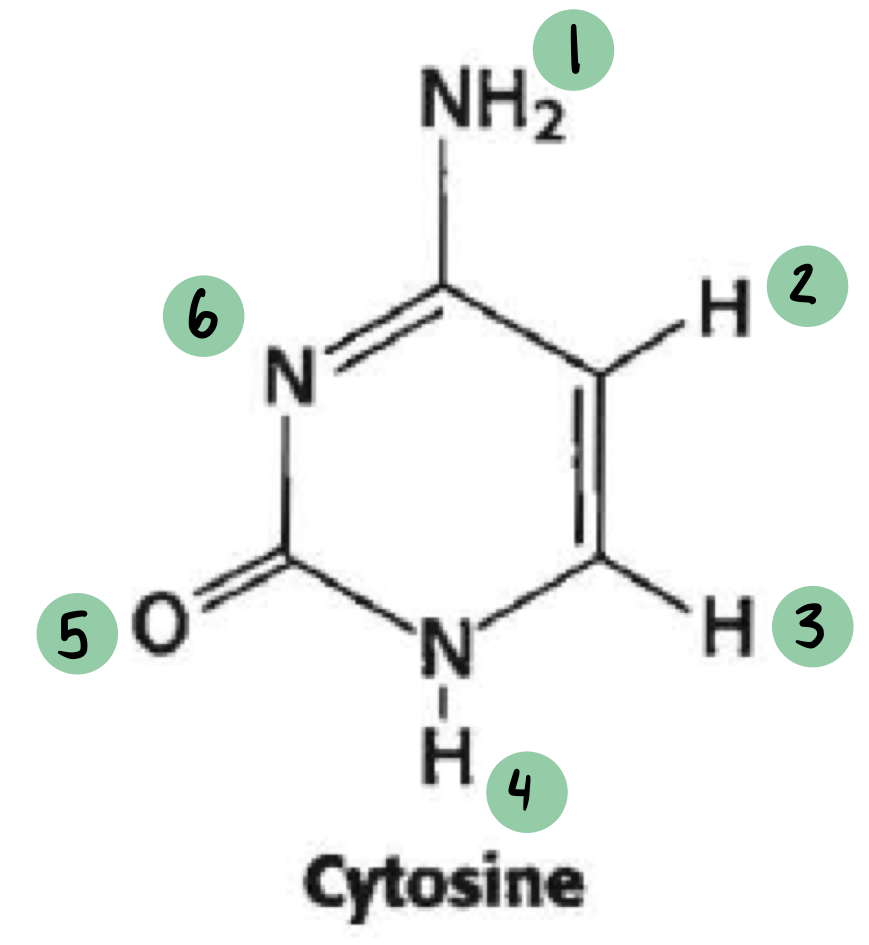
which of these have a 𝛿- and which have a 𝛿+ (not all will have a charge)
𝛿- ~ 1
𝛿+ ~ 5, 6
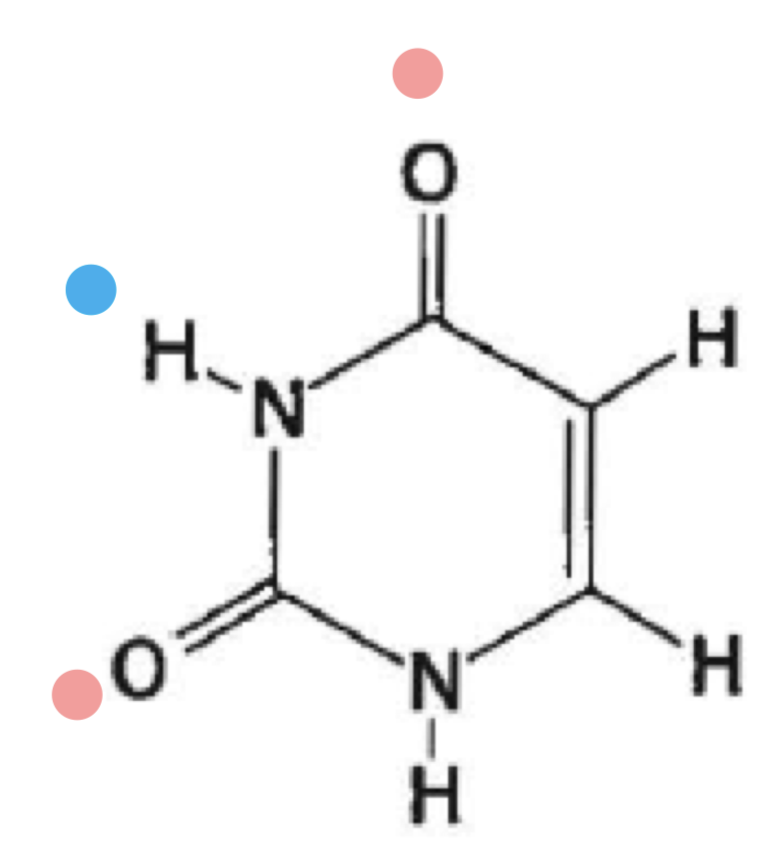
what is this
uracil
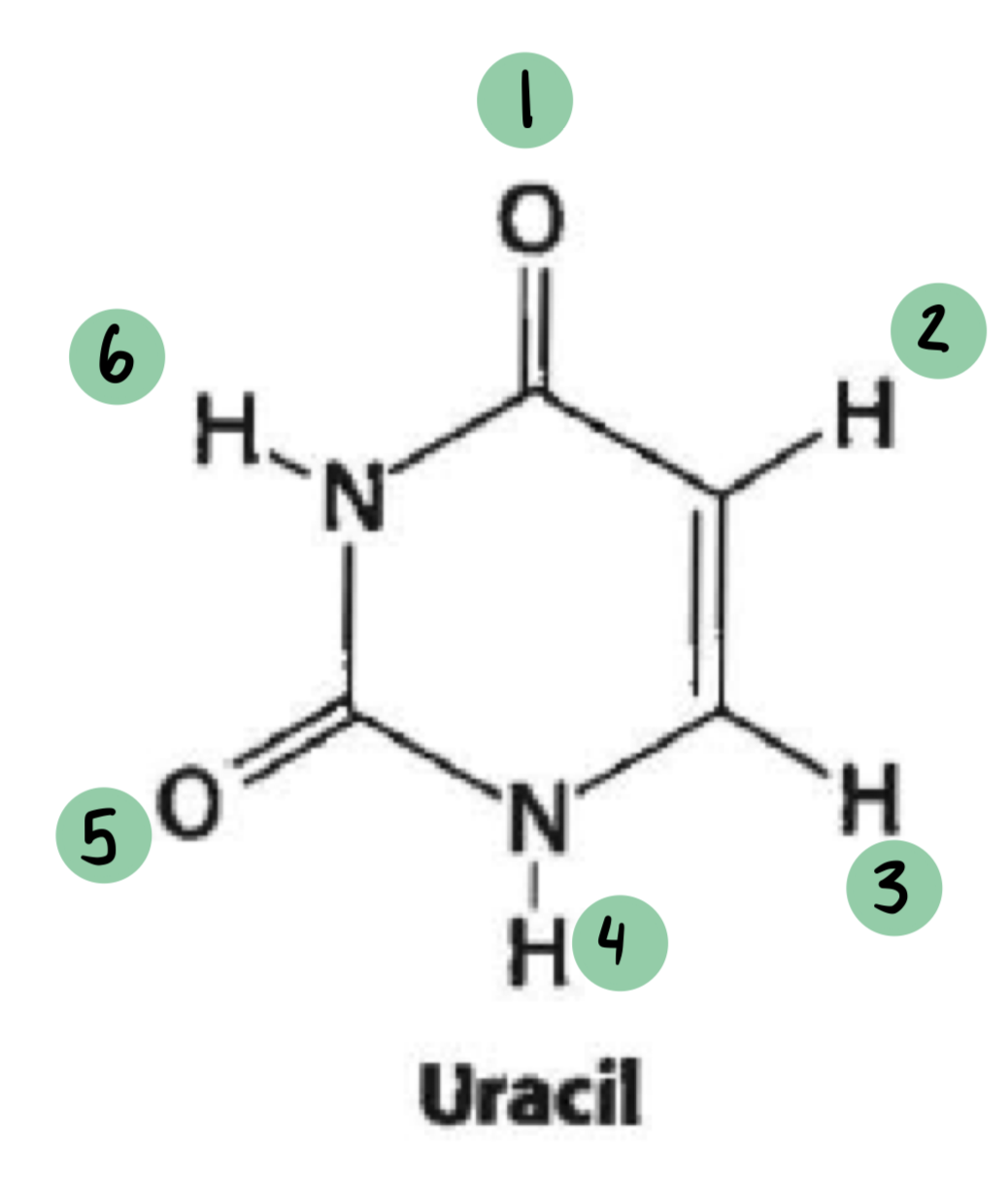
which of these have a 𝛿- and which have a 𝛿+ (not all will have a charge)
𝛿- ~ 1, 5
𝛿+ ~ 6
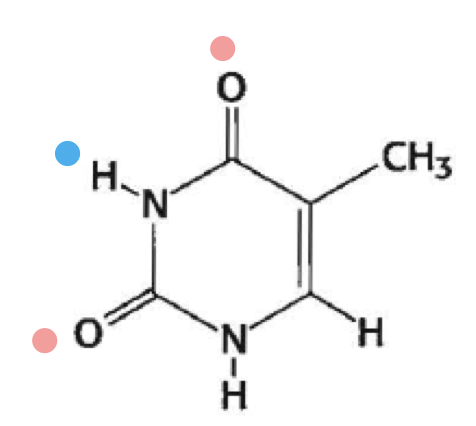
what is this
thymine
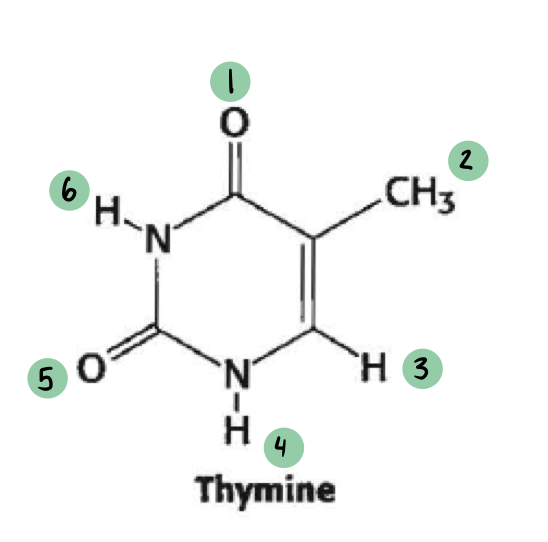
which of these have a 𝛿- and which have a 𝛿+ (not all will have a charge)
𝛿- ~ 1, 5
𝛿+ ~ 6
in watson-crick base-pairing, a GC pair has ____ H-bonds and an AT pair has _____ H-bonds
GC - 3
AT - 2
what is base stacking / pi stacking
where when bases layer on top of eachother, van der Waals interactions contributes stabiity of base pairing
where is the glycosidic bond in DNA / RNA
between N1-C1’ in pyrimidines, and between N9-C1’ in purines
the majority of dsDNA in vivo is in what form
B-form
in the B-form of dsDNA, the sugar takes …?
C2’ endo
when in dehydrated conditions, dsDNA in vitro will take what form?
A-form
in vivo, dsRNA and RNA-DNA hybrids will take what form?
A-form
in the A form, the sugar takes the …?
C3’ endo
what is the Z form of dsDNA?
a left-handed helix with sequence specific (GC repeat)
which is more stable, B-form DNA or A-form DNA?
B-form
which is thinner, A-form DNA or B-form DNA?
B-form
which takes C-2’ endo puckering, A-form DNA or B-form DNA?
B-form
only ___________ allows B form structure (What kind of sugar puckering?)
C2’-endo
in C3’-endo puckering, is the sugar on the same side or the opposite side of the 5’ carbon?
opposite
what is the rise per base pair of the A-DNA structure?
a. 2.3Å
b. 3.4Å
c. 3.8Å
d. 4.2Å
a. 2.3Å
what is the rise per base pair of the B-DNA form?
a. 3.8Å
b. 4.2Å
c. 3.4Å
d. 2.3Å
c. 3.4Å
what is the rise per base pair of the Z-DNA form?
a. 3.8Å
b. 4.2Å
c. 3.4Å
d. 2.3Å
a. 3.8Å
what is the helix diameter of DNA in the A-form?
a. 18Å
b. 20Å
c. 26Å
d. 28Å
c. 26Å
what is the helix diameter of DNA in the B-form?
a. 18Å
b. 20Å
c. 26Å
d. 28Å
b. 20Å
what is the helix diameter of DNA in the Z-form?
a. 18Å
b. 20Å
c. 26Å
d. 28Å
a. 18Å
what is the screw sense of each form of DNA? (A, B, and Z)
A and B - right handed
Z - left handed
how many base pairs are there per turn of helix in A-form?
a. 10
b. 10.4
c. 11
d. 11.4
e. 12
c. 11
how many base pairs are there per turn of helix in B-form?
a. 10
b. 10.4
c. 11
d. 11.4
e. 12
b. 10.4
how many base pairs are there per turn of helix in Z-form?
a. 10
b. 10.4
c. 11
d. 11.4
e. 12
12
what is the pitch per turn of helix in A-form DNA?
a. 25.3Å
b. 35.4Å
c. 45.6Å
a. 25.3
what is the pitch per turn of helix in B-form DNA?
a. 25.3Å
b. 35.4Å
c. 45.6Å
b. 35.4
what is the pitch per turn of helix in Z-form DNA?
a. 25.3Å
b. 35.4Å
c. 45.6Å
c. 45.6
what is the ΔGº for the conversion of A-form DNA → B-form DNA?
~-0.5 kcal/mol(bp) - not much
which form does dsRNA take? why?
A-form, RNA has a bulky oxygen on 2’ carbon which interfere C’-2 endo by steric hinderance
what are the three typical motifs of DNA binding proteins?
helix-turn-helix, leucine zipper, and zinc finger
majority of DNA binding proteins recognize __________ with their ɑ-helices
major grooves
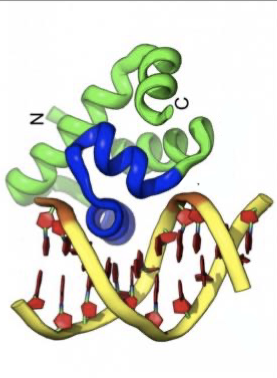
what structure is this?
helix-turn-helix
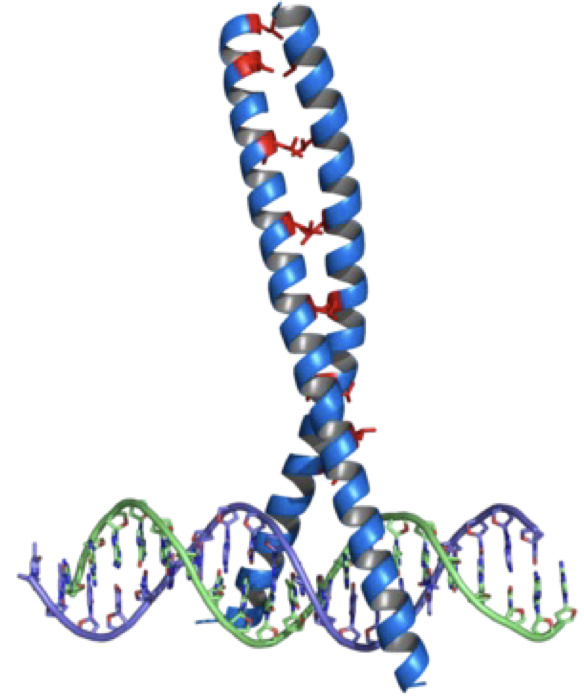
what structure is this?
leucine zipper

what structure is this?
zinc finger motif
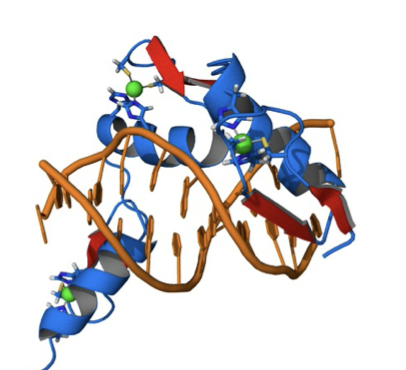
what structure is this?
zinc finger
what is the approximate size of the major groove of DNA?
22Å
what is the approximate size of the minor groove in DNA?
12Å
why is protein binding to the major groove of DNA easier?
size fits well to an ɑ-helix and polar groups in major groove are well exposed
DNA binding proteins can read DNA sequence without unwinding dsDNA up to ______ base pairs
3 to 4
(1 rise = 3.4Å, 10-14Å)
because DNA is twisted, what is the MAXIMUM number of base pairs that a single protein can recognize?
5
a sequence of 5 amino acids will be seen once every how many base pairs?
1024 (4^5)
about how many proteins would it take to recognize a TRULY unique sequence of DNA
3
true or false: histone recognize specific sequences of DNA?
false
how do histone operate?
interact with polynucleotide backbone of DNA by positively charged amino acids
what amino acids do histones typically interact with?
R and K (Argenine and Lysine)
what form is the core of a histone?
octamer
the outside of the histone core (octamer) has lots of __________
positive charges
sequence specific proteins often bind to _________, typically as _________
symmetrical (inverted) DNA sequences, dimers
non-sequence specific proteins bind to DNA based on _____________ rather than a repeating or mirrored sequence pattern
general structural features (i.e. polynucleotide backbone)
____________________ of ɑ-helix allow sequence-specific DNA binding
amino acid-side chains
many recognition sequences are _______________ that are recognized by ________________ of proteins
inverted repeats, symmetric dimer (tetramer)
what kind(s) of protein(s) can recognize non-palindromic sequences
heterodimeric proteins and Zn-finger proteins
non-specific DNA binding proteins often contain _____ side chains to interact with polynucleotide backbone
basic (+)
what are the main functions of DNA polymerase?
nucleotide incorporation mechanism - elongate DNA sequence by adding nucleotides
proofreading
what are the main functions of the sliding clamp/the loader
processivity of DNA synthesis - how efficient a polymerase is and how well it duplicates DNA - high processivity = running a bunch without stopping
what are the main functions of helicase?
opens the replication fork and allows both strands of DNA to be duplicated at the same time
what are the main functions of primase?
start DNA replication by synthesising an RNA primer
what are the main functions of topoisomerase
twists unzipped DNA to prevent supercoiling/untwists to release tension
what are the main functions of telomerase
remakes the buffers (telomeres) on the end of chromosomes that are shortened in each replication of DNA, without them actual genetic material would be lost instead of just the telomeres
what is the approximate speed of DNA replication by polymerase?
500 nt/sec/catalytic subunit
what is the definition of “fidelity” in reference to replicative DNA polymerase?
chance of mistake per synthesis
does replicative DNA polymerase have higher fidellity with or without proof reading?
higher without - high chance of making mistakes becuase it is going so fast
match the following levels of fidelity with their respective DNA polymerase method
a. 10^-3 - 10^-4
b. 10^-6 - 10^-7
c. 10^-9 - 10^-10
with proofreading
without proofreading
in vivo (including mismatch repair)
a-2 ~ 10^-3 - 10^-4 without proofreading
b-1 ~ 10^-6 - 10^-7 with proofreading
c-3 ~ 10^-9 - 10^-10 in vivo (including mismatch repair)
what are three things DNA polymerase ALWAYS needs to function?
needs template (DNA)
needs primer (RNA or DNA)
extends pre-existing 3’OH (cant start from nothing and can’t extend if OH end is blocked)
what does pyrophosphotase do in DNA replication?
enzyme that takes phosphates off the 3’ end to allow the next base to bond
makes it so the polymerase reaction is irriversible, which is GOOD for the formation of DNA
what is the core reaction of DNA synthesis?
(DNA)n + dNTP ←→ (DNA)n+1 + PPi
ΔGº’ = -3.5 kcal/mole
when two bases bind while forming DNA, what kind of bond is formed?
phosphodiester
the reaction: (DNA)n + dNTP ←→ (DNA)n+1 + PPi (ΔGº’ = -3.5 kcal/mole) is done by ________, while the reaction (DNA)n + dNTP ←→ (DNA)n+1 + 2 Pi (ΔGº’ = -7 kcal/mole) is done by ___________
DNA polymerase, Pyrophosphatase
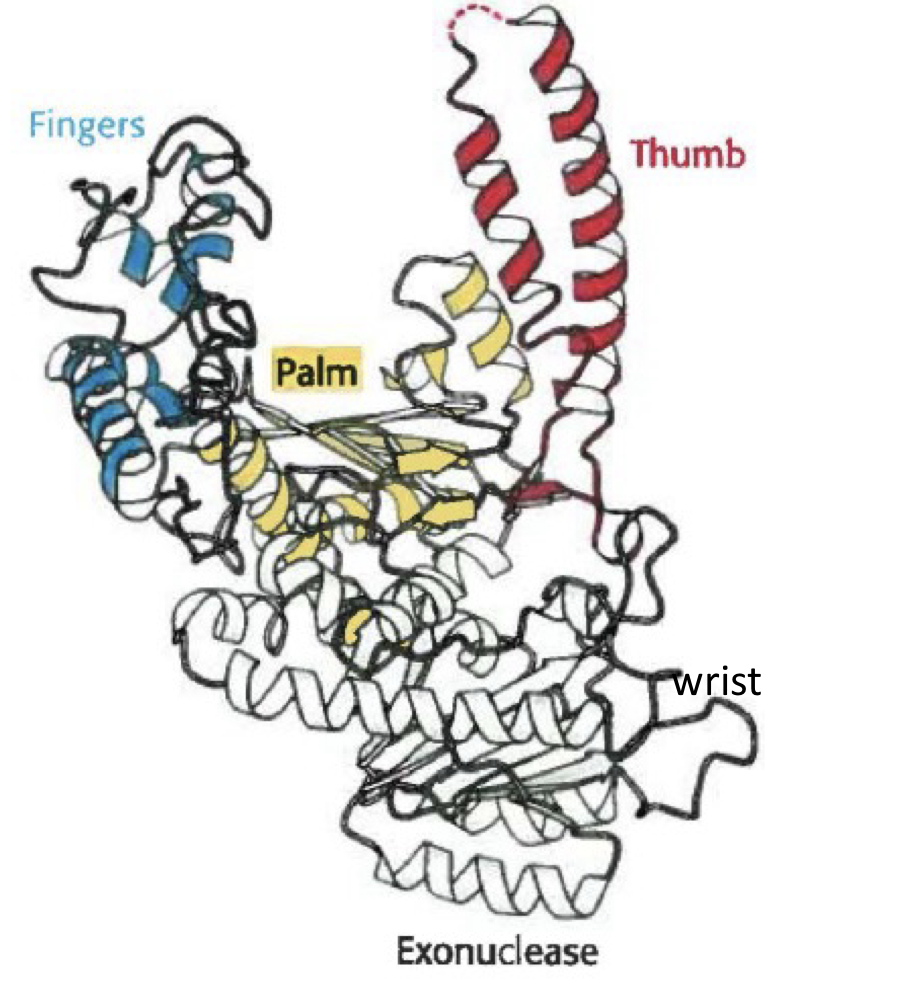
what is the name for this structure? what is it a part of?
Klenow fragment of DNA polymerase I