L22 - Nucleotide Metabolism
1/67
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
68 Terms
where are nucleotides or nucleic acids in the cycle of metabolism
they do not contribute to energy but energy commits to them
how do nucleotides store genetic information
store genetic information as DNA, but some viruses use RNA
how do nucleotides carry and translate genetic information
as mRNA, transfer RNA, and ribosomal RNA
what do nucleotides and nucleic acids use as energy storage molecules
ATP
how do nucleotides carry out enzymatic reactions
ribozymes such as ribosomeal RNA
what do nucleotides and nucleic acids use as signaling molecules
cAMP and cGMP
what do nucleotides use as redox coenzymes
NADH, NADPH, CoA, and SAM
what activates inflammation
viral and bacterial DNA
what are the three parts of nucleotides
phosphate group, pentose sugar (C1 - glycosidic bond, C2 - RNA vs DNA, and C5 - where and how many P bond), and nitrogenous base
what does it mean if the pentose sugar has 2 OH groups
it is RNA
what is a nucleoside
nucleic acid + ribofuranose → the base is always attached to the 1’ position of the sugar to make a beta-N-glycosidic linkage
how do we transition between nucleosides with different numbers of phosphate groups
nucleomonophosphate (NMP), NDP, and NTP can be interconverted by using appropriate kinases - the same processes catalyze formation of deoxyNMP into deoxyNDP and eventually into deoxyNTPs - these building blocks are then used for DNA and RNA synthesis where the ADP is reconverted to ATP using mitochondrial ATP synthase
what is nucleoside monophosphate kinase
the enzyme that phosphorylates NMPs to NDPs using ATP and the enzyme nucleoside monophosphate kinase
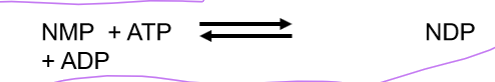
what is nucleoside diphosphate kinase
the enzyme that converts NDP’s into NTP’s
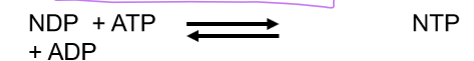
what is required to build nucleotides
very simple molecules - amino acids, one carbon donors, and CO2
why does nucleotide synthesis need to be very tightly regulated
imbalances facor mutation
what is the difference between building purines vs pyrimidines
purine synthesis begins on the ribose sugar, while the pyrimidine rings are synthesized separate from sugar and then attached after it is built
how are purines built
multi-step pathways the starts with ribose-5-phosphate, produced in PPP, and ending with a nucleotide - pathway is very complex and uses many different cofactors and substrates to provide the basic purine ring, which is used in IMP and additional reactions that are required to make AMP or GMP
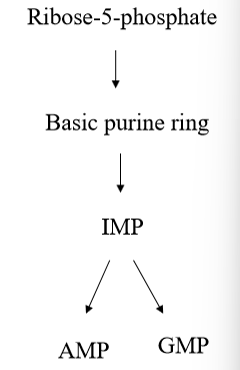
how are purines synthesized
phosphoribosylpyrophosphate (PRPP) is produced, then phosphoribosylamine, then IMP (inosine monophosphate), then AMP/GMP
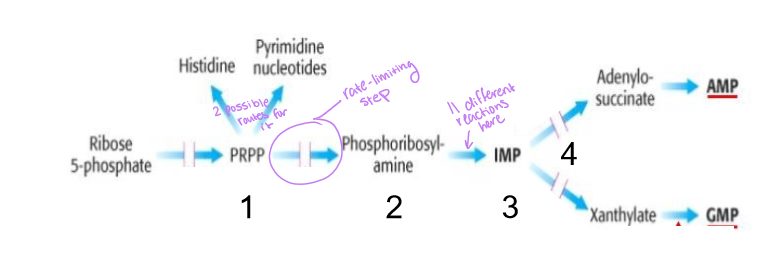
what is the rate-limiting step of purine synthesis
conversion of phosphoribosyl pyrophosphate into phosphoribosyl amine
how is phosphoribosyl pyrophosphate produced
using phosphoribosyl pyrophosphate synthetase and ATP in the first step in producing 5-phosphoribosyl-1-pyrophosphate - ribose-5-phosphate is the starting molecule, and synthetase has kinase activity so it adds pyrophosphate to make PRPP
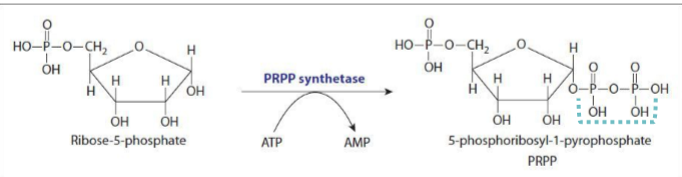
why is the conversion of ribose-5-phosphate into 5-phosphoribosyl-1-pyrophosphate not a committed step
PRPP is also used for histidine biosynthesis and pyrimidine synthesis
what is 5-Phosphoribosyl-1-pyrophosphate (PRPP)
used to make the sugar backbone in all nucleotides - know the full name
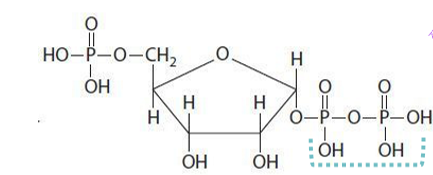
how is phosphoribosyl amine produced in step 2 of purine synthesis
phosphoribosyl pyrophosphate reacts with glutamine to start the formation of the purine ring, which requires the enzyme glutamine-PRPP amidotransferase - THIS IS THE RATE-LIMITING STEP - forms a glycolsidic bond
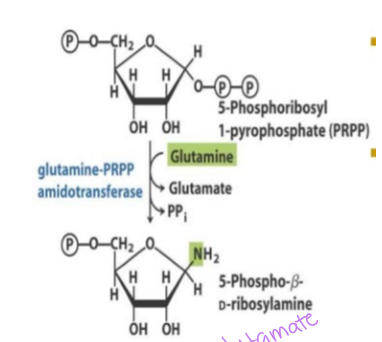
how is the conversion of phosphoribosyl pyrophosphate regulated in purine synthesis
regulated allosterically by the substrates and end products of the pathways, and because it is the committed step, it goes all the way to IMP formation at this point (many steps are required for IMP formation)
what happens to IMP (inosine monophosphate) after it is produced in purine production
gets converted into wither adenylo-succinate and then AMP, or xanthylate and then GMP
how many reactions does it take to build the purine base onto the amine group
11 reactions, and then each additional group comes from a different molecules - N10FH4 (tetrahydro folate from 1C metabolism) donates a methyl group, and then nitrogen is sourced from 4 locations to form the ringV
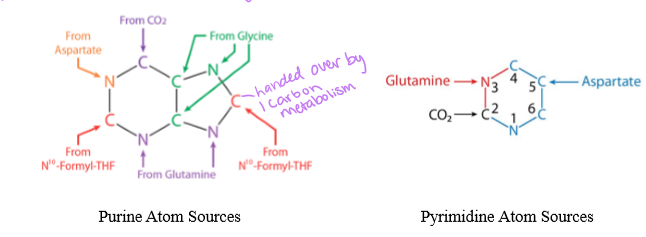
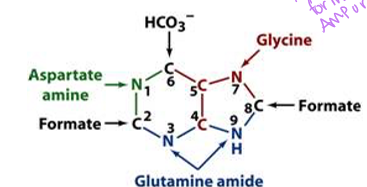
what are the four sources of N for the nine-membered purine ring stucture
glutamine for amide N, aspartate for amino N, glycine, and ammonium
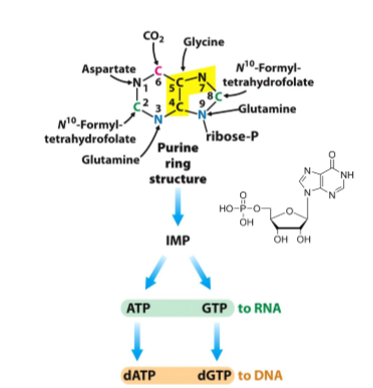
what are all of the substrates that go into purine synthesis
PRPP, glutamine, glycine, N10 -formyl H4 folate, glutamine, HCO3, aspartate, N10-formyl H4 folate
how is purine synthesis regulated
the pathways is subject to several redundant systems and cross-regulation, but metabolites won’t be invested into purine biosynthesis unless ALL CONDITIONS ARE FAVORABLE - all metabolites needed to form the ring and must be ABUNDANT enough to carry out the synthesis reactions (ATP and GTP)
what is the conversion of R5P to PRPP and PRPP to phosphoribosyl amine inhibited by
IMP, AMP, and GMP
what is the conversion of IMP to adenylo-succinate or xanthylate inhibited by
adenylo-succinate is inhibited by AMP and requires GTP to be produced, and xanthylate in inhibited by GMP and requires ATP to then be converted into GMP
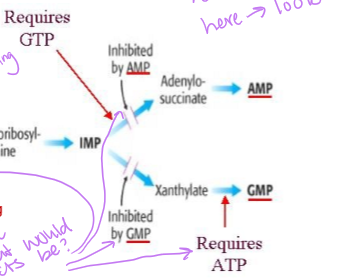
how is AMP produces from IMP
IMP is converted into adenylosuccinate suing GTP and aspartate, which is then converted into AMP releasing fumarate
how is GMP produced from IMP
IMP is converted into xanthosine monophosphate using NAD+ and water, and is then converted into GMP using glutamine and ATP
how are pyrimidines synthesized
the pyrimidine ring base is assembled completely and then PRPP is added as the final step - starts with glutamine, then carbamoyl phosphate, carbamoyl aspartate, the make orotate (2 reactions that close the ring), attach ribose-5-phosphate via PRPP, and then it is decarboxylated to form UMP
inosine monophosphate is the starting substrate generating AMP or GMP - what should you know about them
these reactions are regulated by levels of AMP and GMP and energy is required for these reactions
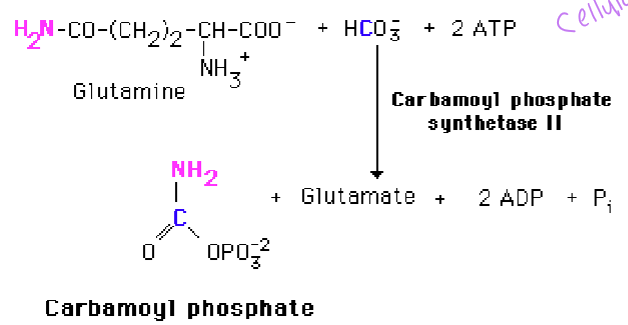
what does carbamoyl phosphate synthetase II use to make carbamoyl phosphate
glutamine and bicarbonate and + ATP - uses glutamine as an N-donor and occurs in the cytoplasm as opposed to CPSI that functions in urea synthesis and uses in ammonia and occurs in the mitochondria
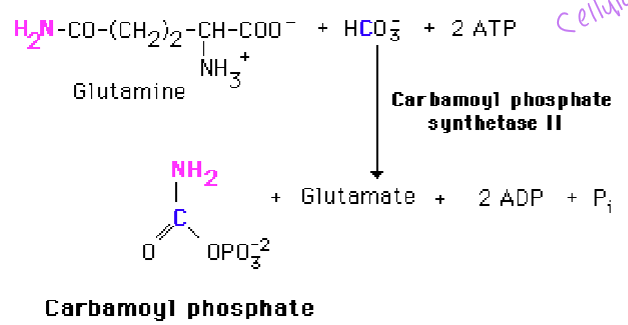
what is the regulatory step of pyrimidine step
carbamoyl phosphate synthetase II, which is activated by PRPP (PRPP dictates synthesis and activity)
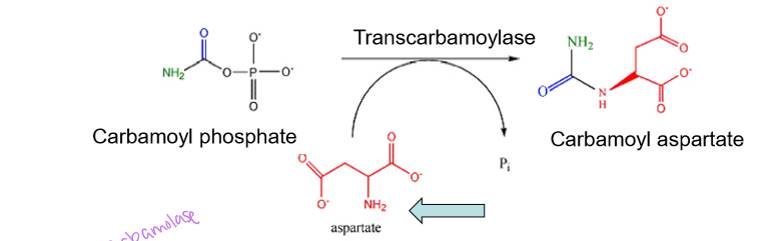
what does aspartate transcarbamoylase do
attaches aspartate to carbamoyl phosphate and removes the phosphate group to produce carbamoyl aspartate- this enzyme starts building the pyrimidine ring
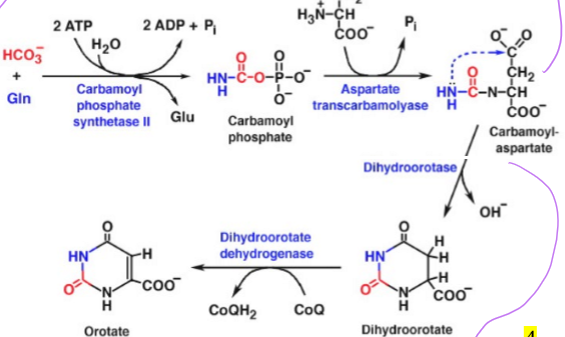
how is the orotate ring of pyrimidine synthesis formed from carbamoyl aspartate
in two enzymatic steps, takes carbamoyl aspartate and use dihydroorotase to form dihydroorotate (removing and OH- group), and then takes dihydroorotate and converts it to orotate using Dihydroorotate dehydrogenase and CoQH2
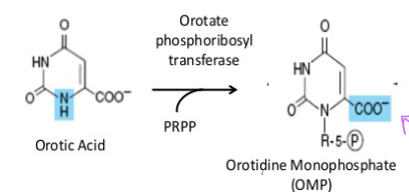
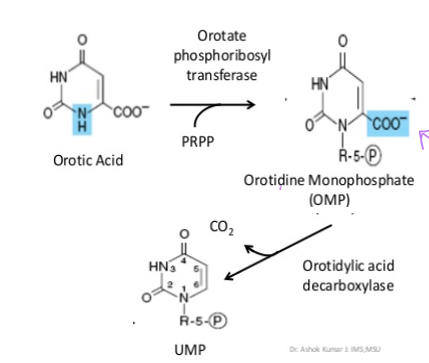
what happens to orotate once it is formed by dihydroorotate dehydrogenase
ribose-5-phosphate is added orotate phosphoribosyl transferase and PRPP to form orotidine monophosphate
what happens to orotidine monophosphate after orotate phosphoribosyl transferase replaces H with R5P
orotidylic acid decarboxylation removes the COO- group in order to form UMP
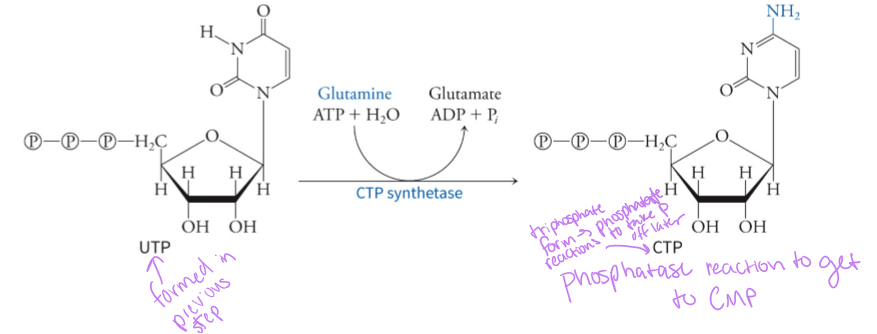
what is the precursor for CMP (Cytidine monophosphate)
UMP (uridine monophosphate) - CTP synthetase aminates UMP, which requires ATP and glutamine, to make CMP - this is what happens to UMP after it is converted by orotidylic acid decarboxylase in pyrimidine synthesis
what does thymidine required that cytosine and uracil don’t
an extra methylation step - requires the generation of dUMP (deoxy uridine monophosphate) which is the starting place for the thymidylate synthase reaction
what does ribonucleotide reductase generate from UDP
deoxy form of CMP for thymined
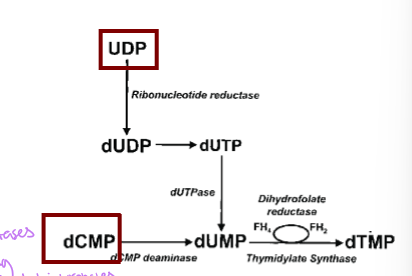
how is deoxy uridine monophosphate generated from dCMP
deoxycytodine monophosphate is converted to dUMP via dCMP deaminase
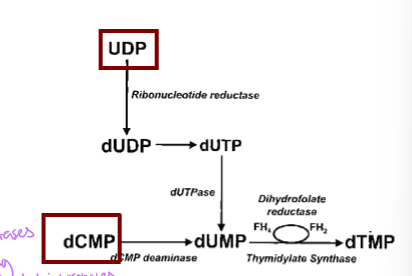
how is dUMP generated from UDP
UDP is converted in two steps - UDP to dUDP via ribonucleotide reductase and then to dUTP, and then dUTP to dUMP via dUTPase to remove the P
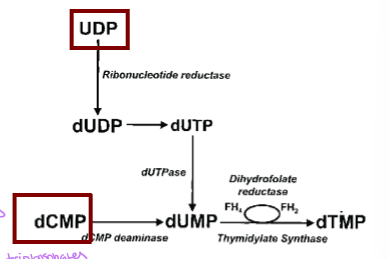
how is dUDP converted to dUTP
through phosphorylation by the enzyme nucleoside diphosphate kinase, using ATP as a P donor
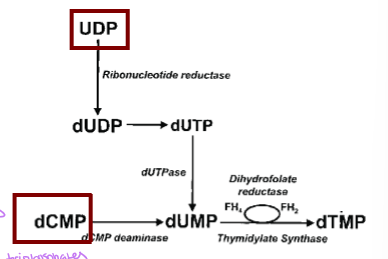
what does dTMP synthesis require
one-carbon metabolism to acquire a methyl group - 5,10-methylenetrahydrofolate and dUMP uses thymidylate synthase to generate dihydrofolate and dTMP

what does one-carbon metabolism do for nucleotide synthesis
provides carbons and contributes to the synthesis of purines and pyrimidines
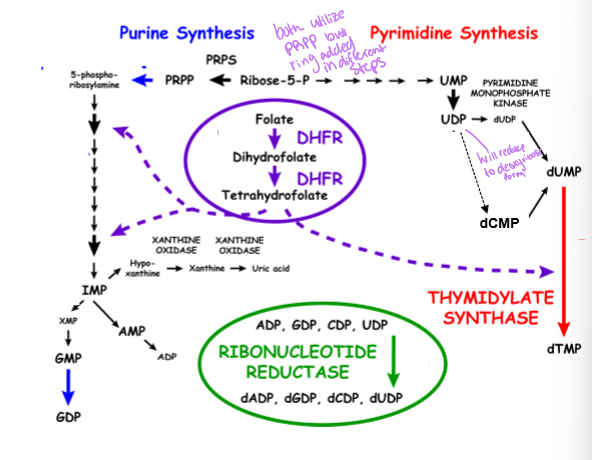
what does ribonucleotide reductase do
converts ADP, DGP, CDP, and UDP to dADP, dGDP, dCDP, and dUDP
what does DHFR (Dihydrofolate reductase) convert
folate to dihydrofolate to tetrahydrofolate - these are the one carbon carriers
how is pyrimidine biosynthesis regulated
primarily controlled and regulated at the carbamoyl phosphate synthetase II step where is it inhibited by CTP (final product) and activated by PRPP (substrates) - purine and pyrimidines both have the commitment to the first N as the rate limiting step
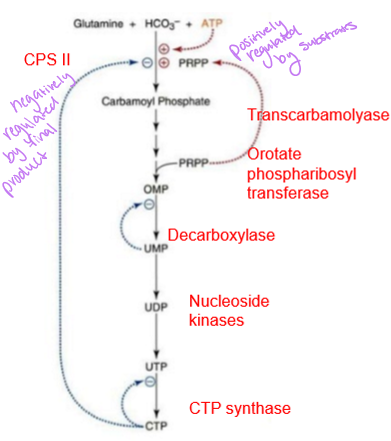
how is deoxyribonucleotides formed from ribonucleotides
ribonucleotide reductase which acts on the diphosphate form (not mono or tri) of ribose, using NADPH as a reducing power - used in DNA synthesis, making this step required to generate usable nucleotides
why is ribonucleotide reductase important
it generates precursors for replication and repair of DNA, is vital for cell proliferation and genome maintenance, and drugs that target this enzyme are used to treat cancers because it prevents cell proliferation - it is also a conserved enzyme in all living organisms that has a shared mechanism of generating DNA (helps us understand DNA damage repair)
what drugs target ribonucleotide reductase and how does that work
cancer-fighting drugs that block the enzyme responsible for creating deoxyribonucleotides (dNTPs), which are essential building blocks for DNA synthesis and repair. By inhibiting RNR, these drugs deplete the nucleotide pool, stalling DNA replication and causing tumor cells to die - hydroxyurea, gemcitabine, etc.
what are thioredoxin and thioredoxin reductase
used as intermediates as the substrate and enzyme in the reduction reaction of ribose to deoxyribose - reduced form reduces RNR, becoming oxidized/regenerated in the process
what does ribonucleotide reductase actually do
removes the 2’-hydroxyl group of the ribose ring of nucleoside diphosphates
what is the starting substrate for the RNR reaction
a nucleoside diphosphate, which is reduced by the reducing power of NADPH
what is ribonucleotide reductase composed of
a large and small subunit that controls balance of deoxyribonucleotides with complex allosteric controls
what does the large subunit of RNR do
has two allosteric sites and the active site
what does the small subunit of RNR do
has a tyrosine that gets radicalized in the reaction mechanism
what is the purine salvage pathway
reversible reaction that salvages the bases that are already formed, and took a lot of energy to make, and recycles them back into nucleotides - this requires PRPP transferase enzyme and the product is the nucleoside monophosphate - this occurs when a cells dies, and the RNA is recycled11
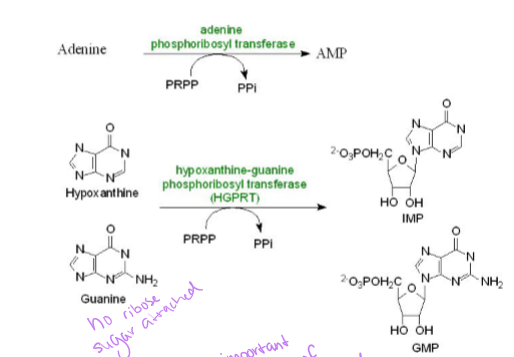
what does the purine salvage pathway do to purines like uracil
reversible phosphorolysis them via kinase activity which could add phosphate to make UDP (uracil to uridine)

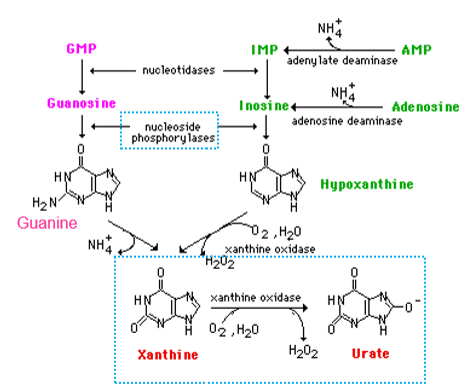
what does purine degradation pathway lead to
uric acid synthesis - this is the primary mechanism to get rid of extra N - it uses nucleoside phosphorylase to break the glycosidic bond, and uses xanthine oxidase and the synthesis of ROS in the form of hydrogen peroxide - urate is excreted via urine
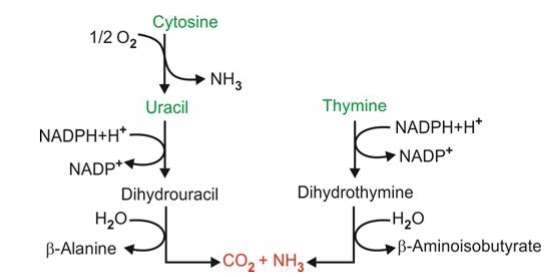
what does the pyrimidine degradation pathways require
NADPH and results in the synthesis of beta-alanine, beta-aminobutyric acid, ammonia, and CO2 - uses nucleoside phosphorylase to break the glycosidic bond using NADPH as the reducing power
what is the summary of salvage
nucleosides are recycled back to nucleotide by addition of ribose sugar backbone, purines require transferase enzyme and PRPP, pyrimidines require uridine phosphorylase and Ribose-1-phosphate
what is the summary of degradation
gets rid of excess nitrogen found in nucleosides, nucleoside phosphorylase breaks the glycosidic bond, which makes the nucleoside available for release in the form of uric acid or ammonia, and there is an overlap with amino acid degradation in using xanthine oxidase and releasing ROS/uric acid in urine