T2 Shimeld- Principles of animal development + Development and diversity
1/5
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
6 Terms
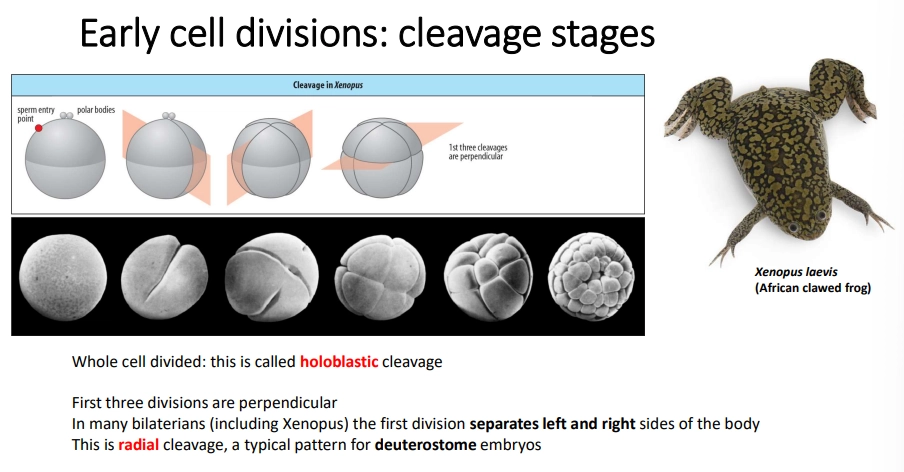
how is the oocyte initially divided in most bilaterian animals?
most animals undergo holoblastic cleavage
bilaterians typically go through three initial divisions
polar bodies (small cells with a discarded nucleus) determine the top
radial cleavage is seen in deuterostomes, where the three divisions are perpendicular
spiral cleavage is often seen in protostomes (ecydosozoa + lophotrochozoa)- the third division is twisted, and the divisions may be equal or unequal, to produce different cells
some species, eg. teleost fish, undergo teleoblastic (instead of holoblastic) cleavage, where cleavage is restricted to just part of the egg due to a high amount of yolk
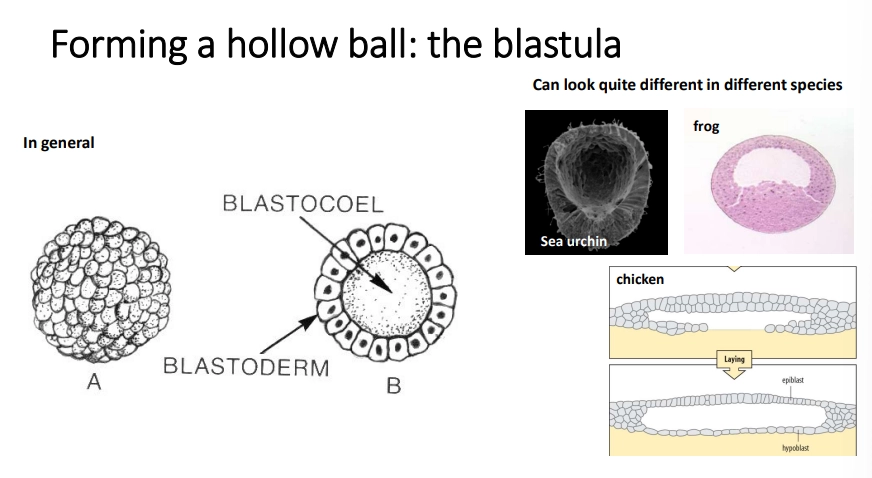
what is the blastula?
bilaterian animals enter a blastula stage during early development, where the zygote has a fluid-filled blastocoel space in the middle surrounded by a layer of cells called the blastoderm
this undergoes gastrulation to move cells into the blastocoel
this produces three cell layers: the ectoderm, mesoderm and endoderm
this movement of cells is an example of morphogenesis (another being neural crest cell migration)
what are the three processes involved in development in animal cells?
differentiation- expression of different transcription factors that activate certain groups of genes (eg. myogenic genes controlled by MyoD) and repress others
pattern formation- lineage dependent mechanisms (programmed asymmetric cell division) + organising fields of cells (using intercellular signalling/morphogen gradients- Wolpert’s french flag model eg. maternal bicoid with mRNA localised at anterior + protein diffusion gradient)- along the three embryo axes, in many cases using particular intermediate Hox genes that encode homeodomain TFs
movement/morphogenesis- gastrulation + neural crest cell migration
what are the two main routes to identifying the genes that control animal development?
biochemistry:
studying compounds that have been extracted from tissues
this relies on fractionation and repeated assay to identify the protein
this works best with intercellular signalling systems, because they can be applied topically
not very useful with intracellular proteins/TFs
forward genetics:
observing the phenotype change from a mutant genotype
most of the genes identified were either for transcription factors or intercellular communication
these are very highly conserved across animal species (ancestral trait)
how do embryonic axes get determined in animal development?
A/P signalling is controlled by Wnt localisation
bilaterians- Wnt localised at the posterior end
non-bilaterians- Wnt localised at the oral end (they have an oral/aboral axis)
D/V signalling is controlled by Bmp localisation
deuterostomes- at the ventral side
protostomes- at the dorsal side
this is developmentally equivalent, mostly due to our labelling of dorsal and ventral according to gravity
how did the developmental gene toolkit evolve?
tandem gene duplication- during meiotic recombination, mistakes in crossing over cause one chromosome to have two copies of a section of DNA, while the other has none
lack of the DNA is likely to be fatal in the inheriting offspring
however the extra copies aren’t necessarily deleterious in the resulting offspring (leads to copy number variants)
whole genome duplication occurred twice in early vertebrate evolution
these events lead to redundancy
these permit the evolution of genes with new developmental functions by mutations (because redundancy means the mutation doesn’t affect the other copy of the original gene)
eg. changing the amino acid sequence of the homeodomain of Hox genes changes the DNA sequence it binds to
how the gene is expressed in different cells can also be changed
eg. changing the position of a particular Hox gene changes how far along the A/P axis the corresponding trait is expressed
repurposing of genes for new functions, eg. using Hox genes to control limb patterning
mutations in gene enhancers can affect gene expression eg. to allow binding of a new TF, prevent binding of an existing one, or create a new enhancer region for a TF