Nucleotide Metabolism
1/37
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
38 Terms
Nucleotide Metabolism
The nitrogen atoms of nucleotide bases are derived from amino acids, whereas the sugar backbones come from ribose.
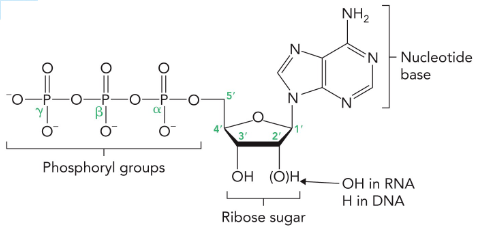
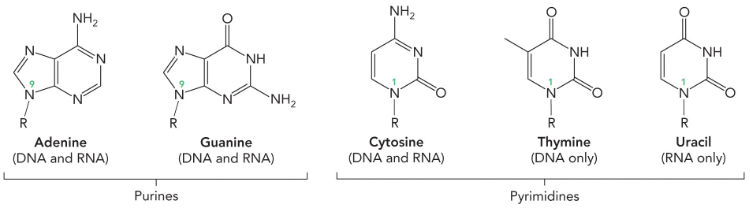
Nitrogenous Bases
Salvage Pathway
Activated ribose (PRPP) + base
recycling nucleotides/nucleotide parts from RNA, DNA and cofactor degradation
Free bases reused to synthesize nucleotide monophosphates
Energy investment not as high as the de novo pathway
Brain requires a continuous supply of pre-formed nucleotides
De novo Biosynthesis
Activated ribose (PRPP) + amino acids + ATP + CO2 + . . .
synthesized “from new” from amino acids, ribose-5-phosphate, CO2, and NH3.
antiparasite drugs
Many parasites (e.g., malaria) lack de novo biosynthesis pathways and rely exclusively on salvage → Compounds that inhibit salvage pathways
Nucleotide Salvage Pathway
uses RNA degradation to produce oligonucleotides, which are cleaved by phosphodiesterases to yield nucleotides in the form of nucleoside 5ʹ-monophosphates.
salvaged by kinases to generate nucleoside triphosphates, which are reincorporated into RNA.
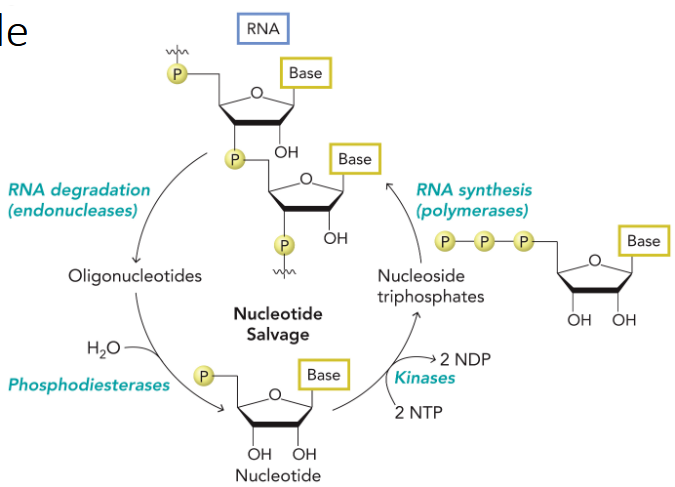
Nucleotide Base Salvage Pathway
produces ribose-5-phosphate and free nucleotide bases, which are attached to phosphoribosyl pyrophosphate (PRPP) by the enzyme phosphoribosyl transferase to yield nucleoside 5ʹ- monophosphates.
Deoxyribonucleotides are salvaged by similar pathways.
De Novo Biosynthesis of Purines Pathway
Approximately the same in all organisms studied
• Glu provides most amino groups.
• Gly is precursor for purines
• Asp is precursor for pyrimidines
• Formate supplied as N10-formyltetrahydrofolate
Nucleotide pools kept low, so cells must continually synthesize them: may limit rates of transcription and replication
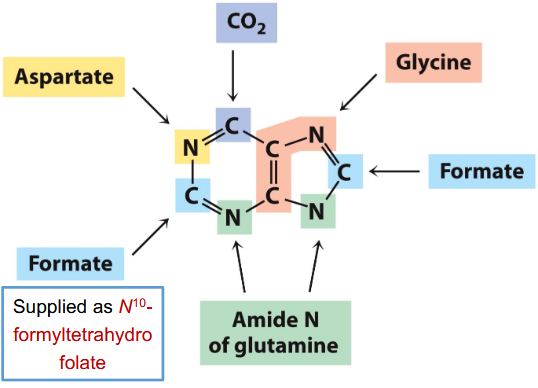
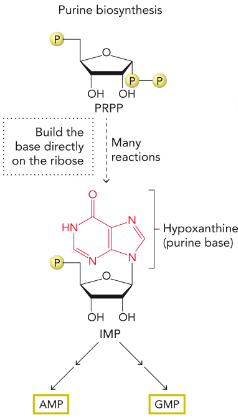
Purine De Novo biosynthesis
synthesized directly on the ribose sugar PRPP to generate the common intermediate IMP, which contains the purine base hypoxanthine
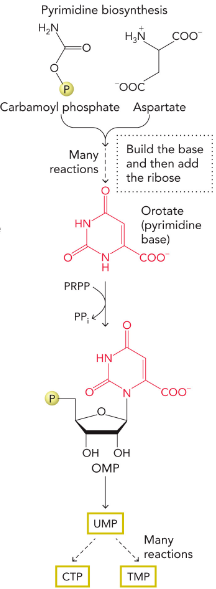
Pyrimidine de novo Biosynthesis
synthesized from carbamoyl phosphate and aspartate to yield orotate, which is then linked to PRPP to generate OMP
5-Phosphoribosyl-1-Pyrophosphate (PRPP)
Synthesized from ribose 5-phosphate of Pentose Phosphate Pathway (PPP) via ribose phosphate pyrophosphokinase: highly regulated allosteric enzyme
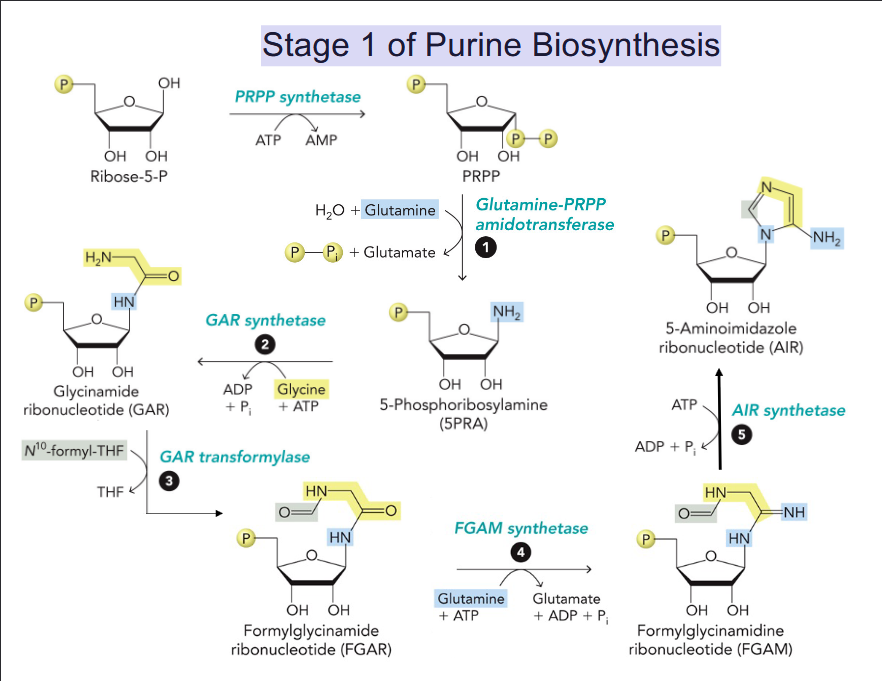
Stage 1 of Purine Biosynthesis
Synthesis begins with reaction of 5-phosphoribosyl 1-pyrophosphate (PRPP) with Glu.
Purine ring builds up following the addition of three carbons from glycine.
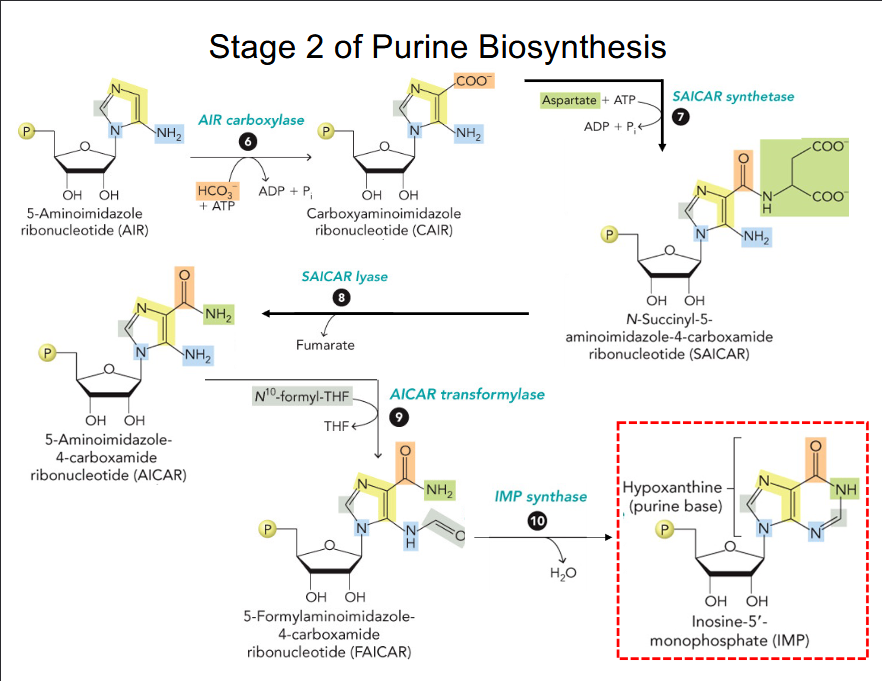
Stage 2 of Purine Biosynthesis
The first intermediate with a full purine ring is inosinate (IMP).
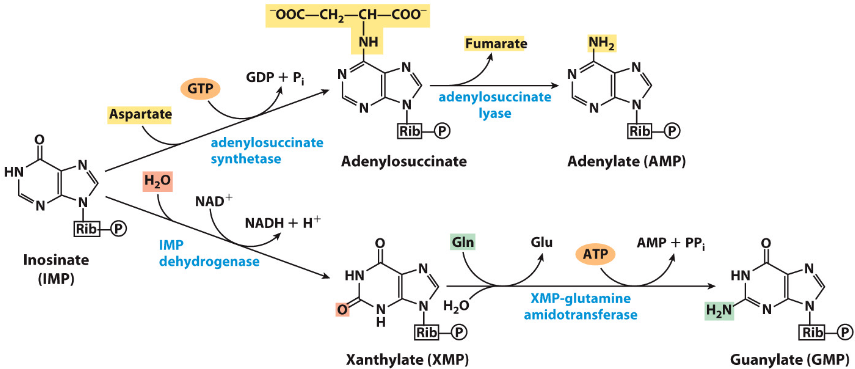
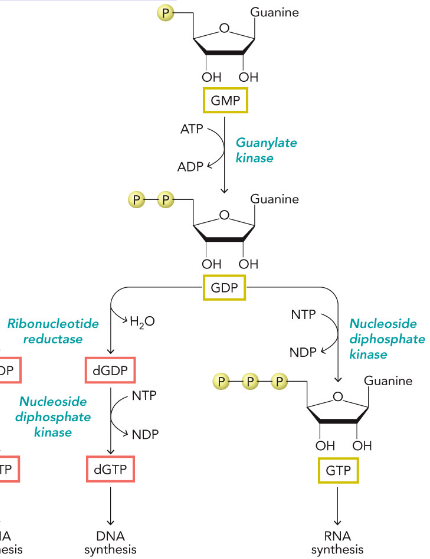
Synthesis of AMP and GMP from IMP
Note that ATP is used to phosphorylate GMP precursor, while GTP is used to phosphorylate AMP precursor
Hence, GTP is needed to make AMP while ATP is needed to produce GMP
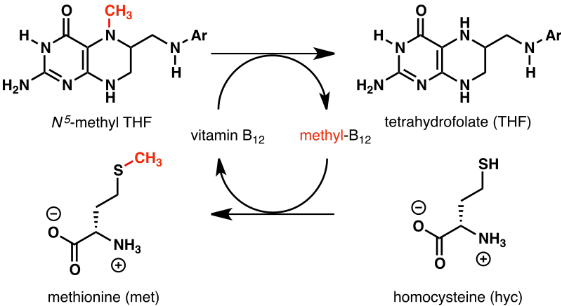
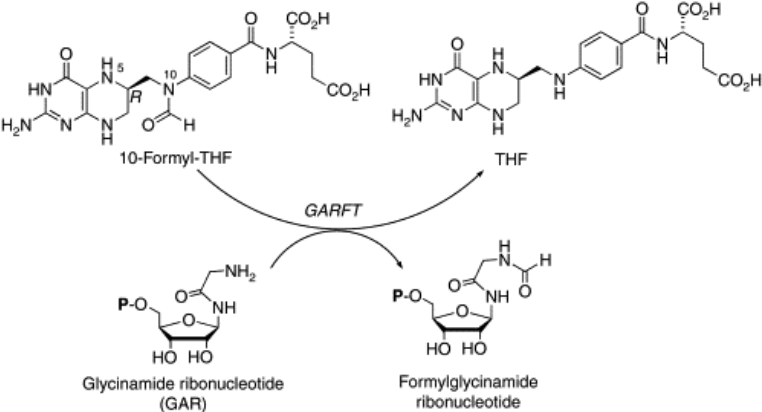
Tetrahydrofolate Reactions
1-carbon carrier - methylation, formylation
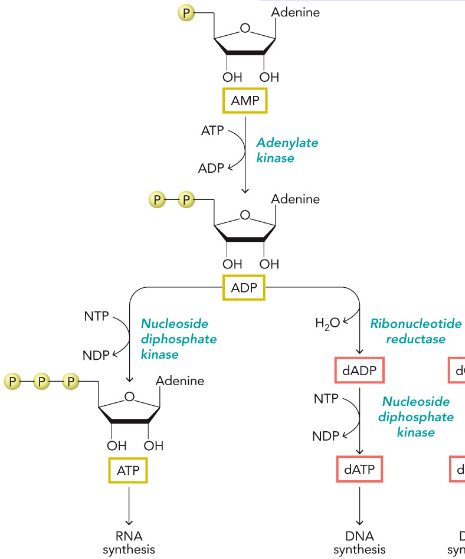
AMP/ATP/dATP Synthesis
GMP/GTP/dGTP Synthesis
Regulation of Purine Biosynthesis (E. coli)
1. Glutamine-PRPP amidotransferase is inhibited by end-products IMP, AMP, and GMP.
2. Excess GMP inhibits formation of xanthylate from inosinate by IMP dehydrogenase.
3. GMP and AMP concentrations inhibit phosphorylation steps.
4. PRPP synthesis is inhibited by ADP and GDP.
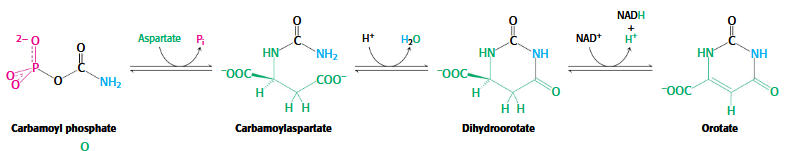
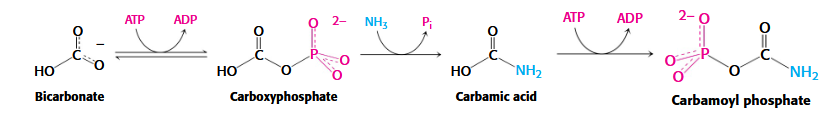
Stage 1 of Pyrimidine Biosynthesis
carbomoyl phosphate → carbomoylasparatate → dihydroorotate → orotate
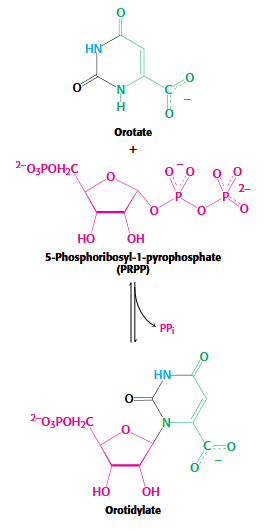
Stage 2 of Pyrimidine Biosynthesis
orotate + PRPP → Orotidylate
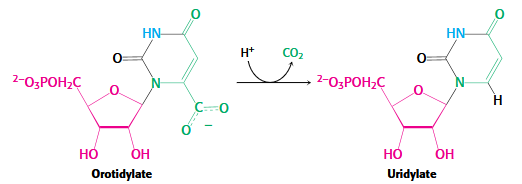
Orotidylate decarboxylase
OMP → UMP
Rate acceleration 10^17
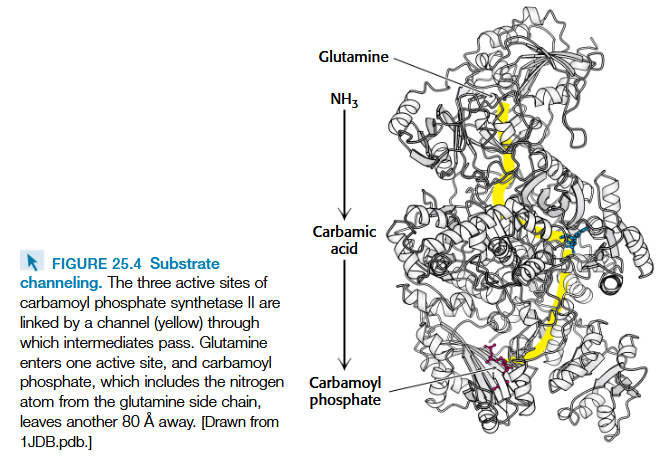
carbomoyl phosphate synthetase II
glutamine nitrogen source
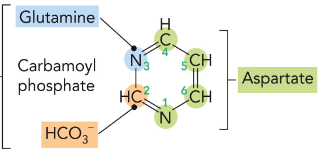
De Novo Biosynthesis of Purines Pathway
All atoms derived from aspartate and carbamoyl phosphate.
UMP is formed in 6 steps
CTP is derived from UTP
Ribonucleotide Reductase
Ribonucleotides are Precursors to Deoxyribonucleotides
2’C-OH bond is directly reduced to 2’-H bond
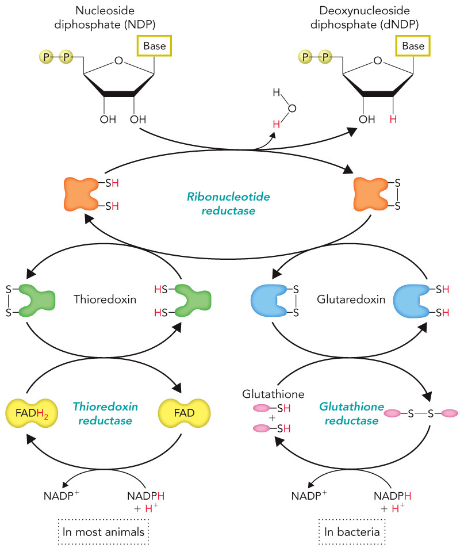
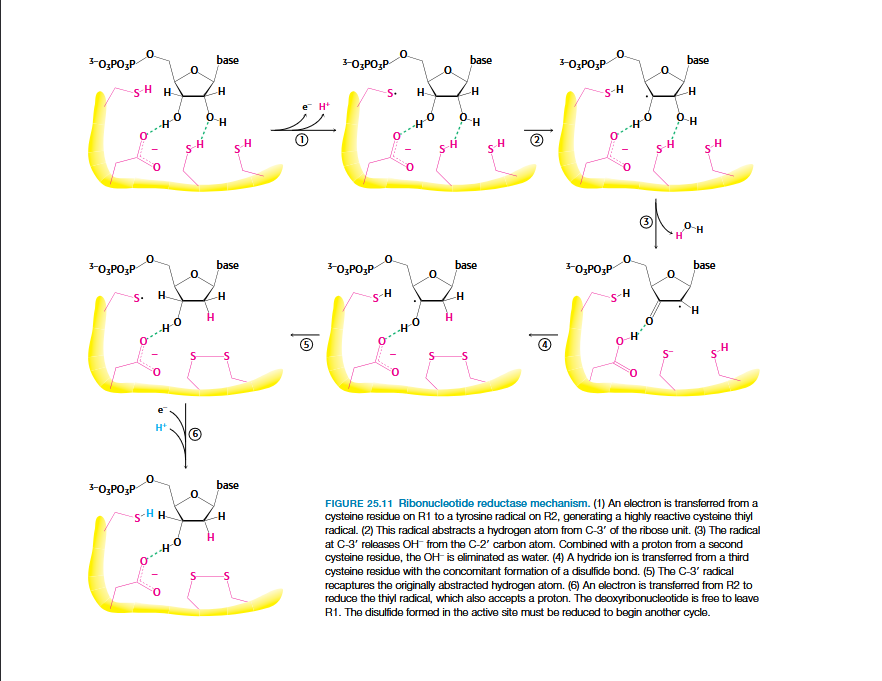
Ribonucleotide Reductase Mechanism
Two H atoms are donated by NADPH and carried by proteins thioredoxin or glutaredoxin.
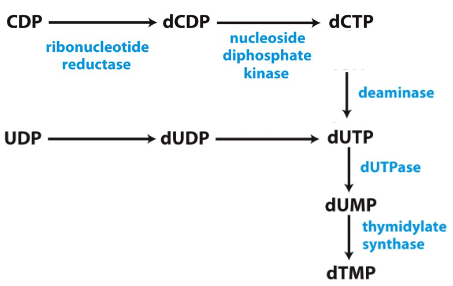
dTMP
Made from dUMP
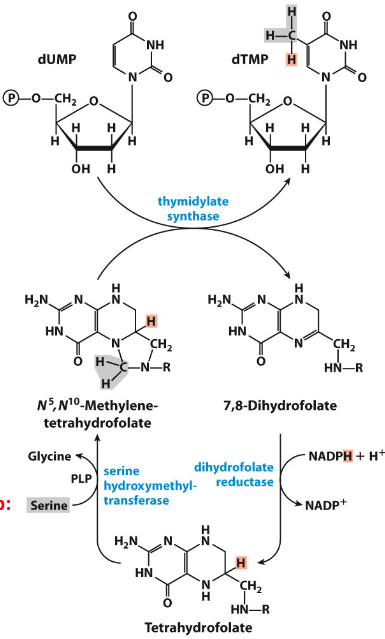
N5-N10-Methylene-tetrahydrofolate
Serine provides the methylene group
Folic Acid Deficiency
Reduced Thymidylate Synthesis → causes uracil to be incorporated into DNA → Repair mechanisms remove the uracil → create strand breaks that affect the structure and function of DNA → associated with cancer, heart disease, neurological impairment
Folic acid Anemia
nutritionally poor populations

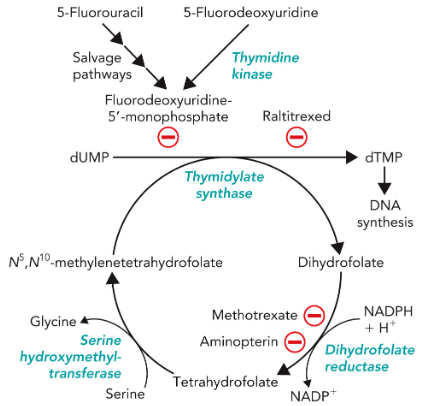
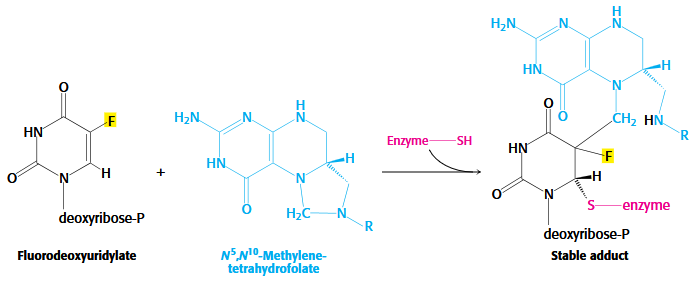
Inhibitors of Thymidylate Synthesis
Folate analogs as potent anticancer medications
Rapidly dividing cells require an abundant supply of thymidylate for the synthesis of DNA.
The vulnerability of these cells to the inhibition of TMP synthesis has been exploited in the treatment of cancer.
Thymidylate synthase and dihydrofolate reductase are choice targets of chemotherapy
ex. Raltitrexed and Methotrexate
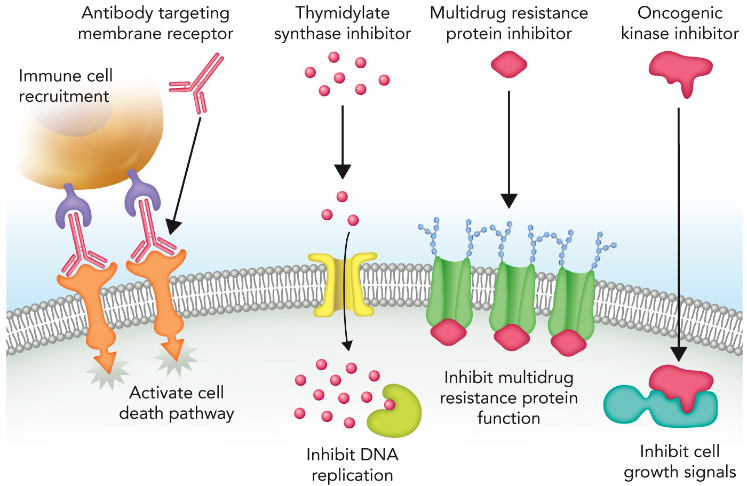
Anticancer Drug “Cocktails”
Each drug blocks a specific process in order to maximize chances of killing the cancer cells before drug resistance begins.
Verapamil, combination of methotrexate and 5-fluorouracil
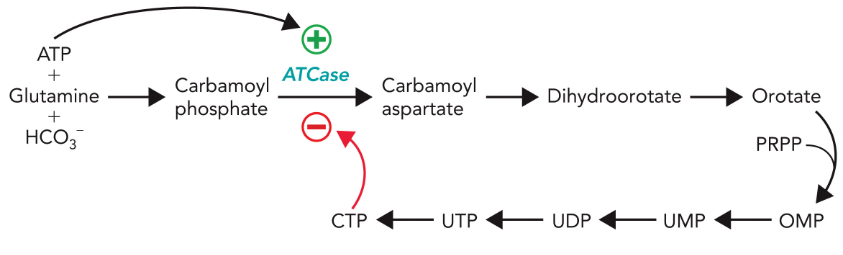
Pyrimidine Biosynthesis Regulation
ATCase is the key regulated enzyme in the pyrimidine biosynthetic pathway in E. coli cells, being activated by ATP and inhibited by CTP - feedback mechanism
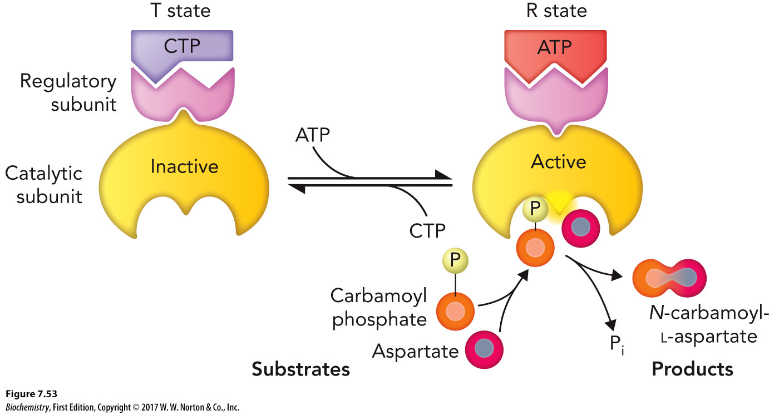
ATCase allosteric regulation
CAD
quaternary organizations of prokaryotic ATCases
•Carbamoyl-phosphate synthetase II
•Aspartate transcarbamoylase
•Dihydroorotase
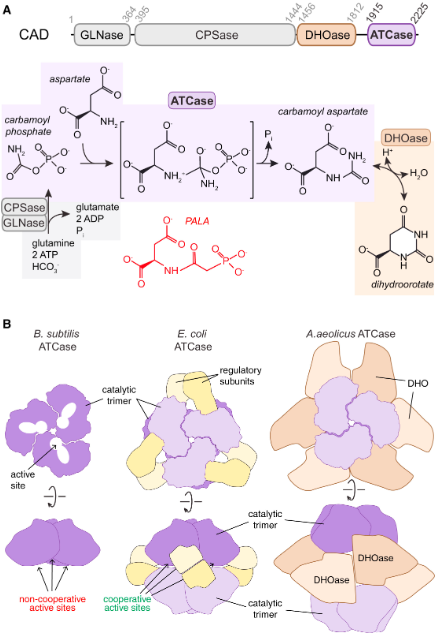
PALA
competitive inhibitor of ATCase
Catabolism of Purines
1. Dephosphorylation (via 5’- nucleotidase)
2. Deamination and hydrolysis of ribose lead to production of xanthine.
3. Hypoxanthine and xanthine are then oxidized into uric acid by xanthine oxidase.
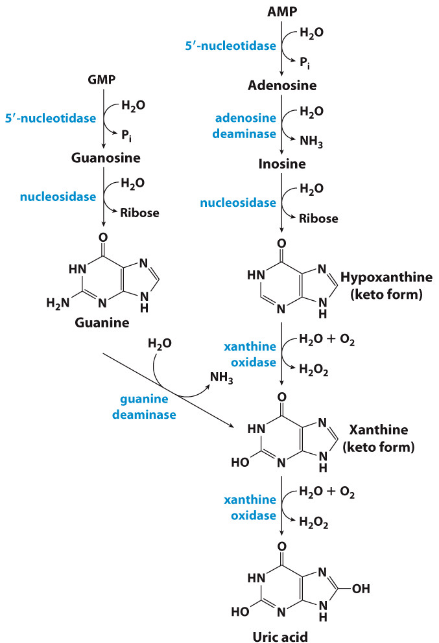
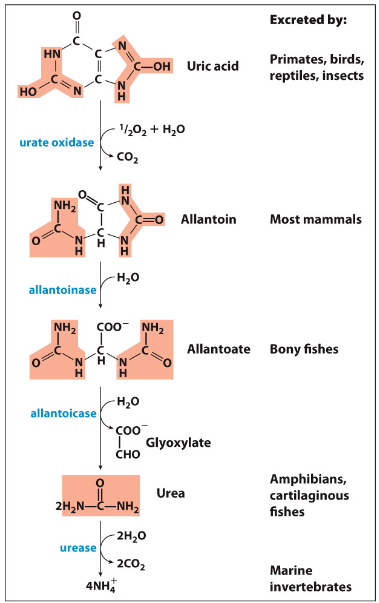
Conversion of Uric Acid
Allantoin, Allantoate and Urea
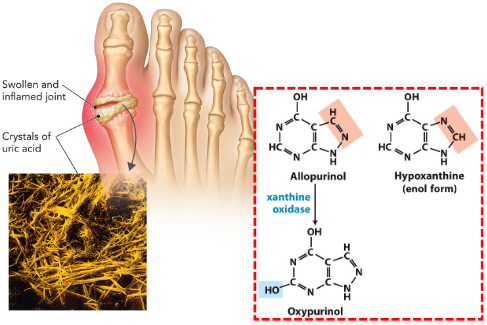
Gout Arthritis
Caused by a defect in purine degradation
Uric acid crystals buildup in joints & kidneys
Treated with avoidance of purine-rich foods (seafood, liver)
Xanthine oxidase inhibitor allopurinol
Catabolism of Pyrimidines
Leads to NH4+ and urea
Can produce intermediates of TCA
Thymine is degraded to methylmalonyl CoA