"Central Dogma"
1/63
Earn XP
Description and Tags
DNA Replication, Transcription, and Translation
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
64 Terms
The two main properties of DNA Replication (elaborate)
Semi-Conservative: Demonstrated by the Messelson-Stahl experiment
Bidirectional: 2 replication forks move in opposite directions away from each (future) origin, forming a replication bubble
Refer to new strands as ‘daughter’, and OG as ‘parental’
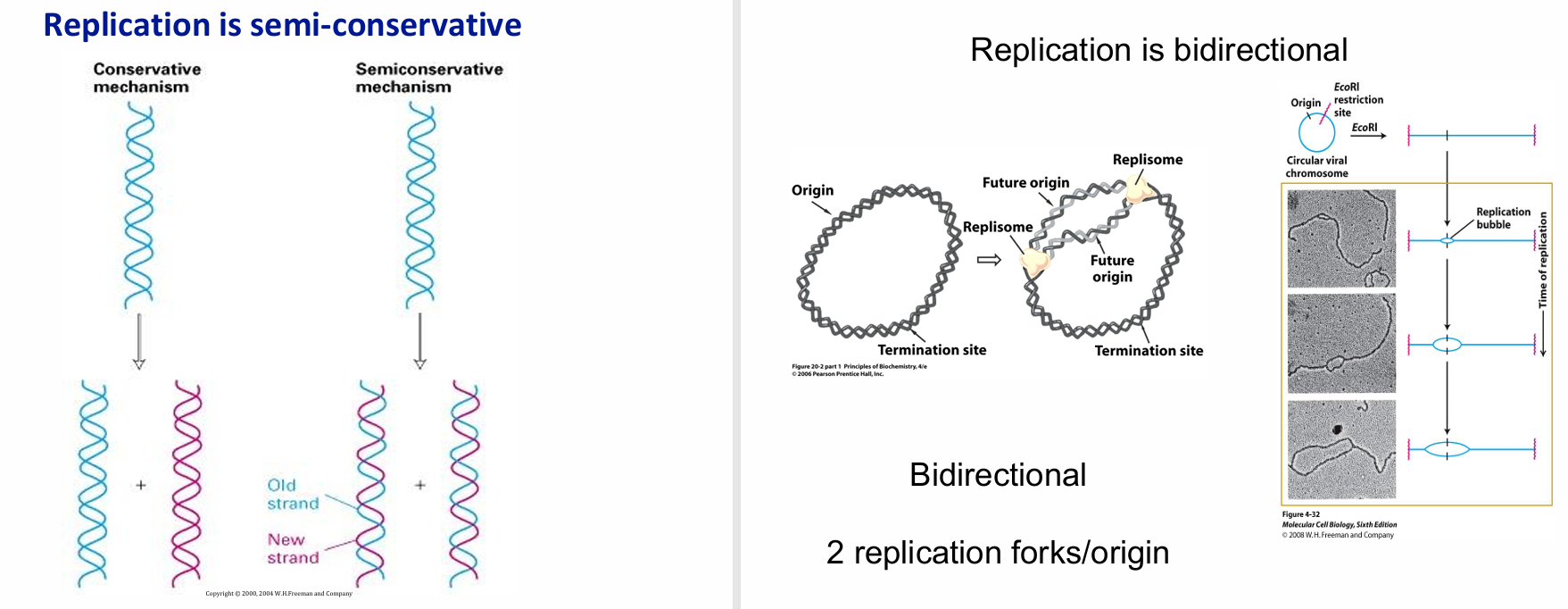

What is meant by OriC and replicon?
Origin of replication in prokaryotes
DNA replicated from one origin of replication
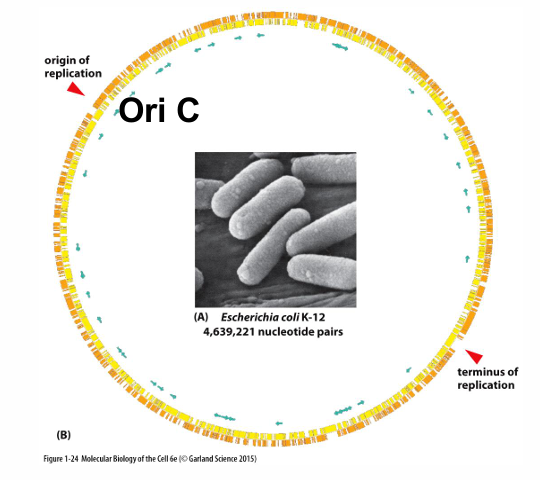
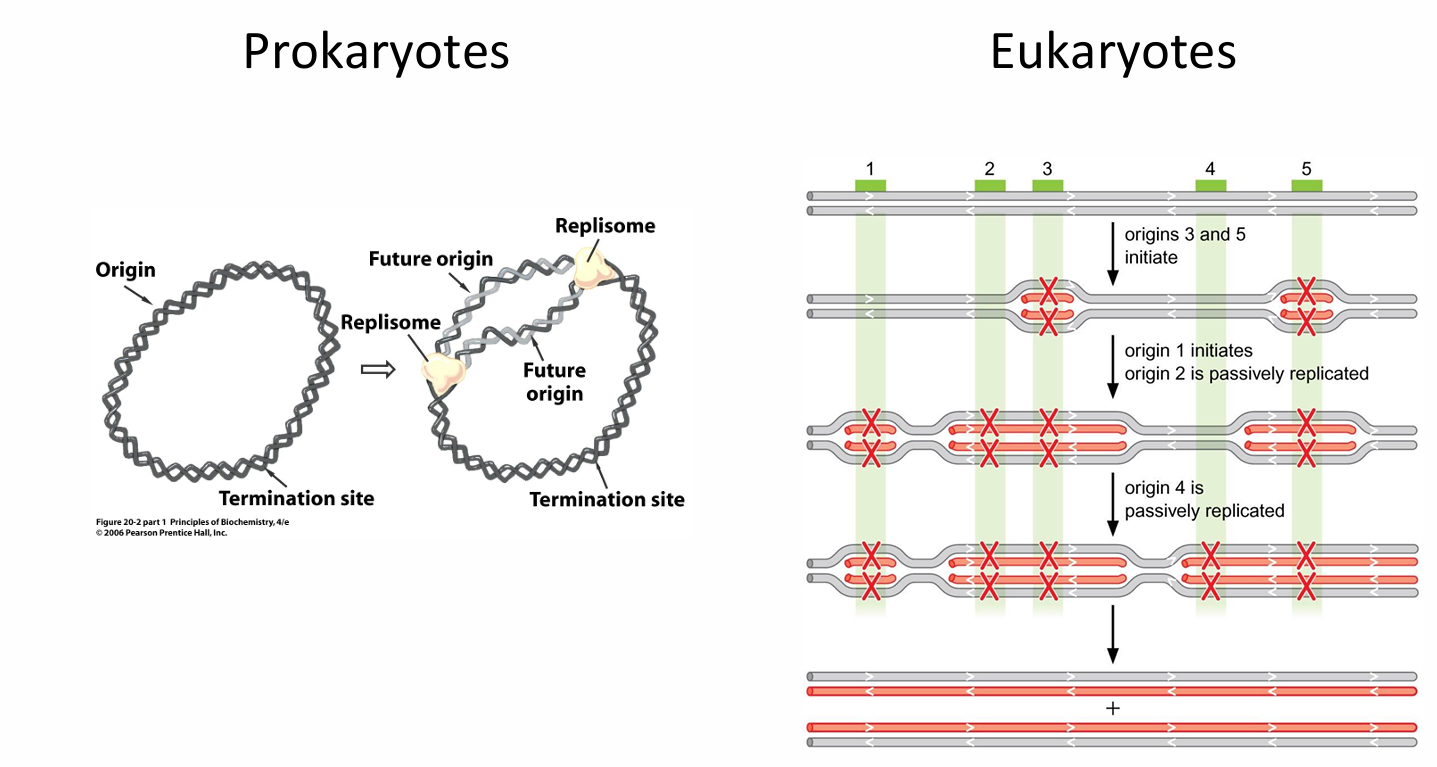
What are the key differences between origin of replication in prokaryotes and eukaryotes?
Prokaryotes | Eukaryotes | |
No of origins | Single (OriC) | Many |
No of replicons | Single | Many |
In eukaryotes
Origins fire in a staggered order - some are passively replicated by neighbouring forks
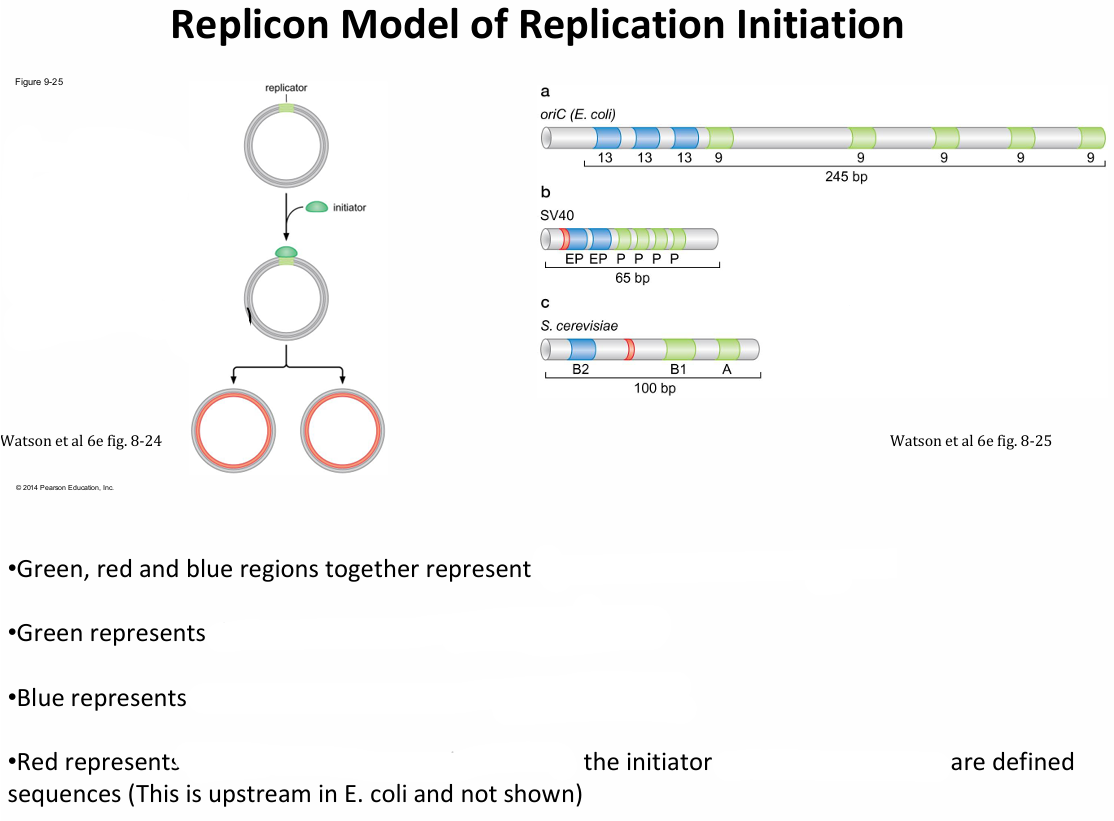
Explain the Replicon Model, and what are the three key components
Replicator: DNA sequence that is sufficient to direct the initiation of DNA replication. (blue, green and red regions)
Origin of replication: The site where the DNA is unwound and DNA synthesis begins (red)
Initiator: Is the protein that recognises and binds the replicator.
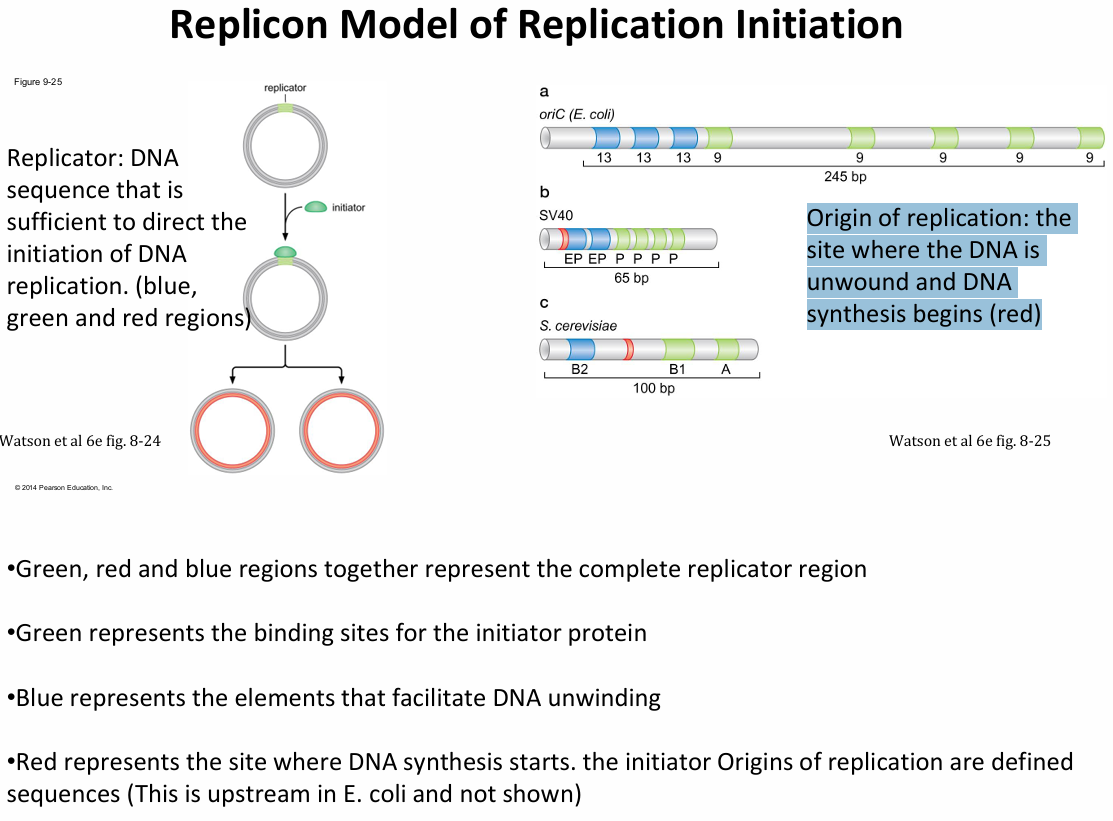
In the E. coli oriC structure, how is the 13-mer repeats advantagous?
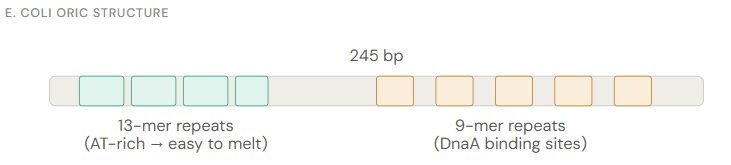
What are the roles of DnaA, DnaB, & DnaC in Initiation? In E. coli
DnaA (initiator protein) bound to ATP
Binds as a multimer to the double-stranded 9-mer repeats. When bound to ATP, DnaA also interacts with the AT-rich 13-mer region, melting it into single-stranded DNA and forming the initial "open complex".
DnaB (helicase) + DnaC (helicase loader)
DnaA recruits a DnaB–DnaC complex to the open complex. DnaC keeps helicase inactive until it loads it onto single-stranded DNA, then DnaC departs — this activates DnaB, which begins unwinding the duplex.
Primase recruitment
Active DnaB, helicase, recruits primase (DnaG), which synthesises short RNA primers ==> Causes helicase to dissociate
DNA polymerase III holoenzyme then extends these primers, and replication elongation begins.
Why is primase essential?
DNA polymerases cannot initiate chains from scratch — they can only extend a pre-existing primer (RNA or DNA).
In which direction does DNA polymerases synthesise DNA, why is this problematic, and why does synthesis occur differently on each strand?
5’ ==> 3’
Because the two strands are antiparallel, ‘leading’ and ‘lagging’ strand
What are the differences between elongation on the leading strand and the lagging strand?
Synthesis? | Continuous | Discontinuous
|
No of short RNA primer(s)? | 1 | Each Okazaki fragment starts with one |
Direction of growth | Same direction as fork movement | Opposite to direction of fork movement |
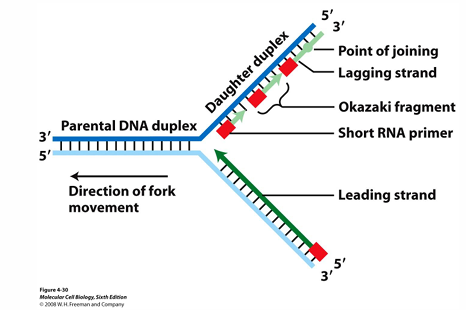
What is the key difference in the direction the template is read and new strand is synthesised?
Template is read from 3’ to 5’
Synthesis is from 5’ to 3’
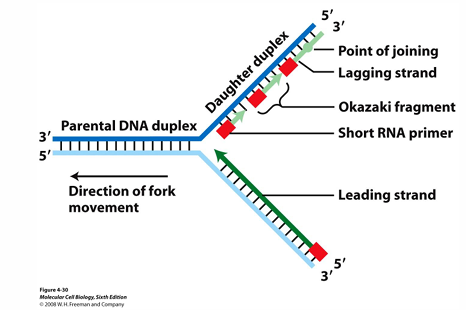
What are the purposes of DNA polymerase I?
Synthesises DNA to complete the lagging strand (i.e. connect the Okazaki fragments)
Removes RNA primers from Okazaki fragment on the lagging strand, and replaces it with itself
How do you calculate fork speed?
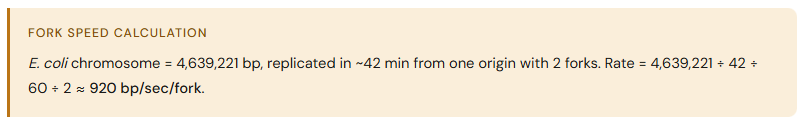
Until when does DNA replication proceed?
Until the entire genome is replicated
In prokaryotes with circular chromosomes, how does elongation come to an end?
The two converging replication forks meet at Ter (termination) regions
What are catenanes, and when is it produced?
2 interlinked circular pieces of DNA/ DNA molecules
Produced after the completion of replication
What do topoisomerases resolve? How many types are there, and what does each type do?
Resolves topological problems during and after replication
Type I
Relaxes supercoils/DNA
By nicking and closing one strand of the duplex DNA
Passes the other strand through the break
Type II
Change DNA topology by breaking and rejoining double-stranded DNA
Cuts both strands of the duplex
Passes another duplex through the break
Separates catenanes; introduces/removes supercoils
Requires ATP
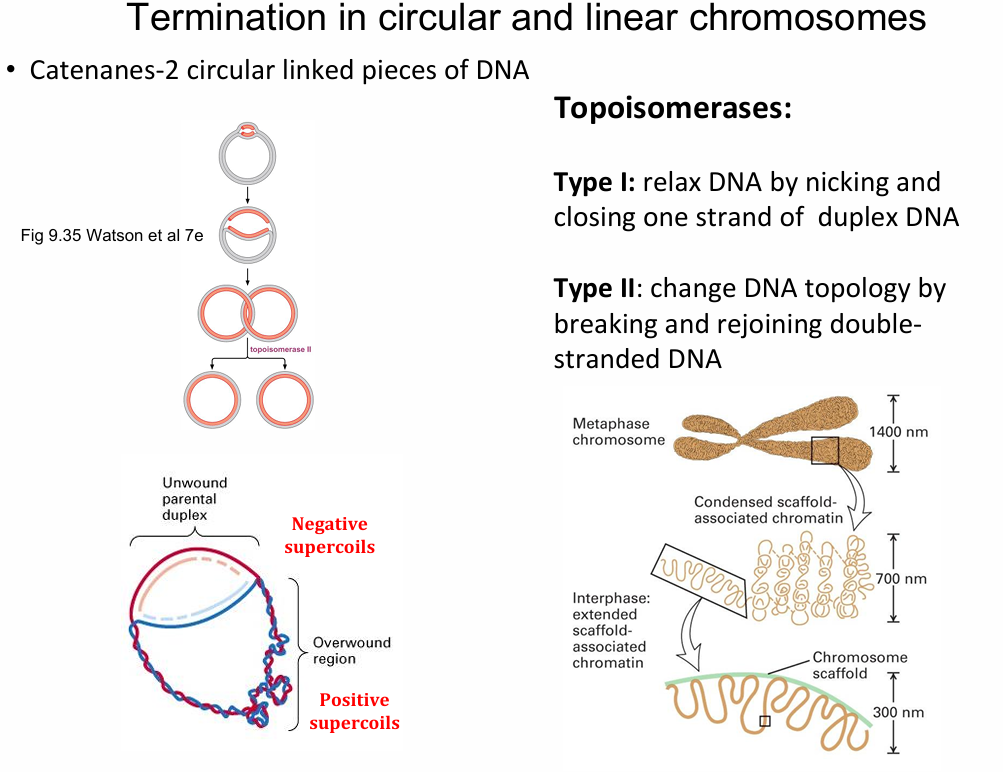
In which part are positive and negative supercoils generated in circular DNA?
Unwinding generates positive supercoils, ahead of the replication fork
What is therapeutic link between DNA replication and anticancer drugs?
Many anti cancer drugs target DNA synthesis; cancer cells are particularly susceptible since they are rapidly dividing.
Target topoisomerases (also by antibiotics)
What is the accuracy of DNA replication like?
Very accurate/ extraordinary fidelity
10-9 to 10-10 errors per base pair per round of replication
What is the main function of DNA polymerase? Explain its structure + how it brings about its function
Proofreading
Hand-like shape/Resembles a hand (thumb, palm, fingers)
Fingers and thumb grip the DNA and help position the template
The palm contains the polymerase active site (Pol) where new nucleotides are added to the growing strand
Exo (exonuclease) = Proofreading site
In the left panel, everything is going normally. The DNA is sitting in the polymerase active site, and nucleotides are being added to the 3' end of the growing strand, reading the template 3'→ 5' and synthesising 5'→ 3' as usual.
In the right panel, something has gone wrong — a wrong base has been incorporated. The enzyme detects this because a mismatch causes a physical distortion in the DNA helix.
A correct Watson-Crick base pair fits snugly in the active site, but a mismatch doesn't sit right, and the polymerase can sense this altered geometry.
When a mismatch is detected, the 3' end of the growing strand shifts from the Pol site over to the Exo site. The exonuclease then clips off the incorrect nucleotide using 3'→5' exonuclease activity — it chews back from the end. Once the mismatched base is removed, the strand slides back to the Pol site and normal synthesis resumes.
What biomolecular technique exploits DNA replication in vitro and how? Explain the process briefly
Polymerase Chain Reaction (PCR): To amplify a specific DNA sequence exponentially
Uses…
A thermostable DNA polymerase (e.g. Taq)
Two synthetic primers flanking the target region
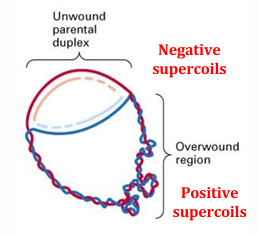
1) Denaturation (~95 °C)
Heat separates the double-stranded DNA into two single strands.
2) Annealing (~55–65 °C)
Temperature lowered so primers hybridise to complementary sequences on each strand.
3) Extension (~72 °C)
Taq polymerase extends from each primer in the 5′→ 3′ direction, synthesising new complementary strands. Each cycle doubles the target DNA — after n cycles: 2n copies.
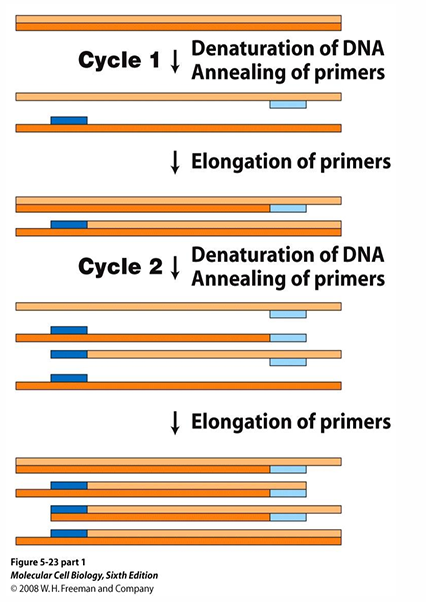
How is PCR different to cellular replication?
Replaces helicase with heat, primase with synthetic DNA primers, and DNA Polymerase III with thermostable Taq polymerase
What are the four simple take home messages from DNA replication?
DNA polymerases cannot melt or unwind duplex DNA
DNA polymerases cannot initiate chains, but can only extend a pre-existing DNA or RNA strand
DNA replication can only start at an origin of replication
All DNA strands grow in a 5’ ==> 3’ direction
What are the 3 types of RNA? Describe each function briefly
mRNA: Carries the coding message
tRNA: Delivers AAs
rRNA: Structural/catalytic core of ribosomes
What are the different types of polymerases that transcribe the different classes of RNA (what are the RNA’s functions as well)?
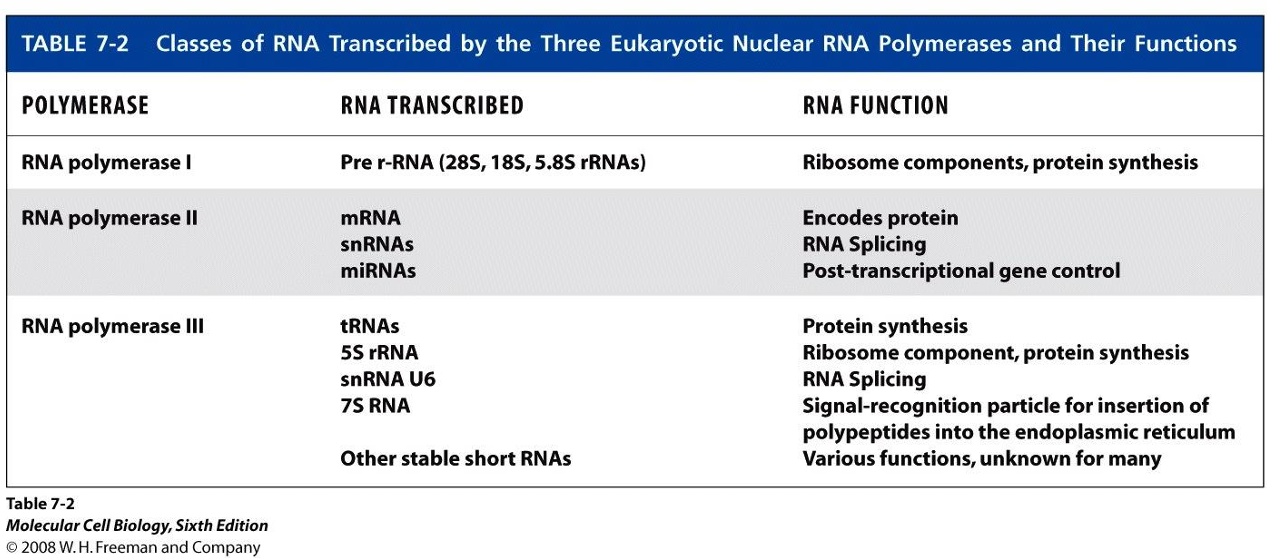

What reads the template strand in transcription and in which direction?
RNA polymerase
3’ ==> 5’
Describe the coding strand, and which way does transcription proceed?
Has the same sequence as the mRNA (with T instead of U)
Transcription proceeds 5’ ==> 3’, reading the template 3’ ==> 5’
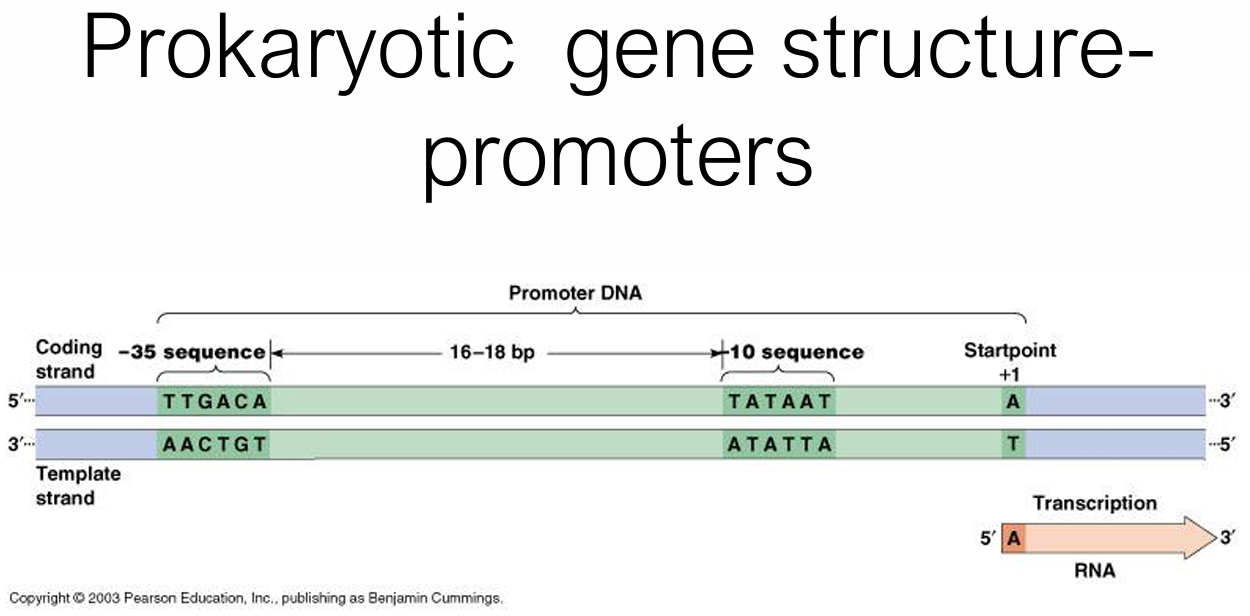
What are the promoter regions like in prokaryotic organisms?
−35 box: TTGACA (σ factor recognition)
−10 box: TATAAT (Pribnow box, melting site = rich in weaker AT)
16–18 bp spacer between them
Transcription starts at +1
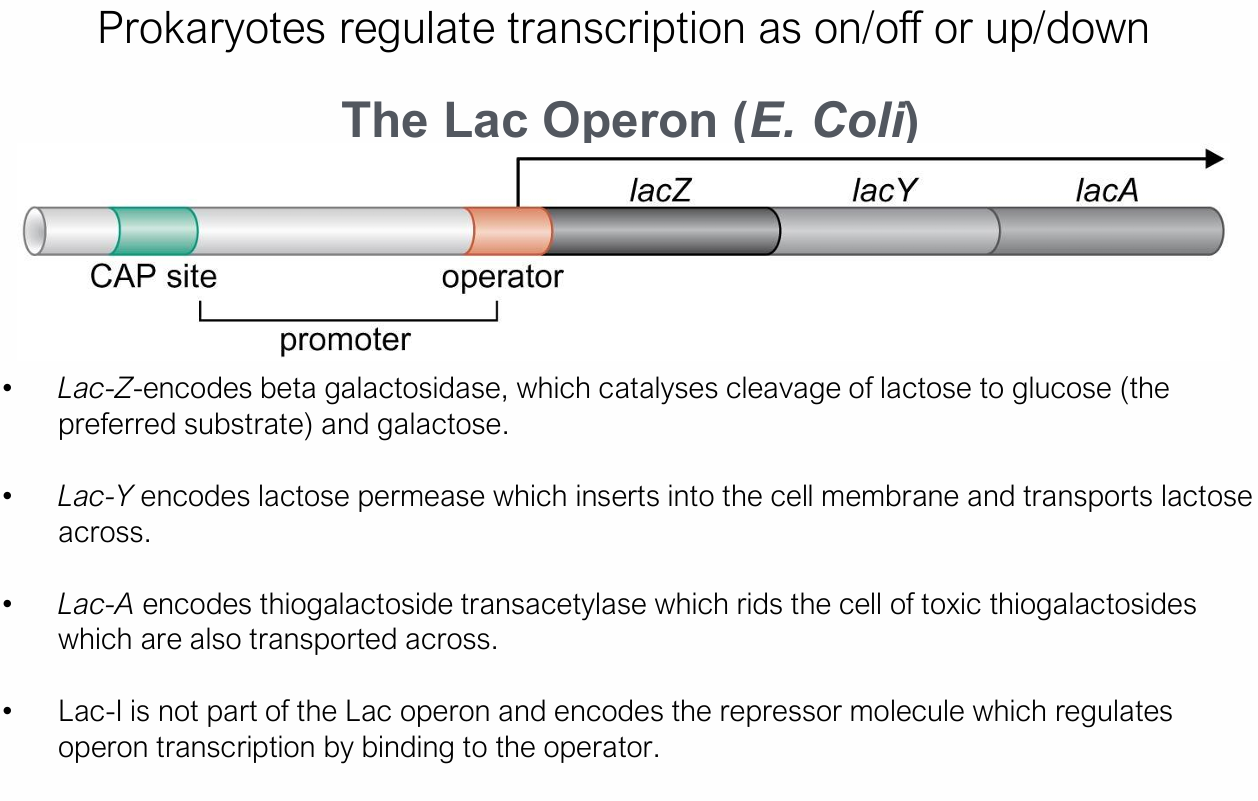
What are the promoter regions like in eukaryotic organisms?
TATA box (~−25): TATAAA, bound by TFIID
CAAT box (~−80): GCCCAATCT, bound by CTF
GC box (~−100): GGGCGG, bound by Sp1
Inr element at +1
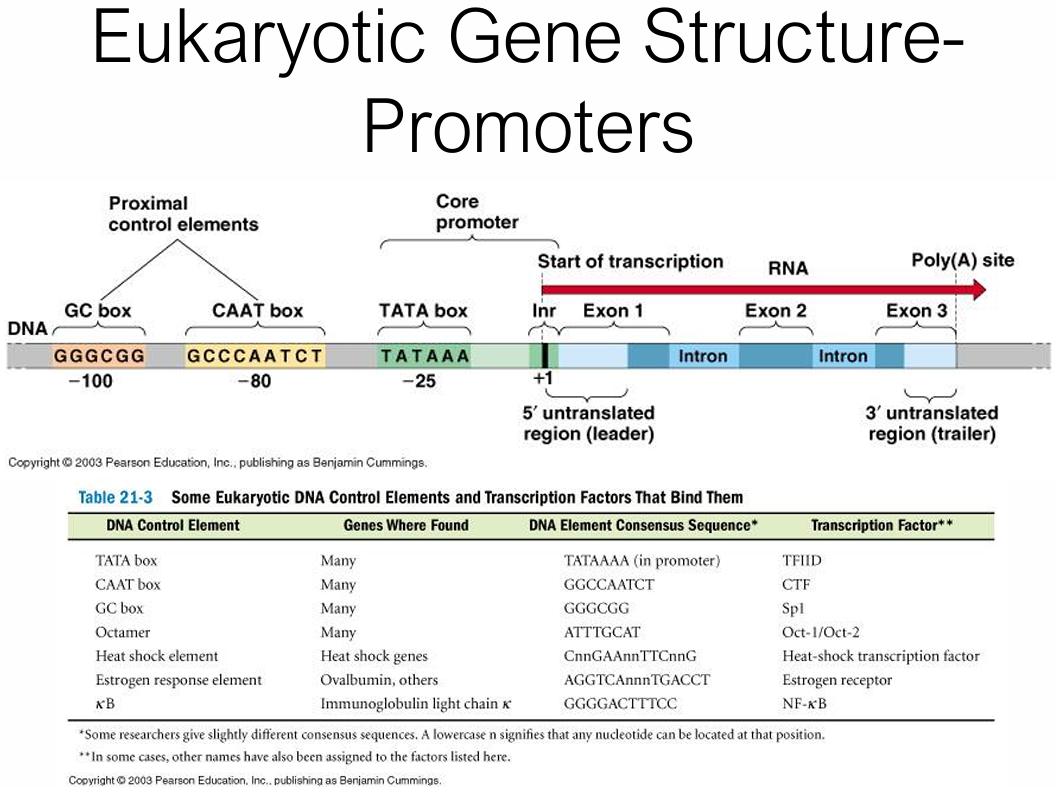
What is the function of prokaryotic operons?
A single promoter controls transcription of multiple genes into one polycistronic mRNA (e.g. the trp operon has genes E, D, C, B, A)
Allowing for the synthesis of several proteins from one mRNA molecule
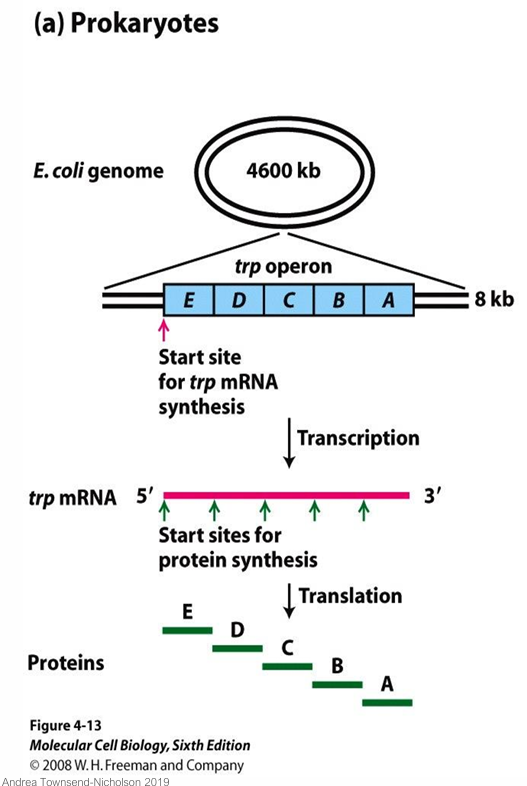
What is transcriptional regulation like in eukaryotic genes?
Each gene has its own promoter and produces a monocistronic mRNA; related genes (e.g. TRP1–5 in yeast) are scattered across different chromosomes.
Single proteins
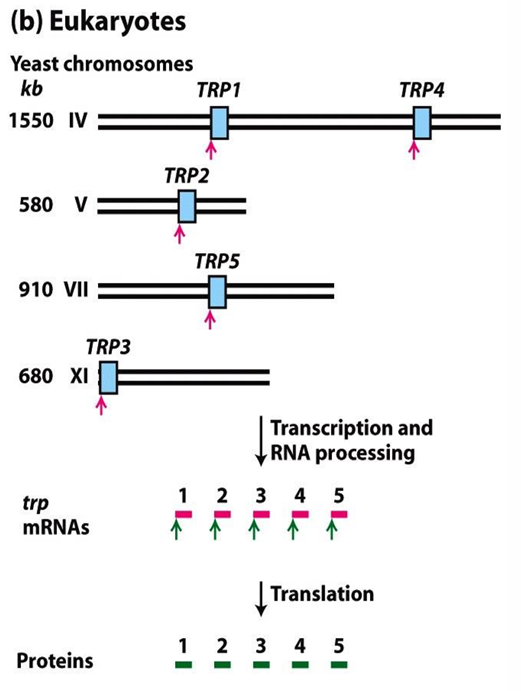
What is The Lac Operon in E. coli for?
Model of prokaryotic gene/transcription regulation, controlled by both a repressor (negative control) and CAP/cAMP (positive control).
Lac-Z = Encodes β-galactosidase
Catalyses cleavage of lactose to glucose (the preferred substrate) and galactose
Lac-Y = Encodes lactose permease
Inserts into the cell membrane and transports lactose across
Lac-A = Encodes thiogalactoside transacetylase
Rids the cell of toxic thogalactosides, also transported across
Lac-I = Encodes the repressor molecule
Regulates operon transcription by binding to the operator
What are the 3 regulatory states of The Lac Operon? In terms of charge states of glucose and lactose, cAMP, repressor, CAP, and transcription
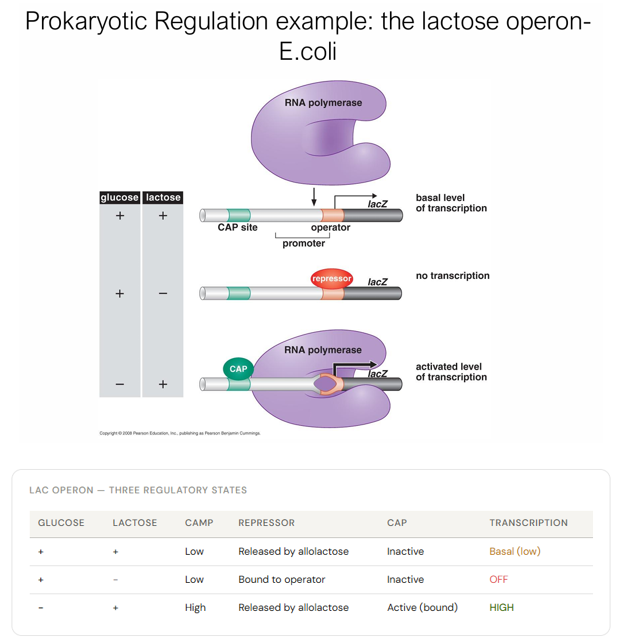
What is the molecular mechanism/perspective of The Lac Operon?
Lactose spontaneously isomerises to allolactose
So that allolactose is present at a low level when lactose is present
Allolactose binds the Lac repressor
Causing a conformational change that prevents it from binding the operator
RNA polymerase can now transcribe
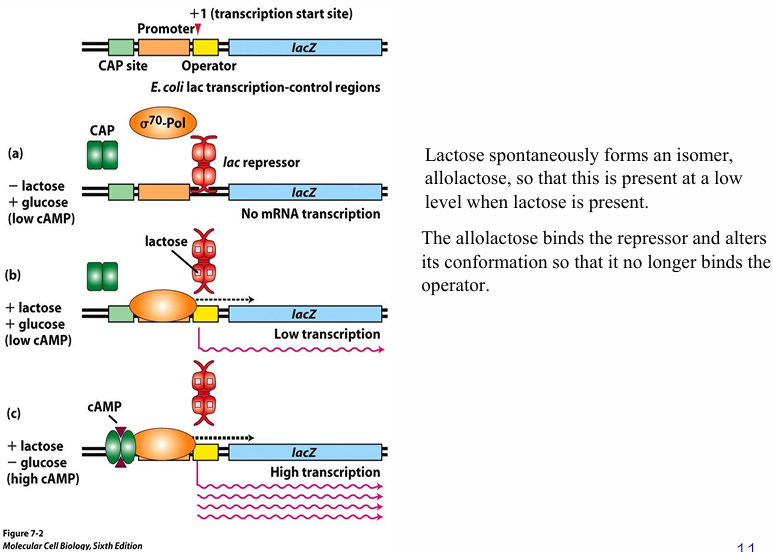
What happens in the absence of glucose, in the molecular mechanism/perspective of The Lac Operon
cAMP rises, binds CAP, and CAP–cAMP binds the CAP site upstream of the promoter → activates transcription to high levels.
What is is eukaryotic gene regulation controlled by? Give examples
General transcription factors (GTFs)
TFIIA, TFIIB, TFIID, TFIIE, TFIIF, TFIIH
What is the function of general transcription factors (GTFs), and what do they resolve?
Eukaryotic RNA Pol II cannot bind the promoter alone
A set of GTFs assemble at the promoter in a specific order to form the preinitiation complex
What is the process of eukaryotic gene regulation up until transcription begins?
TFIID binds TATA box first → TFIIA + TFIIB stabilise → TFIIF + RNA Pol II join → TFIIE + TFIIH complete the complex → TFIIH phosphorylates RNA Pol II CTD (carboxy-terminal domain) → transcription begins
What are activators and what is its role in eukaryotic gene regulation?
DNA-binding proteins
Bind to enhancer DNA sequences (can be thousands of base pairs away from the promoter)
e.g. enhanceosome
DNA looping brings activators close to the core promoter, where they recruit coactivators that bridge the enhanceosome to TFIID, boosting transcription above basal levels
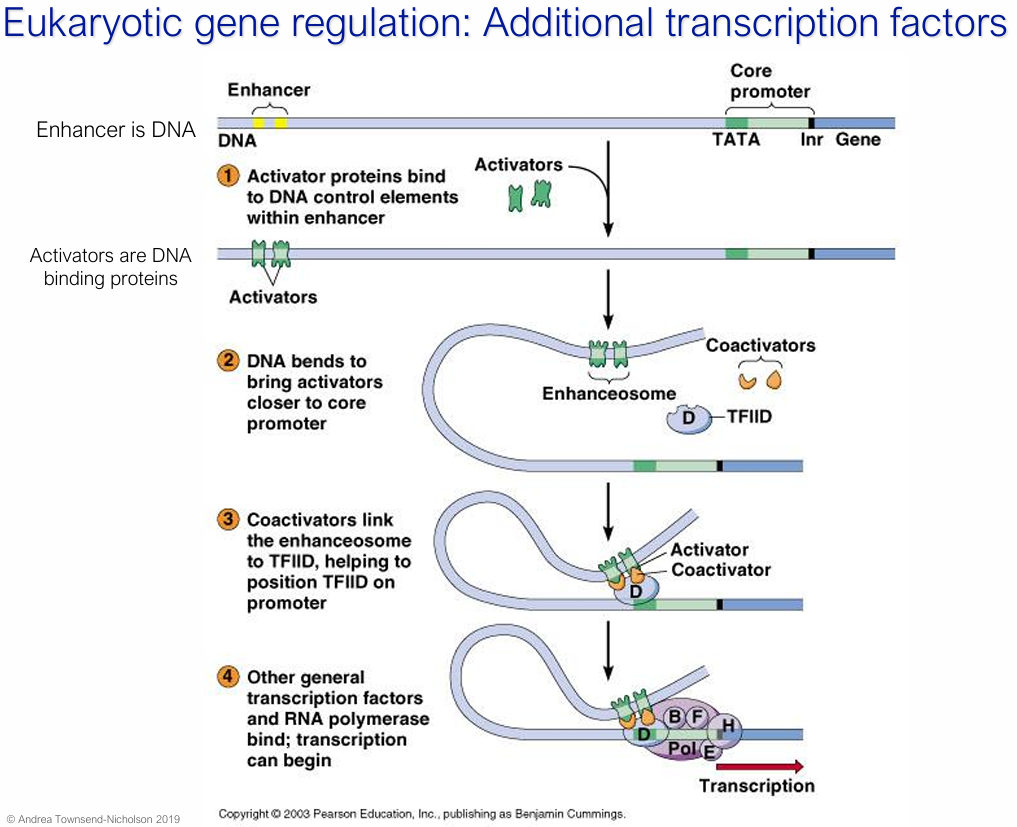
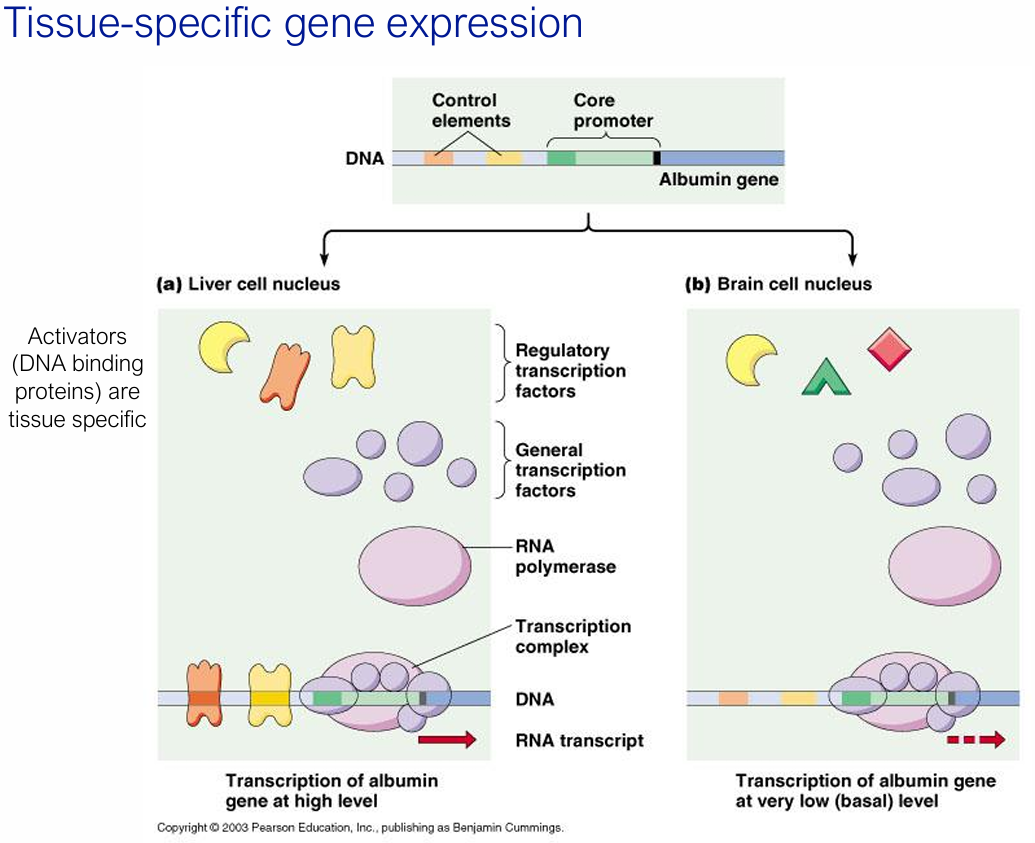
How/Why does tissue-specific gene expression arise?
Because different cell types express different sets of regulatory transcription factors that bind to control elements
For e.g. the albumin gene is transcribed at high levels in liver cells (which express the right activators) but only at basal levels in brain cells.
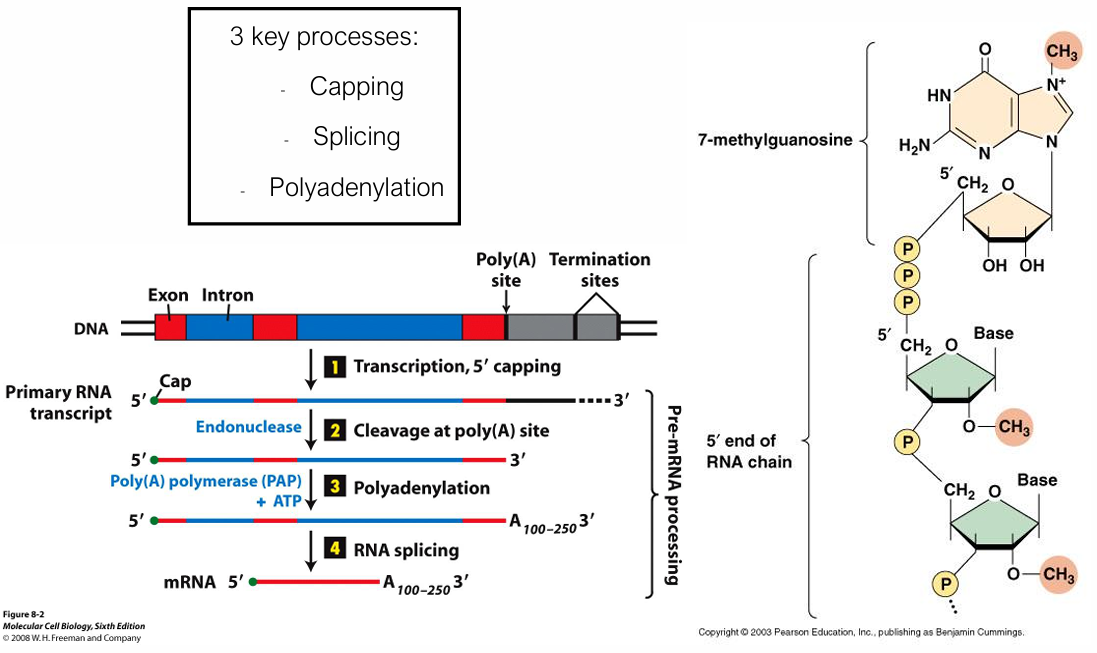
What are the 3 key processes in pre-mRNA processing before exporting from the nucleus? Describe them briefly
Transcription 5’ Capping
A 7-methylguanosine (m⁷G) cap is added via a 5′→5′ triphosphate linkage.
Functions: protects from exonucleases, aids ribosome recognition, promotes nuclear export
Polyadenylation
An endonuclease cleaves at the poly(A) signal site; poly(A) polymerase (PAP) then adds 100–250 adenine residues. REQUIRES ATP
poly(A) tail promotes mRNA stability and translation.
RNA splicing
Introns are removed (by endonucleases) and exons joined by the spliceosome (snRNPs + associated proteins).
This produces the mature mRNA containing only the coding sequence.
What is alternative splicing? Give example
A single gene can produce different proteins in different tissues via alternative splicing
Calcitonin (CT) and Calcitonin gene-related peptide (CGRP)
CT/CGRP Gene
Two peptide hormones from same gene
6 exons
Exons 1-3 constitutive, 4 used in thyroid cells, 5 & 6 used in brain tissue
Thyroid CT: Expressed in thyroid cells + Regulates blood calcium
Exons 1-4, AAAn
Brain CGRP: Expressed in neurons + potent vasodilator, associated with and released during migraine
Exons 1-3, 5, 6, AAAn
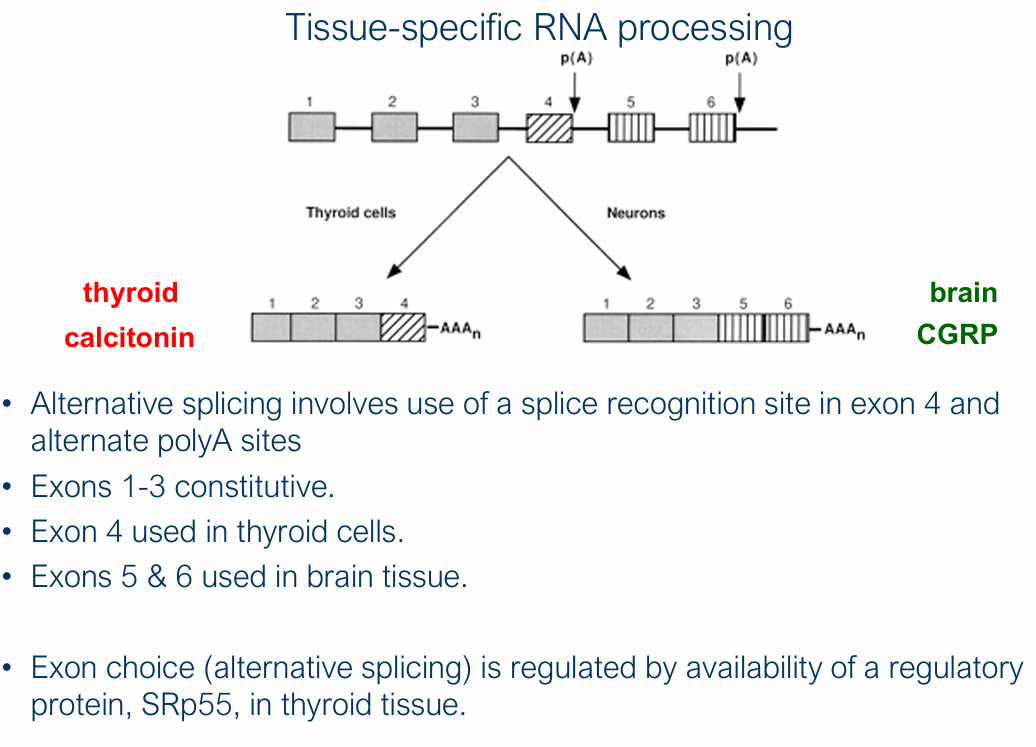
What exon choice/alternative splicing regulated by?
Availability of a regulatory protein, SRp55, in thyroid tissue
What is the clinical significance of alternative splicing?
Up to 40% of genetic defects are linked to incorrect RNA processing. At least 45–60% of human genes undergo alternative splicing
What are the five take home messages from transcription?
Transcription of protein encoding genes is regulated at different levels in both prokaryotes and eukaryotes:
Initiation of basal level of transcription (GTFs in eukaryotes, operons in prokaryotes).
Enhancement of basal transcription (activators in both prokaryotes and eukaryotes).
Tissue specificity (eukaryotes only-presence of activators limited to certain tissues).
Processing of messenger RNA (eukaryotes-pre mRNA to mature mRNA through 5’cap, 3’ poly A tail and removal of introns).
What kind of complexes are ribosomes, and its catalytic activity is provided by what?
Ribonucleoprotein
rRNA (making the ribosome a ribozyme)
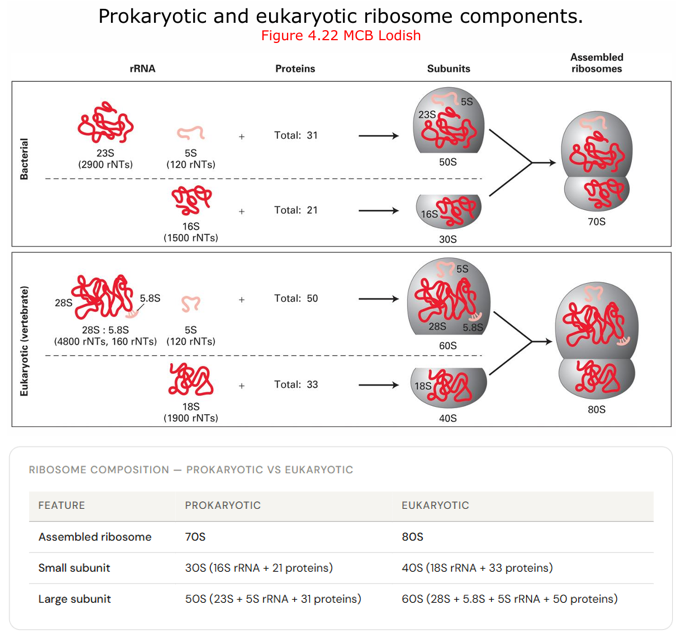
Describe ribosome composition, prokaryote and eukaryotic
What are the 3 tRNA binding sites in ribosomes?
A site (Aminoacyl)
Incoming charged tRNA binds here
P site (Peptidyl)
Holds tRNA carrying the growing polypeptide chain
E site (Exit)
Deacylated tRNA exits from here
What is the common first step in initiation in both prokaryotes and eukaryotes?
Translation always begins at initiator methionine (AUG)
Special initiator tRNA recognises the start codon
(distinct from the regular Met-tRNA used during elongation)
Synthesis of all protein chains in prokaryotic and eukaryotic cells begins with the amino acid Methionine (Met, or M) (f Met in prokaryotes)
“Methionine residues are also present within the protein”. How does the ribosome know which methionine is the initiator methionine?
There are two tRNAs for methionine-one is the Met initiator tRNA and the other is the ‘regular’ Met tRNA. Only the initiator tRNA recognises and binds to the AUG in the ribosome binding site. This is important to ensure that the correct reading frame is used
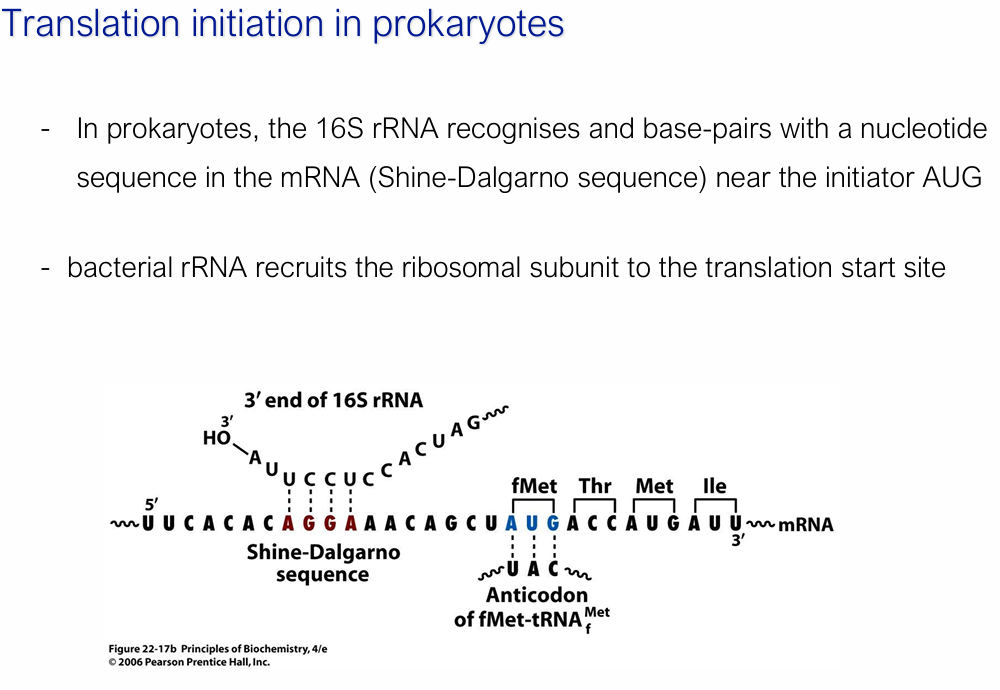
Describe prokaryotic initiation of translation
16S rRNA base-pairs with the Shine-Dalgarno sequence = AGGA (purine-rich, upstream of AUG)
Bacterial rRNA recruits the ribosomal subunit to the translation start site
Start amino acid: fMet (formyl-methionine)
Initiation factors: IF1, IF2, IF3
IF-1: helps in the binding of the ribosome to the mRNA.
IF-2: is responsible for the recruitment of the initiator tRNA.
IF-3: prevents the premature association of the ribosomal subunits
==> IFs essential for forming the initiation complex
Describe eukaryotic initiation of translation
Initiation factors: eIF1–eIF6
Start amino acid: Met (no formyl group)
40S subunit (already loaded with initiator met-tRNAi + GTP + eIF2 = TERNARY COMPLEX) binds near the 5′ cap
Initiator tRNA moves along mRNA from 5’ to 3’ scanning for first AUG = REQUIRES ATP HYDROLYSIS
(Marilyn) Kozak sequence: 5′-GCC(G/A)CCAUGG-3′ = Where translation starts
Once the anticodon of the initiator tRNA base-pairs with the first AUG it encounters, scanning stops. GTP is hydrolysed to GDP (by eIF2), which triggers release of initiation factors and allows the large (60S) subunit to join — and elongation can begin.
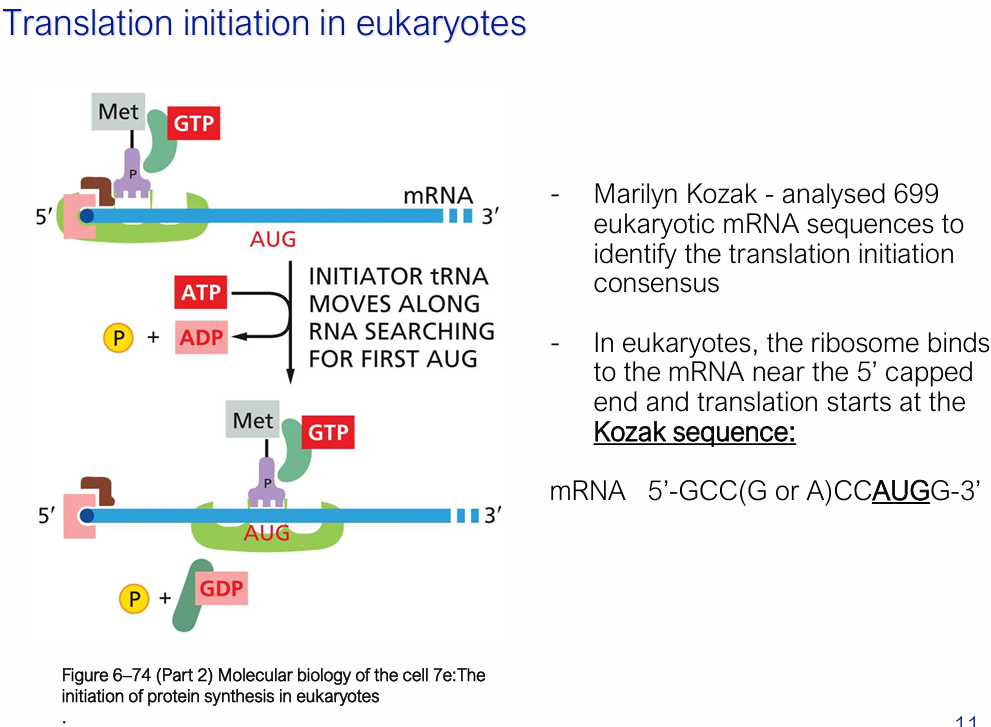
What is the ternary complex in eukaryotic initiation of translation
Met-tRNAᵢ + eIF2 + GTP
Elongation in translation tends to proceed at roughly what rate?
3-5 AAs/second
Describe elongation in translation
The pp chain is attached to the P-site tRNA
Charged aminoacyl-tRNA enters the A site (codon–anticodon matching + by elongation factor EF-Tu + GTP)
GTP hydrolysis acts as a proofreading check before the tRNA is locked in
Peptide bond formation (peptidyl transfer): the growing chain transfers from P-site tRNA to A-site (peptidyl transferase = a ribozyme — made of rRNA, activity of 23S/28S rRNA)
The P-site tRNA is now 'deacylated' (empty)
Translocation: ribosome shifts one codon in the 5′ ==> 3′ direction — A-site tRNA moves to P, P-site tRNA moves to E
E-site tRNA exits, will be recycled — picked up by aminoacyl-tRNA
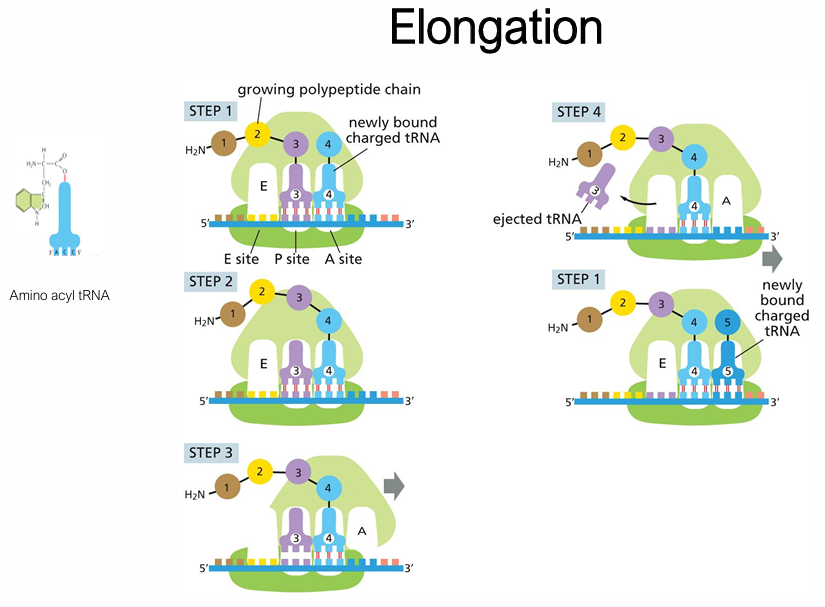
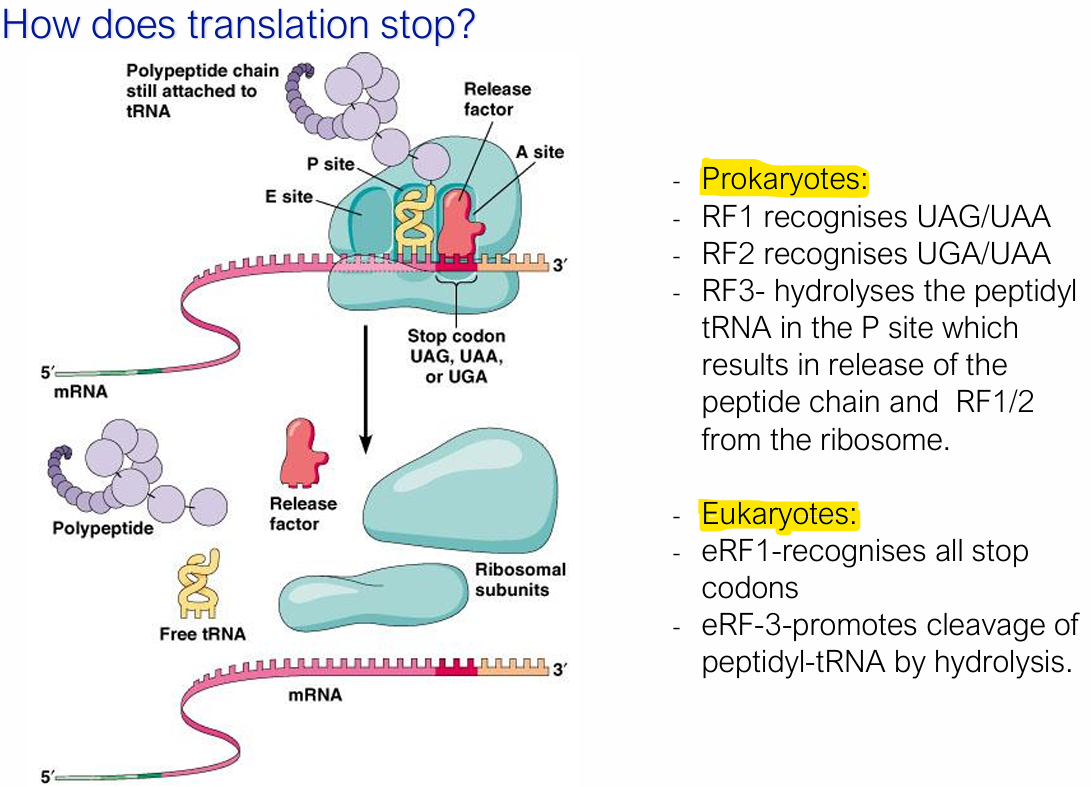
Generally, how does termination occur in translation?
When a stop codon (UAA, UAG, or UGA), is reached/ enters the A site.
No tRNA recognises stop codons — instead, release factors (proteins) recognise the different stop codons, bind, and promote cleavage of the peptidyl-tRNA
What are the different release factors for termination of translation in prokaryotes and eukaryotes?
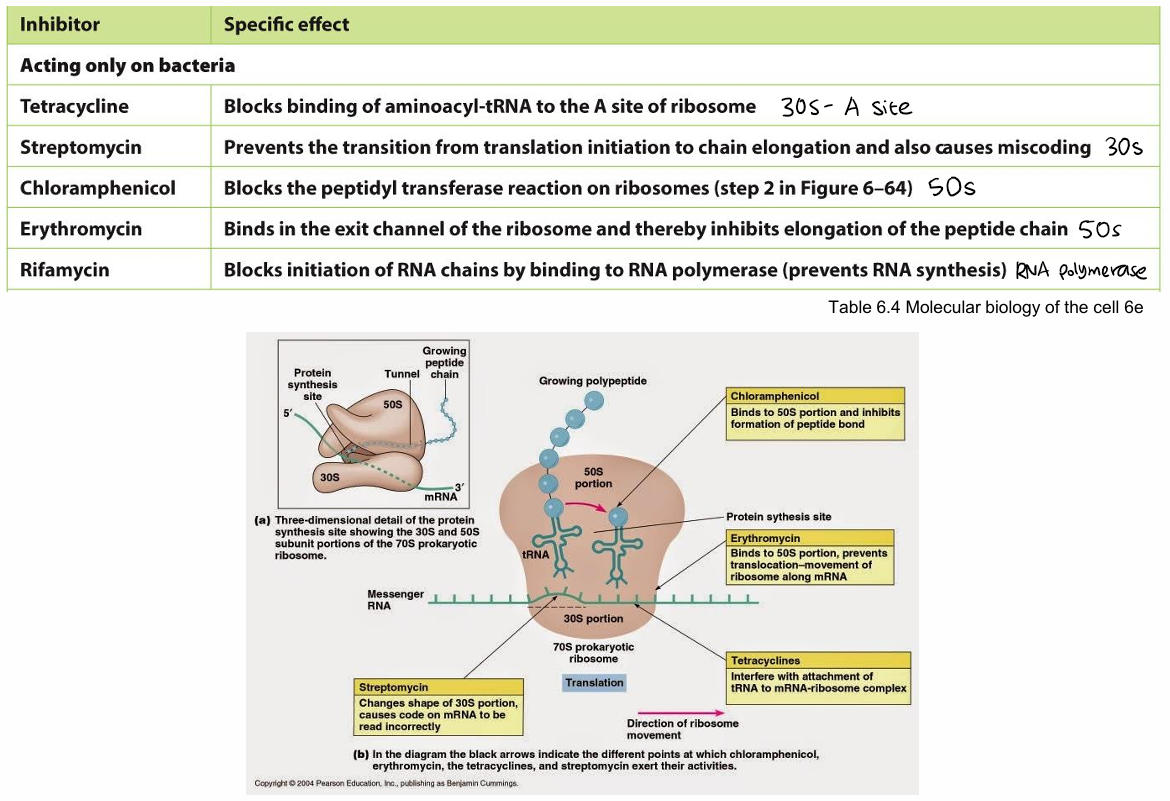
Name the antibiotics that affect protein synthesis and its specific effects/
What is the link between circular mRNA and Poly(A) tail?
In eukaryotes, PABP proteins on the poly(A) tail interact with eIF4G at the 5′ cap, circularising the mRNA.
This promotes efficient ribosome recycling and re-initiation.
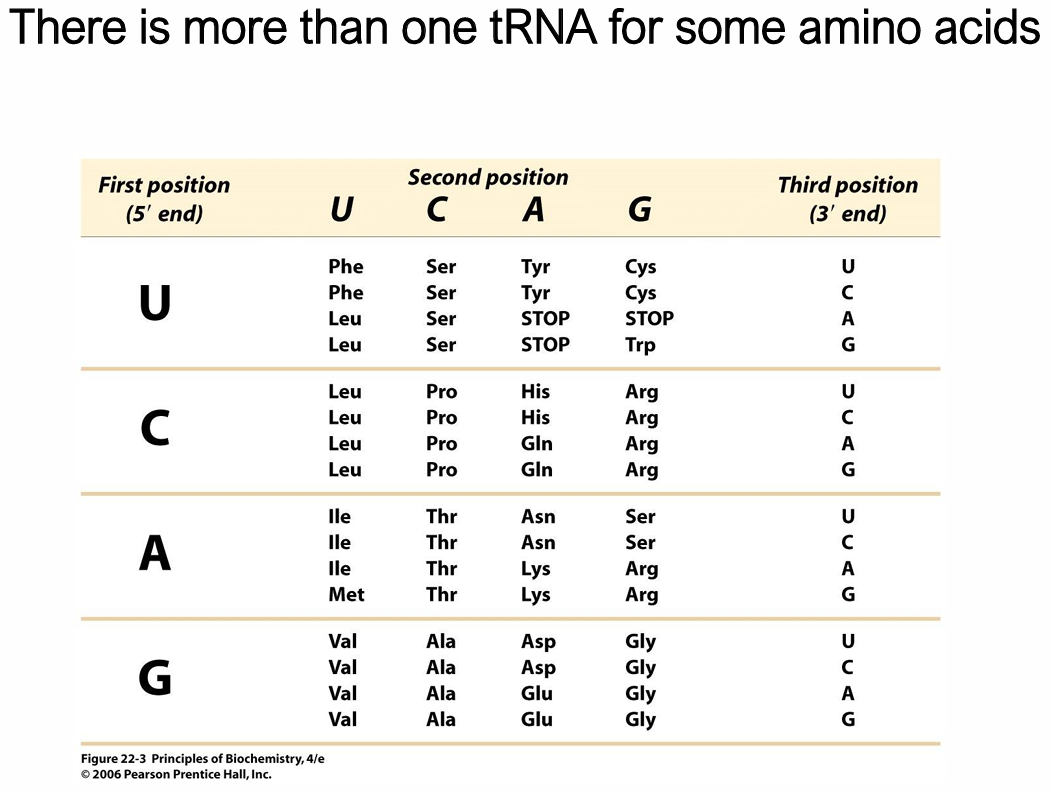
What is The Genetic Code?
Triplet code
4³ = 64 possible codons
Of these, 61 encode AA
20 AAs used in proteins and encoded directly in DNA
3 are stop signals
Degenerate
Most AAs are specified by more than one codon
Universal across life
Human mRNA can be correctly translated by bacterial ribosomes
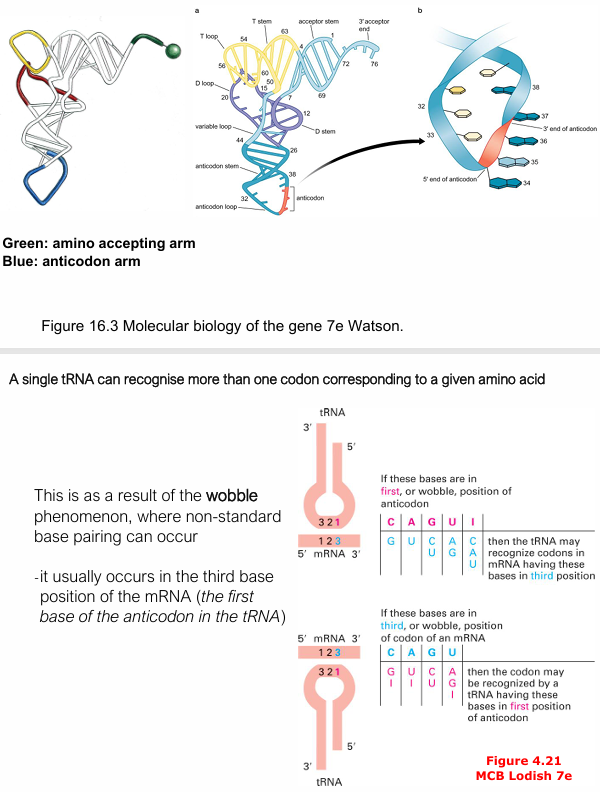
What is wobble hypothesis (crick)?
Humans have only ~48 tRNA anticodons for 61 sense codons. The wobble position is the 3rd base of the codon (= 1st base of the anticodon), where non-standard base pairing is tolerated — e.g. G can pair with U, and the modified base inosine (I) can pair with U, C, or A.
This allows one tRNA to recognise multiple synonymous codons.
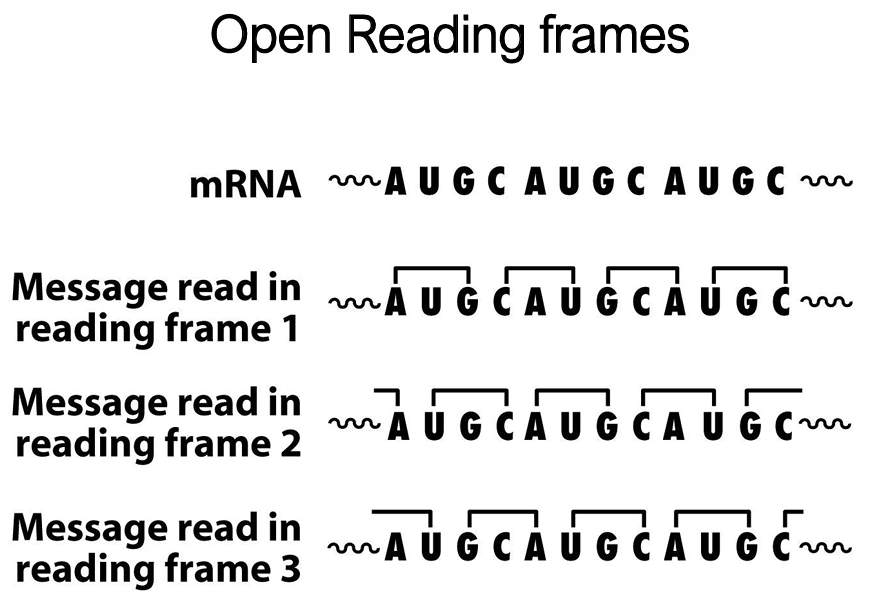
What are reading frames?
The same mRNA sequence can be read in three possible reading frames depending on the start position.
The correct reading frame is set by the initiator AUG — this is why the Shine-Dalgarno / Kozak sequence and initiator tRNA are so important
How do fewer than 61 tRNAs recognise the 61 codons?
What are the five take home messages from translation?
Ribosomes are large ribonucleoproteins comprised of two subunits containing several different rRNAs and more than 50 proteins
The catalytic activity is provided by the large RNA subunit(23s/28s)
As with transcription, there is an initiation, elongation and termination phase. Each of these is highly regulated by initiation/elongation/termination factors (regulatory proteins).
They read the genetic code from mRNA in triplet codons.
There is degeneracy in the genetic code meaning that multiple codons can encode the same amino acid, and multiple tRNAs can read one codon.