High-Yield DAT Biology Notes
1/897
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
898 Terms
Matter
anything that takes up space and has mass
Element
a pure substance that has specific physical/chemical properties and can't be broken down into a simpler substance
Atom
the smallest unit of matter that still retains the chemical properties of the element
Molecule
two or more atoms joined together
Intramolecular forces
attractive forces that act on atoms within a molecule
Intermolecular forces
forces that exist between molecules and affect physical properties of the substance
Monomers
single molecules that can potentially polyermize
Polymers
substances made up of many monomers joined together in chains
Carbohydrates
- Contain carbon, hydrogen, and oxygen atoms (CHO)
Ex: Monosaccharides, disaccharides, polysaccharides
Monosaccharides
carbohydrate monomers
- empirical formula: (CH2O)n.
- "n" represents number of carbons
Ribose
five carbon monosaccharide
Fructose and Glucose
six carbon monosaccharide
- isomers of each other (same chemical formula, different arrangement of atoms)
Disaccharides
two monosaccharides joined together by a glycosidic bond
- Result of dehydration (condensation) reaction
Dehydration (condensation) reaction
where a water molecule leaves and a covalent bond forms
Hydrolysis reaction
a covalent bond is broken by the addition of water
Sucrose
disaccharide made of glucose + fructose
Lactose
disaccharide made of galactose + glucose
Maltose
disaccharide made of glucose + glucose
Polysaccharides
contain multiple monosaccharides connected by glycosidic bonds to form long polymers
Starch
form of energy storage for plants and is an alpha (α) bonded polysaccharide
- Linear starch: amylose
- Branched form: amylopectin
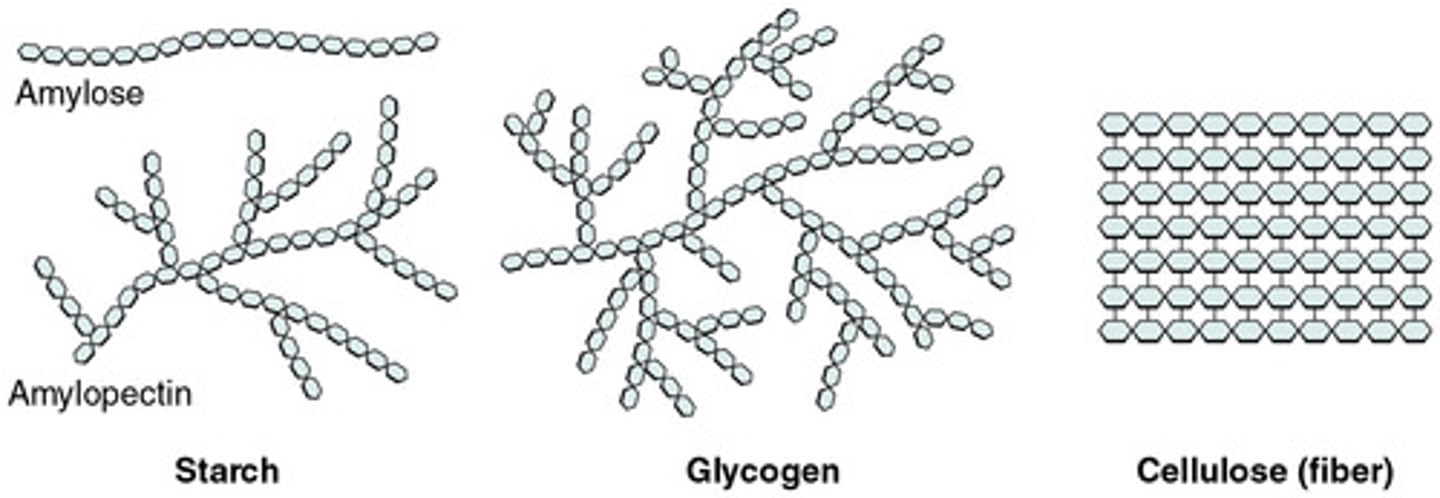
Glycogen
form of energy storage in animals and is an alpha (α) bonded polysaccharide
- much more branching than starch
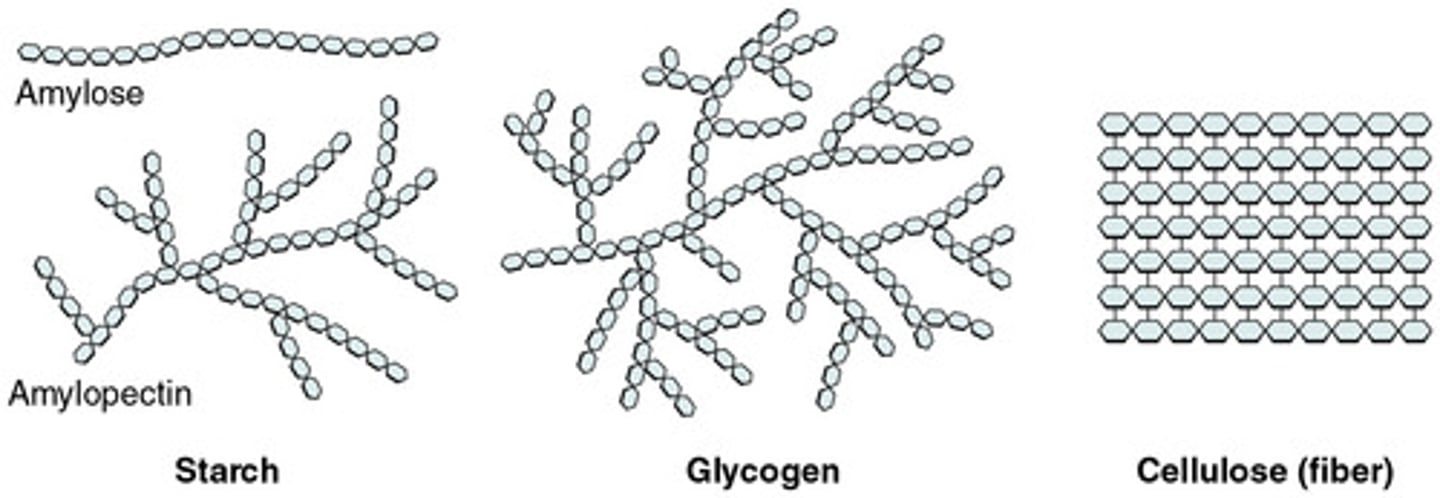
Cellulose
structural component in plant cell walls, and is a beta (β) bonded polysaccharide
- Linear strands packed rigidly in parallel
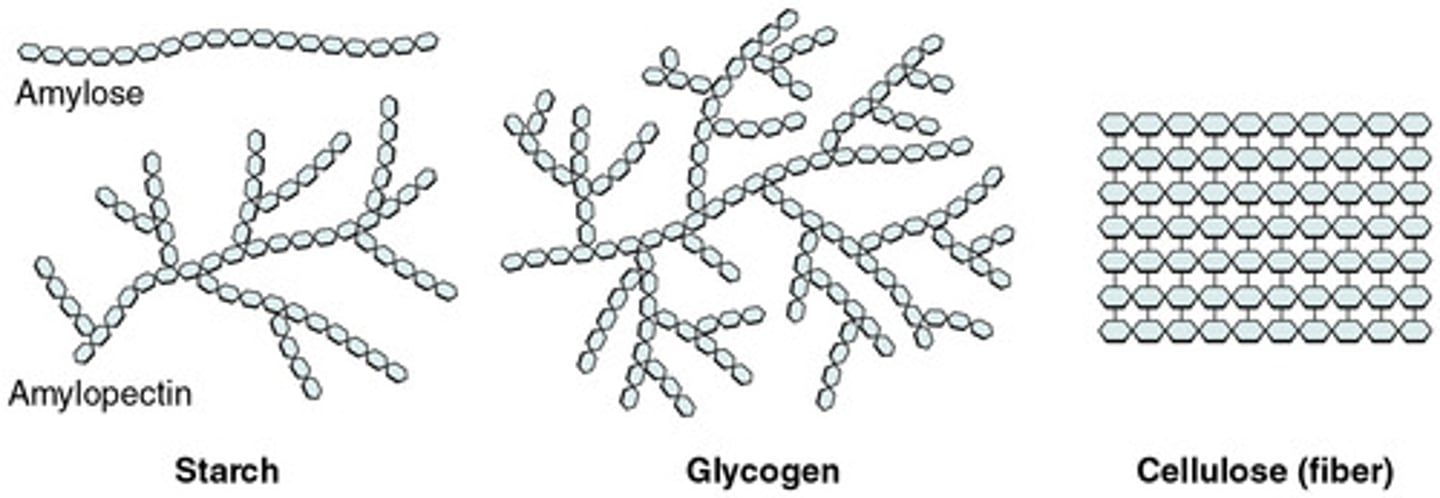
Chitin
structural component in fungi cell walls and insect exoskeletons, and is a beta (β) bonded polysaccharide with nitrogen added to each monomer
Proteins
contain carbon, hydrogen, oxygen, and nitrogen atoms (CHON)
- atoms combine to form amino acids, which link together to build polypeptides (or proteins)
Proteome
all the proteins expressed by one type of cell under one set of conditions
Amino acids
monomers of proteins
- twenty different kinds of amino acids, each with a different "R-group"
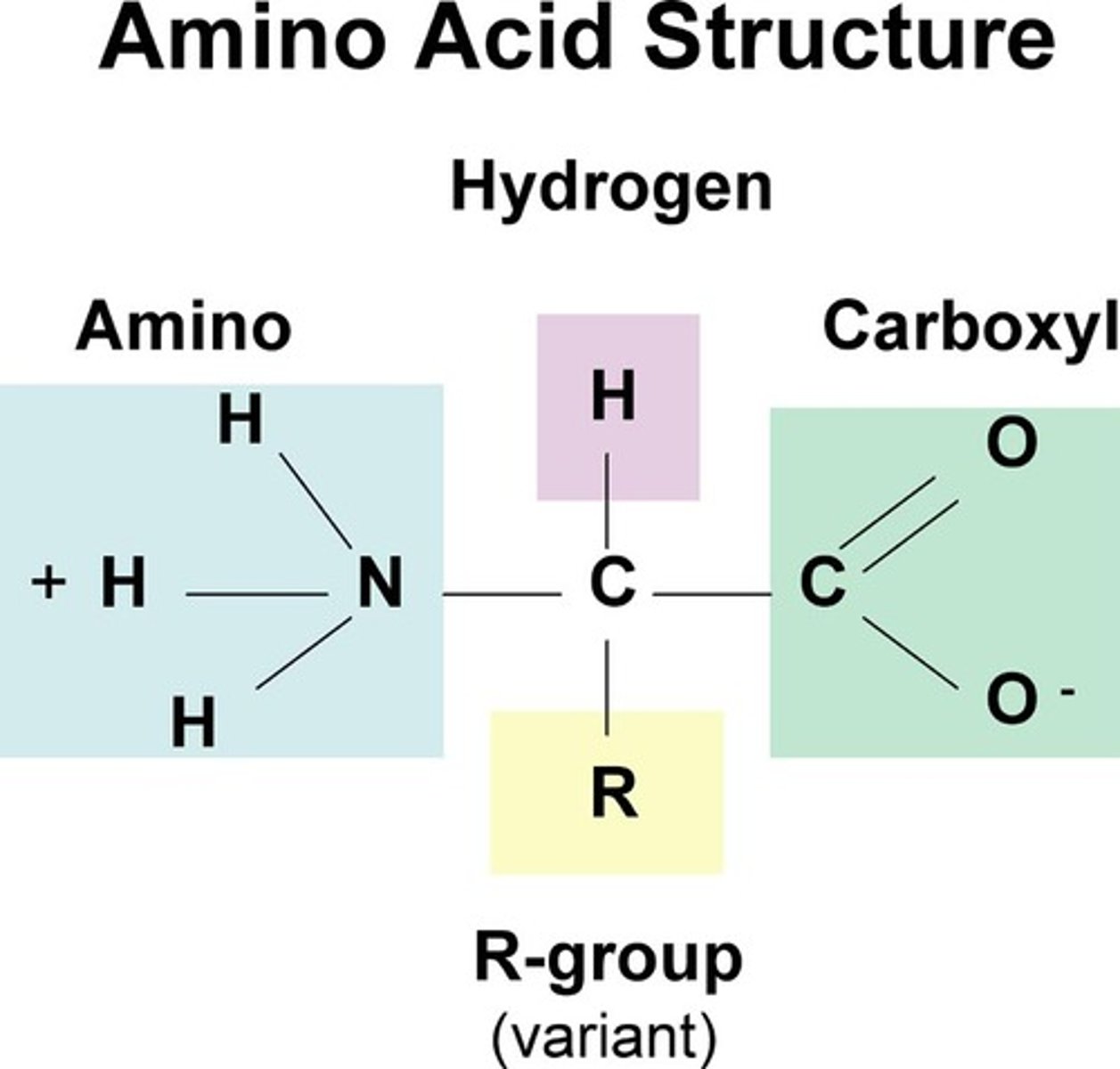
Polypeptides
polymers of amino acids and are joined together by peptide bonds through dehydration reactions
- hydrolysis reactions break peptide bonds
- becomes an amino acid chain that contains two end terminals on opposite sides
N-terminus (amino terminus)
the side that ends with the last amino acid's amino group of a polypeptide
C-terminus (carboxyl terminus)
the side that ends with the last amino acid's carboxyl group of a polypeptide
Conjugated proteins
proteins that are composed of amino acids and non-protein components
- Metalloproteins (ex: hemoglobin)
- Glycoprotein (ex: mucin)
Metalloproteins
proteins that contain a metal ion cofactor
Ex: hemoglobin
Glycoprotein
proteins that contain a carbohydrate group
Ex: mucin
Primary structure
sequence of amino acids connected through peptide bonds
Secondary structure
intermolecular forces between the polypeptide backbone (not R-groups_ due to hydrogen bonding
- Forms α-helices or β-pleated sheets.
Tertiary structure
three dimensional structure due to interactions between R-groups
- create hydrophobic interactions based on the R-groups
- disulfide bonds are created by covalent bonding between the R-groups of two cysteine amino acids
- hydrogen bonding and ionic bonding between R groups also hold together the tertiary structure
Quaternary structure
multiple polypeptide chains come together to form one protein
Protein denaturation
describes the loss of protein function and higher order structures
- only the primary structure is unaffected
Proteins will denature as a result of...
high or low temperatures, pH changes, and salt concentrations
Storage (protein function)
reserve of amino acids
Hormones (protein function)
signaling molecules that regulate physiological processes
Receptors (protein function)
proteins in cell membranes which bind to signal molecules
Structure (protein function)
provide strength and support to tissues (hair, spider silk)
Immunity (protein function)
antibodies that protect against foreign substances
Enzymes (protein function)
regulate rate of chemical reactions
- act as biological catalysts by binding to substrates (reactants) and converting them into products
- most enzymes are proteins
Catalysts
increase reaction rates by lowering the activation energy of a reaction
- reduce the energy of a transition state
- does not shift a chemical reaction or affect spontaneity
Transition state
the unstable conformation between the reactants and the products
Active site
enzymes bind to substrates at an active site, which is specific for the substrate that it acts upon
Specificity constant
measures how efficient an enzyme is at binding to the substrate and converting it to a product
Induced fit theory
describes how the active site molds itself and changes shape to fit the substrate when it binds
- the "lock and key" model is an outdated theory of how substrates bind
Ribozyme
an RNA molecule that can act as an enzyme (a non-protein enzyme)
Cofactor
a non-protein molecule that helps enzymes perform reactions
Coenzyme
an organic cofactor (i.e. vitamins)
- inorganic cofactors are usually metal ions
Holoenzymes
enzymes that are bound to their cofactors
Apoenzymes
enzymes that are not bound to their cofactors
Prosthetic groups
cofactors that are tightly or covalently bonded to their enzymes
What are some ways enzymes catalyze reactions?
- Conformational changes that bring reactive groups closer
- The presence of acidic or basic groups
- Induced fit of the enzyme-substrate complex
- Electrostatic attractions between the enzyme and substrate
Phosphotase
cleaves a phosphate group off a substrate molecule
Phosphorylase
directly adds a phosphate group to a substrate molecule by breaking bonds within a substrate molecule
Kinase
indirectly adds a phosphate group to a substrate molecule by transferring a phosphate group from an ATP molecule
- these enzymes do not break bonds to add the phosphate group
Feedback regulation of enzymes
the end product of an enzyme-catalyzed reaction inhibits the enzyme's activity by binding to an allosteric site
Allosteric site
a location on an enzyme that is different from the active site
Competitive inhibition
occurs when a competitive inhibitor competes directly with the substrate for active site binding
- adding more substrate can increase enzyme action
- Km increases, while Vmax stays the same
Noncompetitive inhibition
occurs when the noncompetitive inhibitor binds to an allosteric site that modifies the active site
- the rate of enzyme action cannot be increased by adding more substrate
- Km stays the same, while Vmax decreases
Enzyme kinetics plot
used to visualize how inhibitors affect enzymes
- x-axis represents substrate concentration [X] while the y-axis represents reaction rate or velocity (V)
- Vmax is the maximum reaction velocity
- Michaelis Constant (Km) is the substrate concentration [X] at which velocity (V) is 50% of the maximum reaction velocity (Vmax)
- Saturation occurs when all active sites are occupied, so the rate of reaction does not increase anymore despite increasing substrate concentration (causes graph plateaus)
![<p>used to visualize how inhibitors affect enzymes</p><p>- x-axis represents substrate concentration [X] while the y-axis represents reaction rate or velocity (V)</p><p>- Vmax is the maximum reaction velocity</p><p>- Michaelis Constant (Km) is the substrate concentration [X] at which velocity (V) is 50% of the maximum reaction velocity (Vmax)</p><p>- Saturation occurs when all active sites are occupied, so the rate of reaction does not increase anymore despite increasing substrate concentration (causes graph plateaus)</p>](https://knowt-user-attachments.s3.amazonaws.com/5ad2498f-1a4b-46f3-b5ce-0b81e0890118.jpg)
Lipids
contain carbon, hydrogen, and oxygen atoms (CHO), like carbohydrates
- long hydrocarbon tails that make them very hydrophobic
Triacylglycerol (triglyceride)
a lipid molecule with a glycerol backbone (three carbons and three hydroxyl groups) and three fatty acids (long hydrocarbon tails)
- glycerol and the three fatty acids are connection by ester linkages
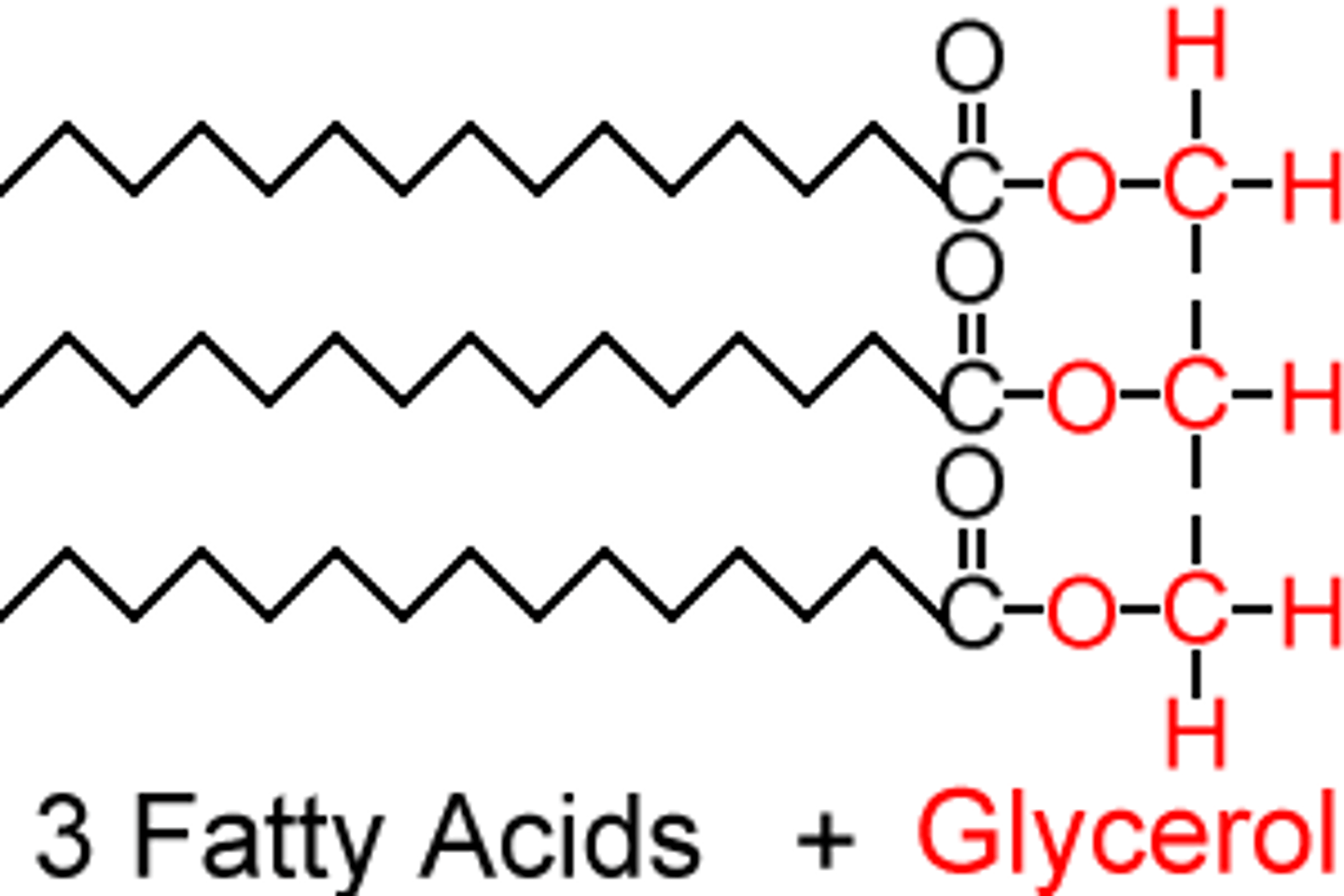
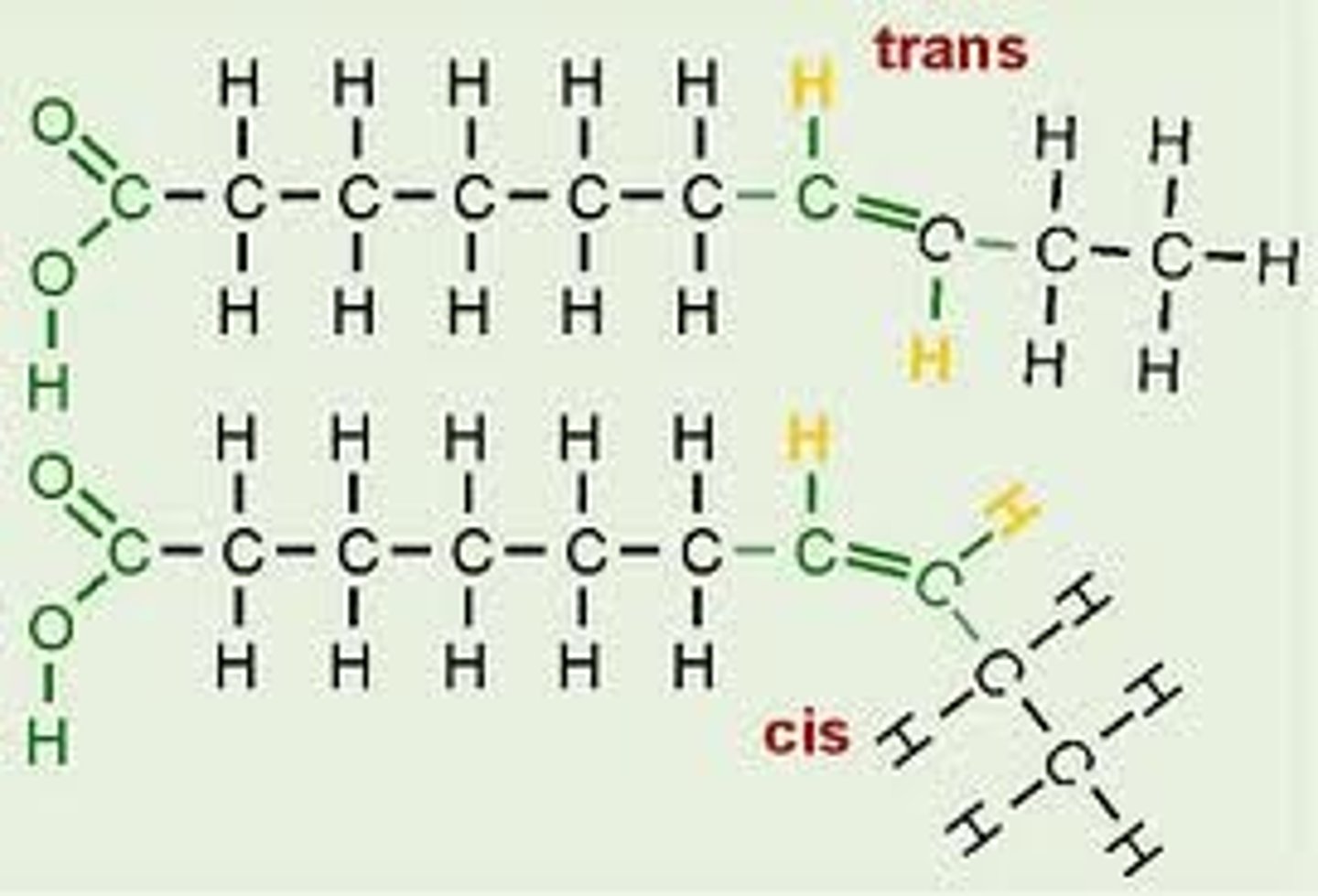
Saturated fatty acids
have no double bonds and as a result pack tightly (solid at room temperature)
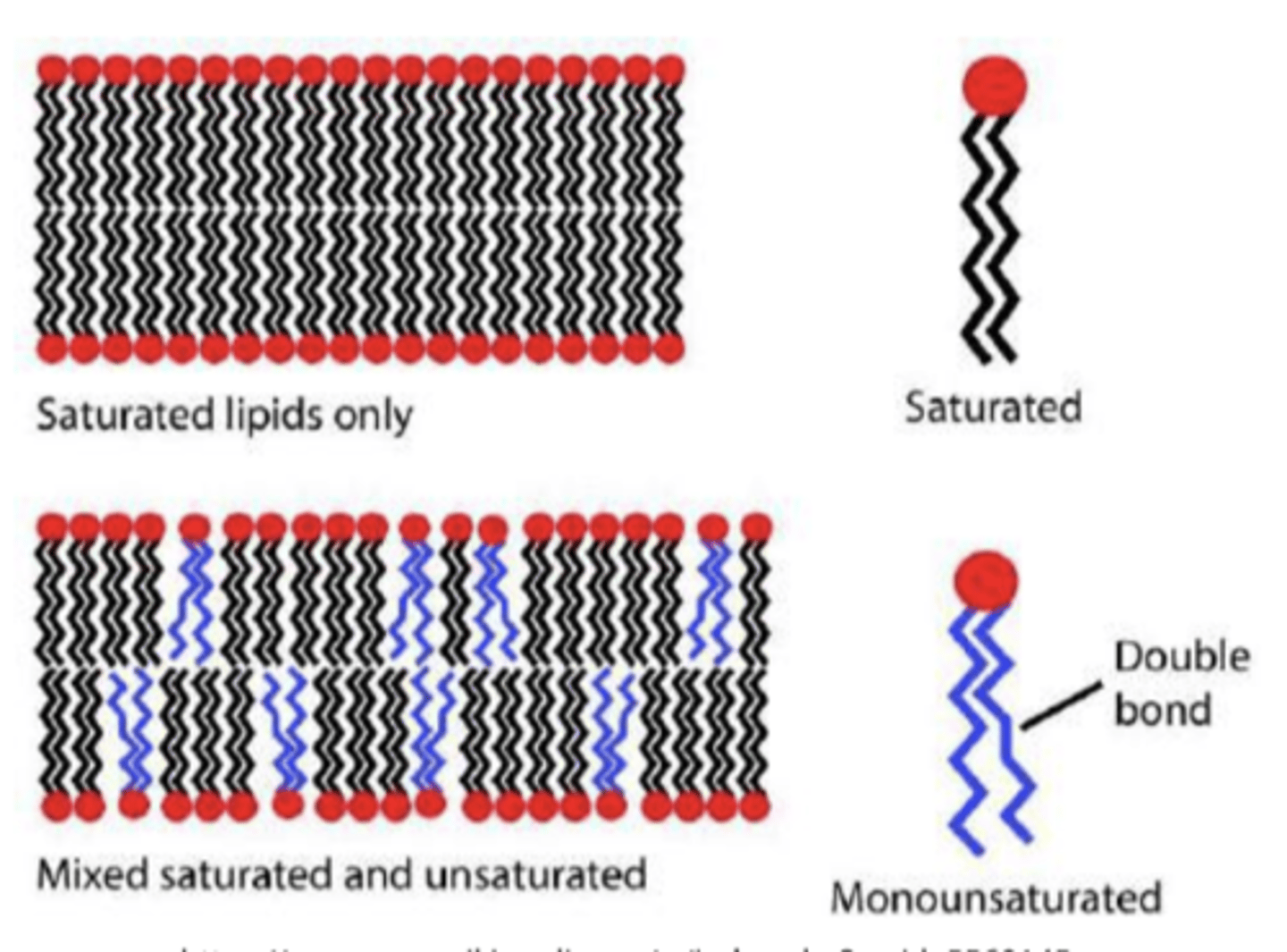
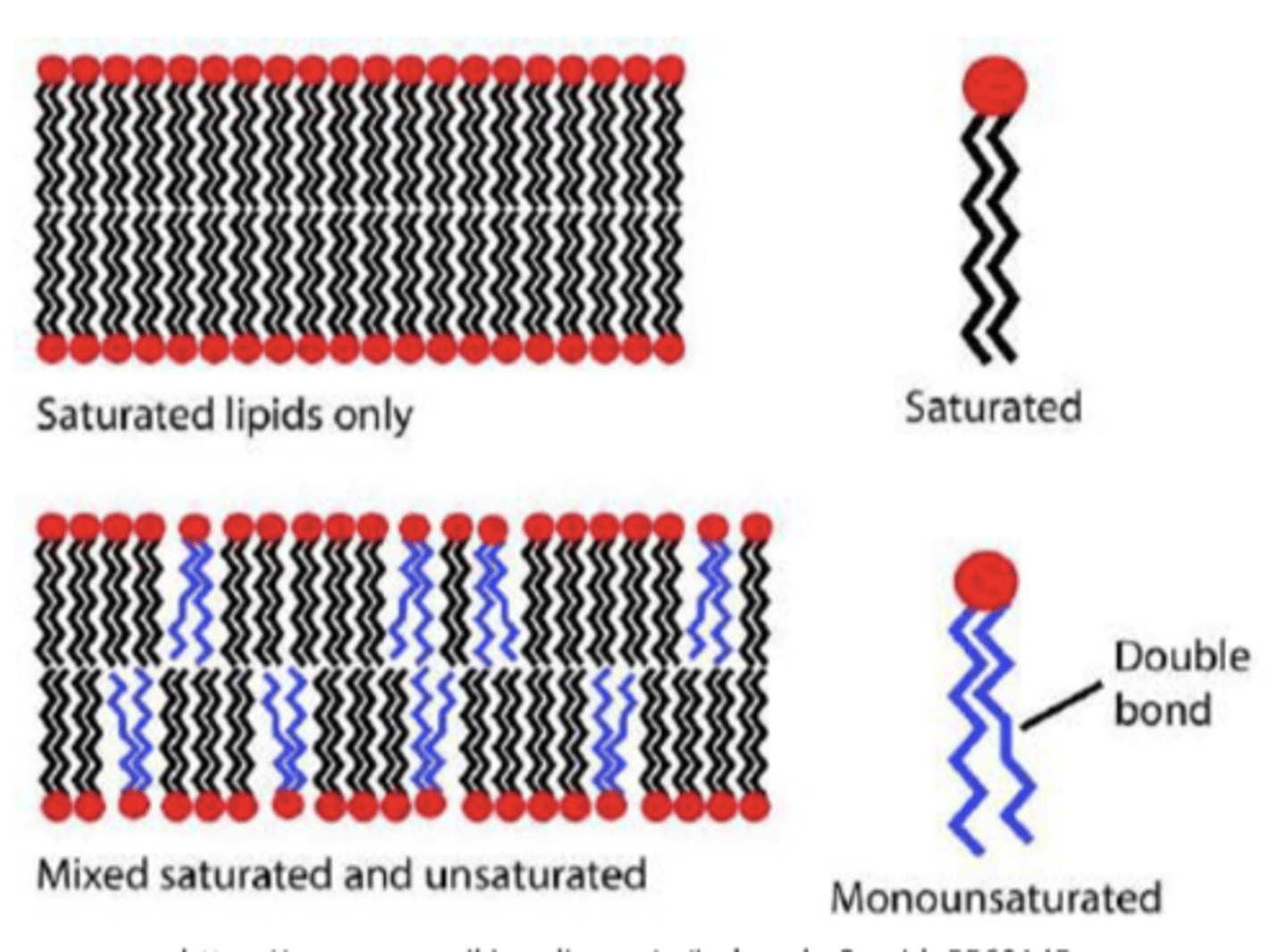
Unsaturated fatty acids
have double bonds and can be divided into monounsaturated fatty acids (one double bond) and polyunsaturated fatty acids (two or more double bonds)
Cis-unsaturated fatty acids
have kinks that cause the hydrocarbon tails to bend
- as a result, they do not pack tightly
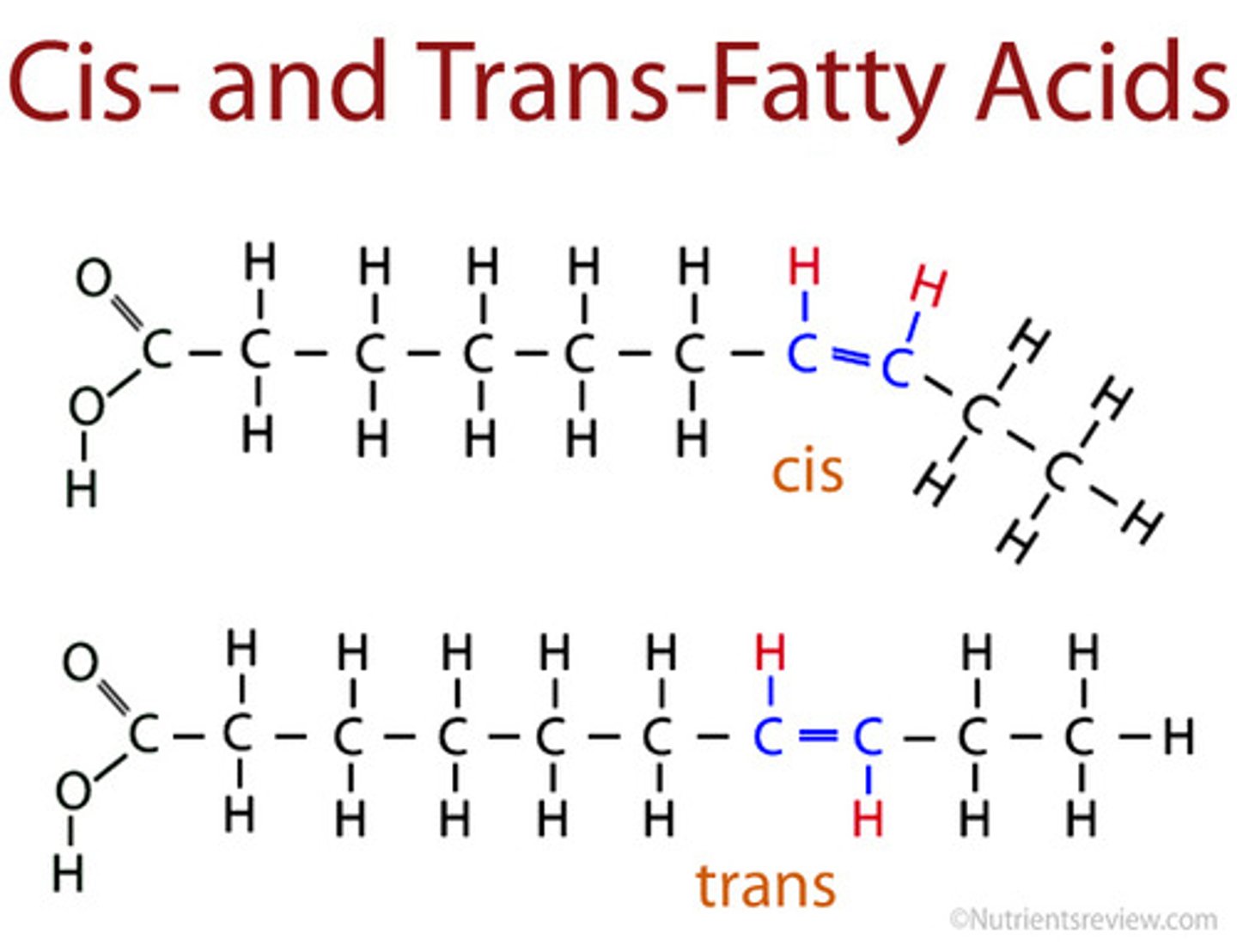
Trans-unsaturated fatty acids
straighter hydrocarbon tails, so they pack tightly
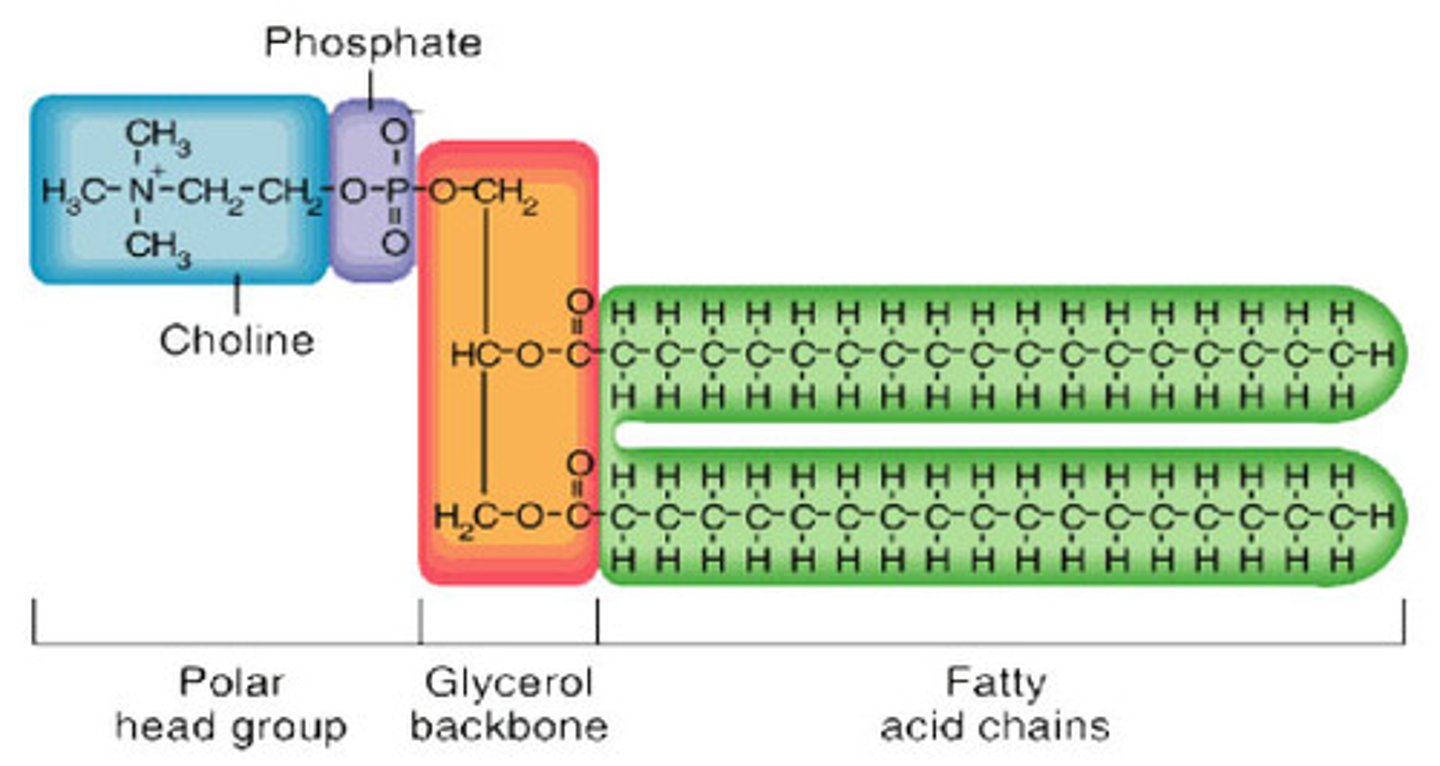
Phospholipids
lipid molecules that have a glycerol backbone, one phosphate group (hydrophilic), and two fatty acid tails (hydrophobic)
- the phosphate group is polar, while the fatty acids are nonpolar
- as a result, they are amphipathic (both hydrophobic and hydrophilic)
- spontaneously assemble to form lipid bilayers in an aqueous environment
Cholesterol
an amphipathic lipid molecule that is a component of cell membranes
- most common precursor to steroid hormones (lipids with four fused hydrocarbon rings)
- starting material for vitamin D and bile acids
- helps regulate membrane fluidity
Factors that influence membrane fluidity
1. Temperature - ↑ temperatures increase fluidity while ↓ temperatures decrease it.
2. Cholesterol - holds membrane together at high temperatures and keeps membrane fluid at low temperatures.
3. Degrees of unsaturation - saturated fatty acids pack more tightly than unsaturated fatty acids, which have double bonds that may introduce kinks.
Lipoproteins
allow the transport of lipid molecules in the bloodstream due to an outer coat of phospholipids, cholesterol, and proteins
Low-density lipoproteins (LDLs)
have low protein density and work to deliver cholesterol to peripheral tissues
- Sometimes considered "Bad cholesterol" - can cause vessel blockage and heart disease
High-density lipoproteins (HDLs)
have high protein density and take cholesterol away from peripheral tissues
- Considered "Good cholesterol" because they deliver cholesterol to the liver to make bile (reduces blood lipid levels)
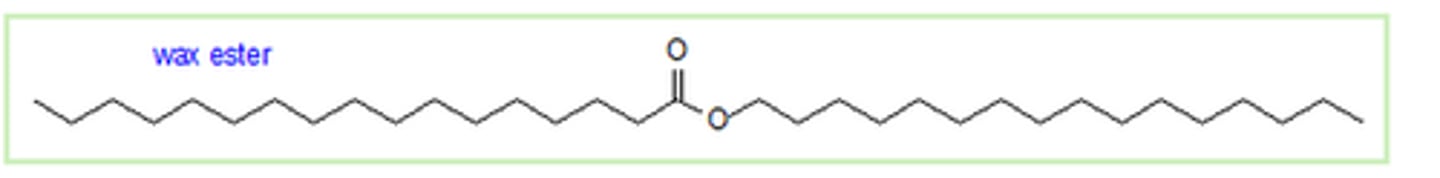
Waxes
simple lipids with long fatty acid chains connected to monohydroxy alcohols (contain a single hydroxyl group) through ester linkages
- used mainly as hydrophobic protective coatings
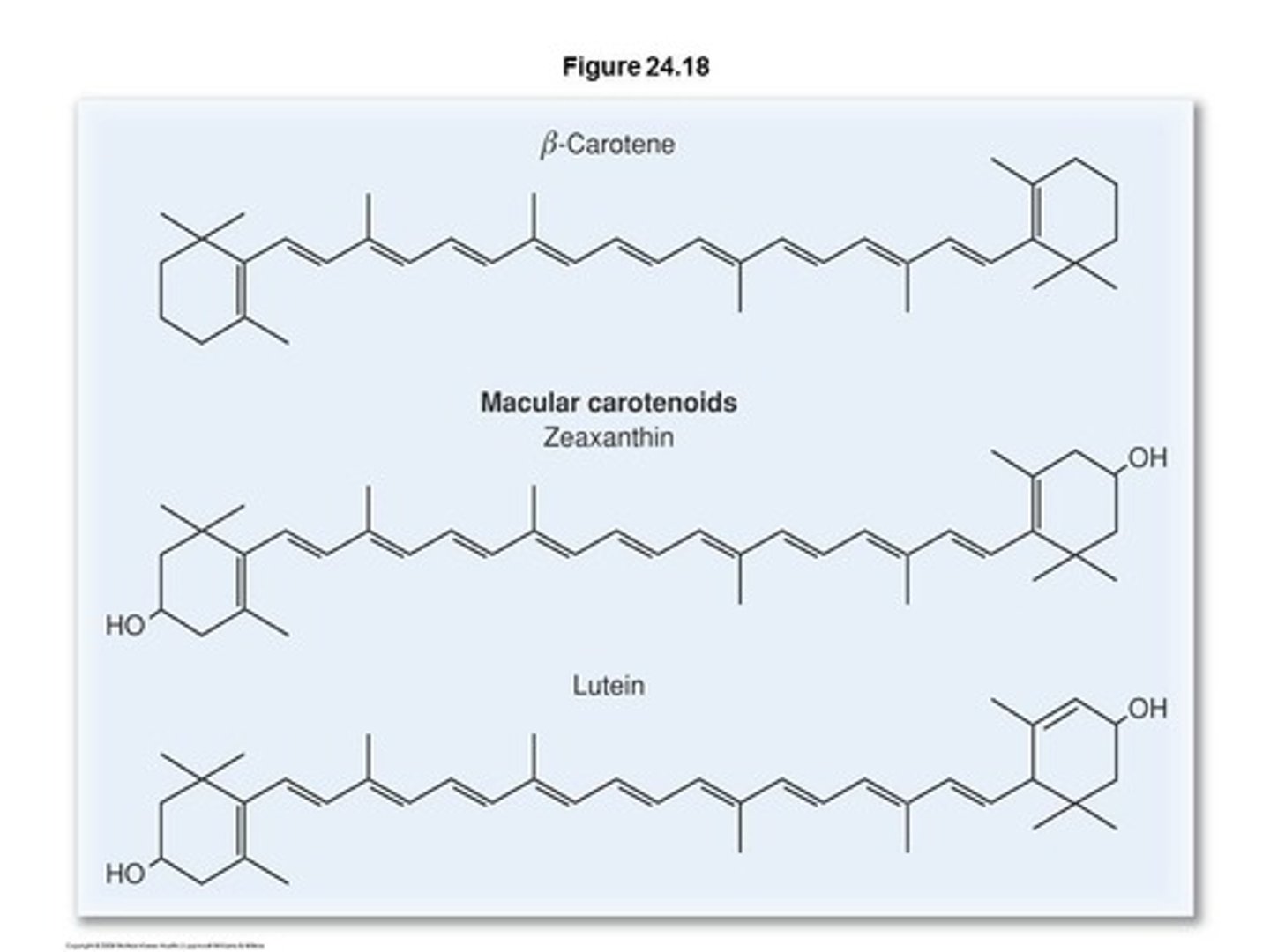
Carotenoids
lipid derivatives containing long carbon chains with conjugated double bonds and six-membered rings at each end
- They function mainly as pigments
Sphingolipids
have a backbone with aliphatic (non-aromatic) amino alcohols and have important functions in structural support, signal transduction, and cell recognition
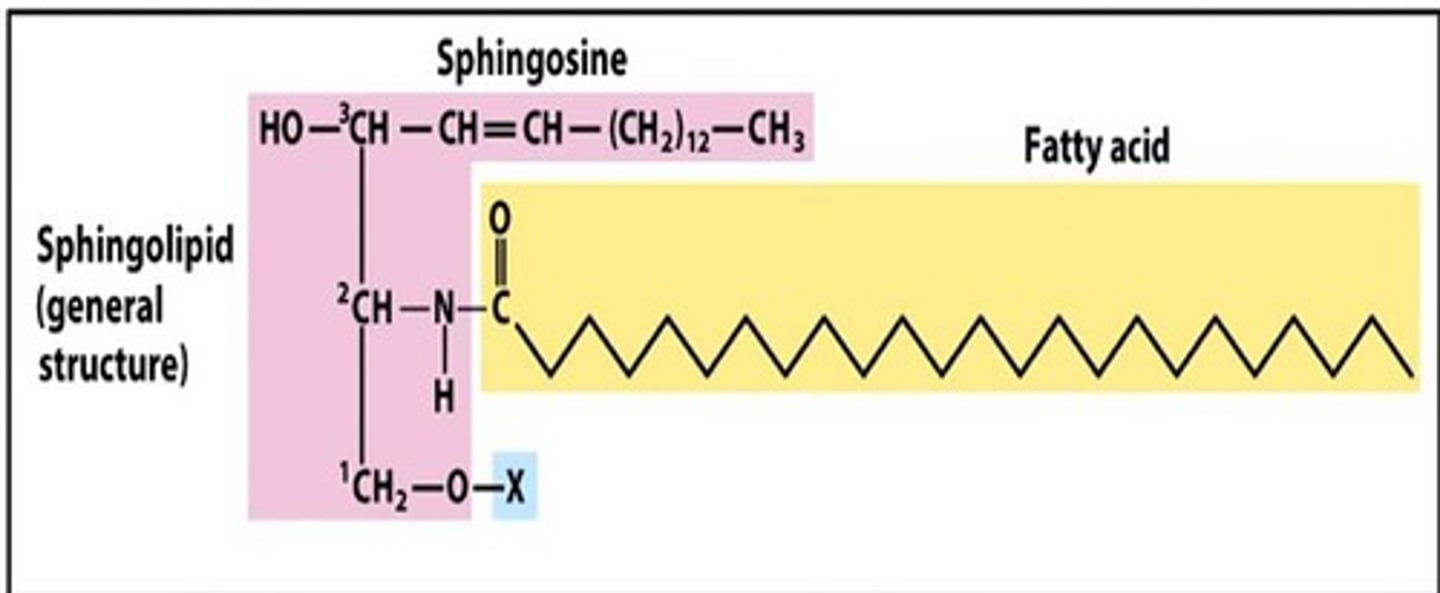
Glycolipids
lipids found in the cell membrane with a carbohydrate group attached instead of a phosphate group in phospholipids
- like phospholipids, they are amphipathic and contain a polar head and a fatty acid chain
Nucleic acids
contain carbon, hydrogen, oxygen, nitrogen, and phosphorus (CHONP)
- contain nucleotide monomers that build into DNA (deoxyribonucleic acid) and RNA (ribonucleic acid) polymers
Nucleosides
contain a five-carbon sugar and a nitrogenous base
Nucleotides
contain a five-carbon sugar, a nitrogenous base, and a phosphate group
Deoxyribose sugar
(in DNA) have a hydrogen at the 2' carbon
Ribose five-carbon sugar
(in RNA) have a hydroxyl group at the 2' carbon
Nitrogenous bases in DNA
- purines (two-ringed structure): adenine (A) and guanine (G)
- pyrimidines (one-ringed structure): thymine (T) and cytosine (C)
Uracil (U)
nucleotide that replaces thymine (T) in RNA
- pyrimidine (one-ringed structure)
Phosphodiester bonds
formed through a condensation reaction where the phosphate group of one nucleotide (at the 5' carbon) connects to the hydroxyl group of another nucleotide (at the 3' carbon) and releases a water molecule as a by-product
- a series of phosphodiester bonds create the sugar-phosphate backbone, with a 5' end (free phosphate) and a 3' end (free hydroxyl)
- nucleic acid polymerization proceeds as nucleoside triphosphates are added to the 3' end of the sugar-phosphate backbone
DNA structure
an antiparallel double helix, in which two complementary strands with opposite directionalities (positioning of 5' ends and 3' ends) twist around each other
- adenine can only H-bond to thymine (using two hydrogen bonds)
- guanine can only H-bond to cytosine (using three hydrogen bonds)
- In RNA, uracil binds to adenine, replacing thymine
mRNA
messenger RNA, a single-stranded RNA molecule that encodes the information to make a protein
- single-stranded after being copied from DNA during transcription
miRNA
microRNA, small RNA molecules that can silence gene expression by base pairing to complementary sequences in mRNA
rRNA
ribosomal RNA, it is formed in the nucleolus of the cell and helps ribosomes translate mRNA
dsRNA
double stranded RNA, some viruses carry their code as double stranded RNA
- dsRNA must pair its nucleotides, so it must have equal amounts of A/U, and C/G
tRNA
transfer RNA, small RNA molecule that participates in protein synthesis
Modern cell theory
1. All lifeforms have one or more cells.
2. The cell is the basic structural, functional, and organizational unit of life.
3. All cells come from other cells (cell division).
4. Genetic information is stored and passed down through DNA.
5. An organism's activity is dependent on the total activity of its independent cells.
6. Metabolism and biochemistry (energy flow) occurs within cells
7. All cells have the same chemical composition within organisms of similar species.
Central dogma of genetics
information is passed from DNA -> RNA -> proteins
- except w/ reverse transcriptase and prions
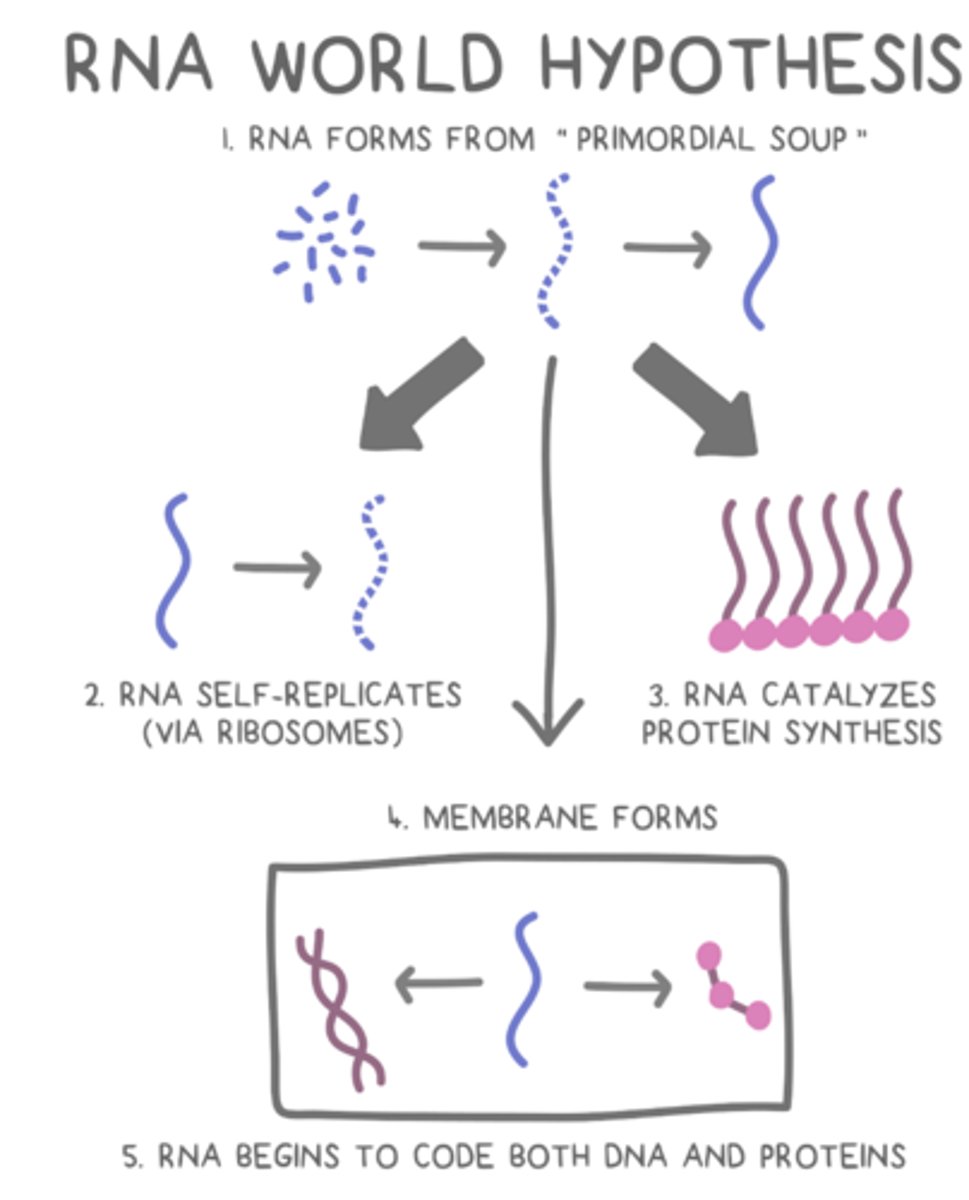
RNA World Hypothesis
states that RNA dominated Earth's primordial soup before there was life.
- RNA developed self-replicating mechanisms and later could catalyze reactions, such as protein synthesis, to make more complex macromolecules
- Since RNA is reactive and unstable, DNA later became a better way of reliably storing genetic information.
Cell membrane
hold cellular contents and are mainly composed of phospholipids and proteins, with small amounts of cholesterol
Membrane proteins
either integral or peripheral membrane proteins
Integral (transmembrane) proteins
transverse the entire bilayer, so they must be amphipathic. their nonpolar parts lie in the middle of the bilayer while their polar ends extend out into the aqueous environment on the inside and outside of the cell. usually assist in cell signaling or transport