LAB QUIZ #8 Biotechnology I: DNA Fingerprinting and Electrophoresis
1/46
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
47 Terms
what is biotechnology
- technology based on biology
- harnesses cellular and bio molecular processes to develop technologies and products that help improve life

where is biotechnology used
-uses biological processes that occur within microorganisms to do some of the following:
■ Food products (bread and cheese)
■ Preserve dairy products
In medicine, it's used for drug development, gene therapy, and diagnostics.
In agriculture, it enables the creation of genetically modified crops with enhanced traits like pest resistance.
Environmental biotechnology utilizes microorganisms for bioremediation and waste management.
Other applications include food processing, energy production (like biofuels), and industrial processes (e.g., producing biodegradable plastics).

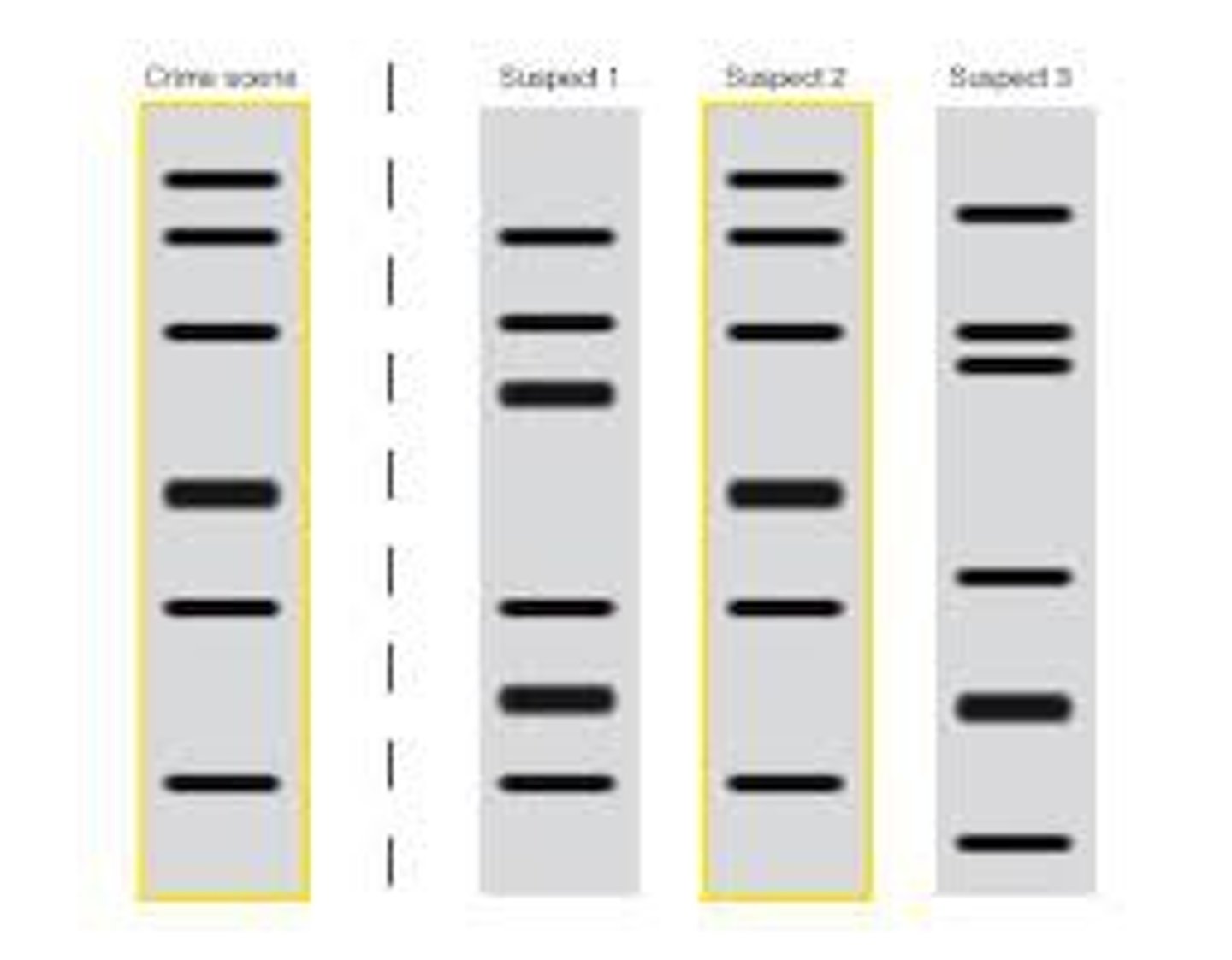
DNA Fingerprinting
technique that detects minisatellites in the genome to produce a pattern unique to individuals
what is the Restriction enzyme in the lab
Endonucleases
Act as a molecular scissors
what do restriction enzymes do
Restriction enzymes cut at specific sequences that it recognizes = restriction sites.
Restriction sites are
Palindromes (both strands read the same in the 5'-3' direction)
describe the 2 restriction enzymes used in the lab
Restriction enzymes are specific and have different restriction sites.
They are named after the bacteria from which they are isolated ■ EcoRI - found in E. coli
■ Pstl - found in P. stuartii
Restriction Fragment Length Polymorphism (RFLP)
refers to differences or variations amongst people in their DNA sequences and where restriction sites are recognized by restriction enzymes.
Variations result in different DNA fragment lengths that are produced by digesting DNA with a restriction enzyme.
Note: # restriction sites =
# of bands - 1
Gel matrix is made of ________
agarose

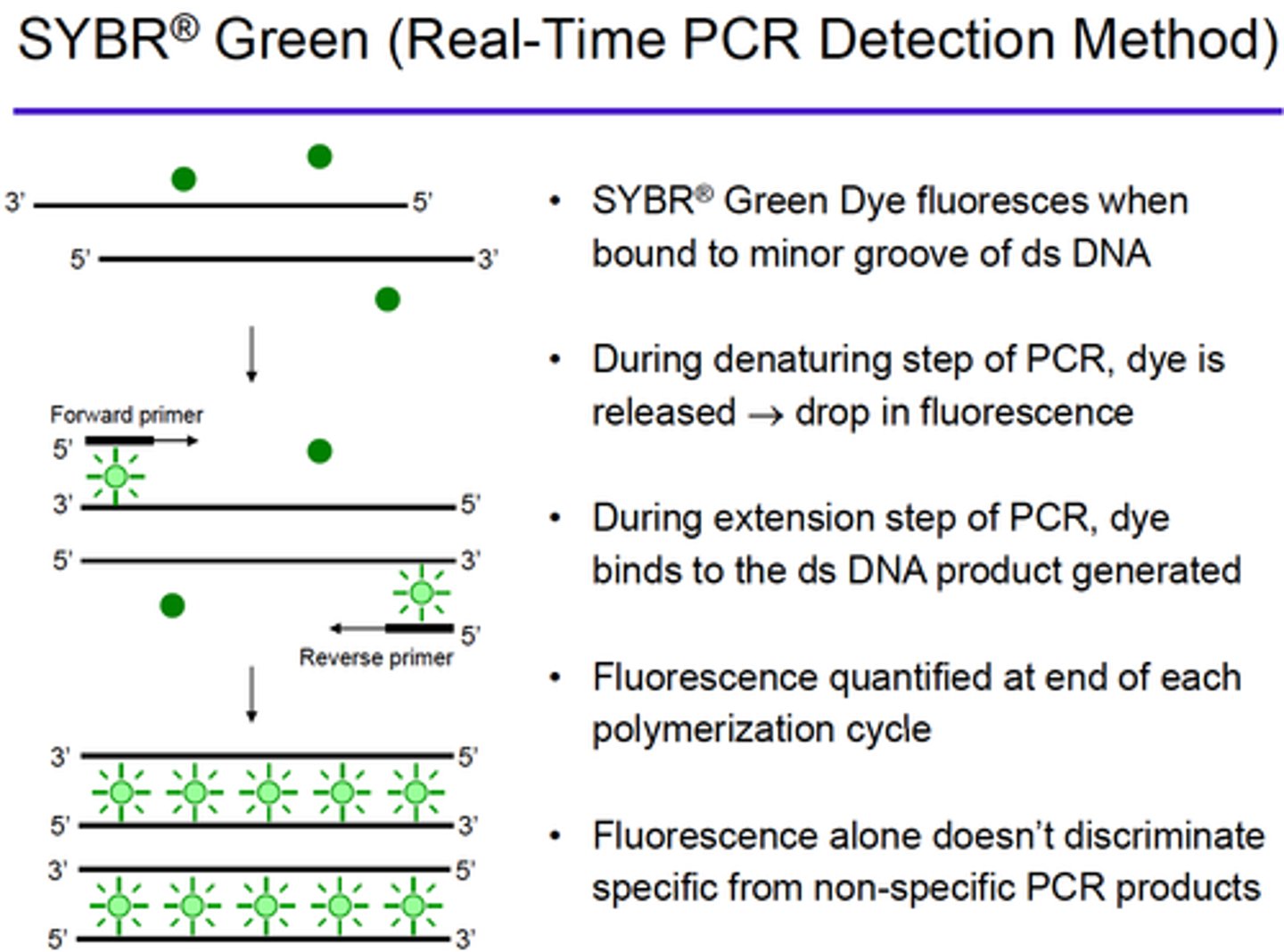
SYBR green
(binds to DNA and fluoresces under UV light)
TAE buffer solution
Maintains pH
Salts in the buffer are used to conduct the electrical current
common running buffer used in agarose gel electrophoresis, particularly for separating nucleic acids like DNA and RNA. It's favored for its compatibility with in-gel manipulations and band recovery, and for the faster migration of double-stranded DNA.
Electrical charge in Agarose Gel Electrophoresis
Anode (+): red end, positive current
Cathode (-): black end, negative current
What charge does DNA have?
negative charge
due to the phosphate groups in its sugar-phosphate backbone
What end would the DNA be loaded into the gel?
In gel electrophoresis, DNA samples are loaded into the wells at the negative end of the gel. Since DNA is negatively charged due to its phosphate backbone, it will be attracted to the positive electrode and migrate towards it during electrophoresis.
Cathode (-): black end, negative current
DNA Loading dye
Adds color to you sample to make it easier to see whether you are successfully loading the gel! ○ Loading dyes are dense and keep the DNA in your sample from diffusing away in the buffer
Gel electrophoresis is used to separate what
DNA fragments based on size
Agarose gels
tiny holes for the DNA to travel towards the anode (+) side.

smaller vs larger fragments
Larger fragments get stuck towards to the top (close to the cathode (-))
Smaller fragments migrate further in the gel (close to the anode (+))
Bands at the top of the gel =
larger DNA fragments
Bands at the bottom of the gel =
smaller DNA fragments
Darker bands =
Darker bands = more DNA fragments
If a single strand of DNA gets cut 2x by a restriction enzyme, how many fragments will appear on the gel?
three fragments that will appear on the gel. Each cut creates a new end, effectively dividing the original strand into smaller pieces.
If you started with a single strand of DNA and after the experiment you visualize 4 bands on the gel, how many restriction sites did the DNA have?
If you visualize 4 bands on a gel after using a restriction enzyme on a single-stranded DNA, the DNA likely had 3 restriction sites. With each restriction site, the DNA is cut into an additional fragment, and one more band appears on the gel. Since the initial DNA was a single strand, and 4 fragments were produced, there must have been 3 cutting events (restriction sites)
RFLP and how they are produced
RFLP, or Restriction Fragment Length Polymorphism, is a technique that analyzes DNA by identifying variations in fragment lengths produced by restriction enzymes. These variations arise from mutations that create or abolish restriction enzyme recognition sites, leading to different sized DNA fragments when the DNA is cut with these enzymes
EcoRI palindrome sequence
The EcoRI palindrome sequence is 5'-GAATTC-3'. This sequence is recognized by the restriction enzyme EcoRI, which cleaves the phosphodiester bond between the G and A nucleotides. The complementary strand also reads the same when read in the reverse direction, making it a palindrome.
If a linear DNA molecule has 5 restriction sites how many fragments will be produced
A linear DNA molecule with 5 restriction sites will be cut into 6 fragments by a restriction enzyme. Each cut made by the enzyme creates a new fragment. Therefore, with 5 restriction sites, there will be 5 cuts, resulting in 6 fragments.
If a linear DNA molecule has 2 restriction sites how many fragments will be produced
A linear DNA molecule with two restriction sites will be cut into three fragments. Each restriction site acts as a point where the DNA molecule is cut, and with two sites, you get one fragment before the first cut, one fragment between the two cuts, and one fragment after the second cut
If a linear DNA molecule has 8 restriction sites how many fragments will be produced
A linear DNA molecule with 8 restriction sites will be cut into 9 fragments. In general, when a linear DNA molecule is cut at 'n' restriction sites, it will produce 'n+1' fragments
if you are given a liner DNA strand with restriction enzyme A and restriction enzyme B and and there are 3 restriction sites for A and 2 sites for B what happens if you only provide restriction enzyme A how many fragments will be produced
f a linear DNA strand with 3 restriction sites for enzyme A and 2 for enzyme B is cut with only enzyme A, it will produce 4 fragments. Since enzyme A has 3 restriction sites, it will make 3 cuts, and each cut creates a new fragment. Thus, a linear DNA strand with 'n' restriction sites will result in 'n+1' fragments.
polymerase chain reaction (PCR)
technique that allows molecular biologists to make many copies of a particular gene
what happens if you only have a really small amount of DNA available to you but you need more for DNA fingerprinting
If only a small amount of DNA is available, techniques like PCR (Polymerase Chain Reaction) can be used to amplify it, creating enough DNA for fingerprinting.
would you expect larger or smaller DNA fragments to travel farther down the gel
In gel electrophoresis, you would expect smaller DNA fragments to travel farther down the gel.
define cathode
Negatively charged electrode
define anode
Positive electrode
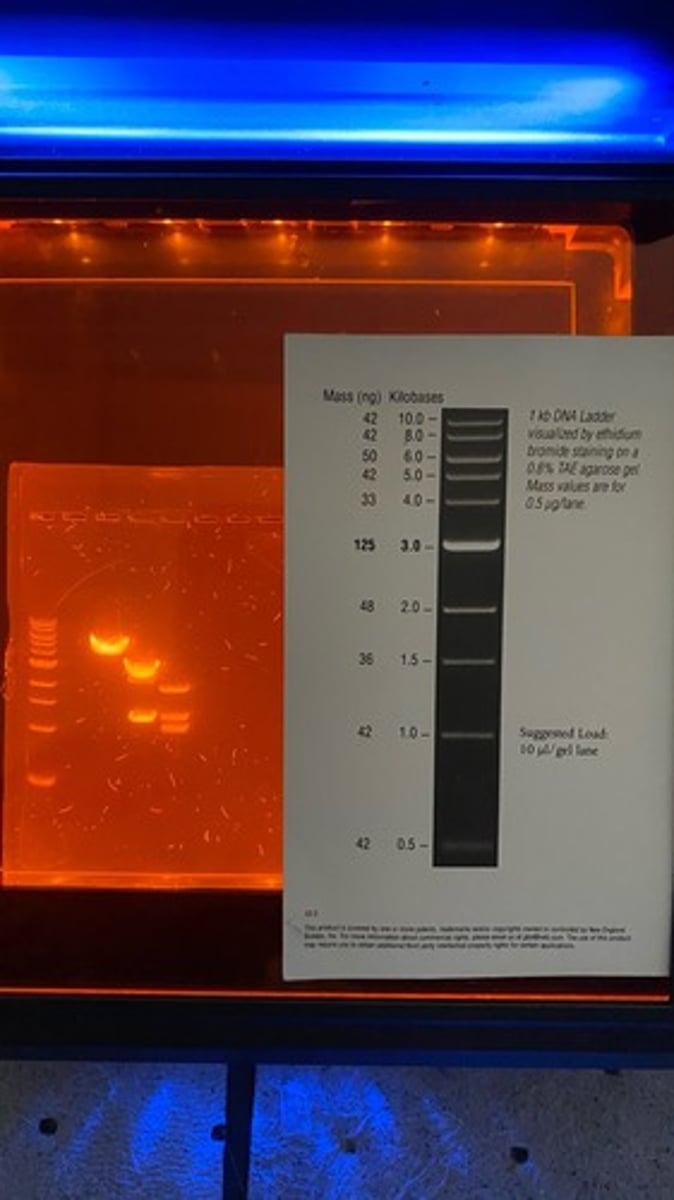
what is a DNA ladder
DNA ladders function to inform on the size of the DNA fragments (base pairs) and serve as a control for the gel.
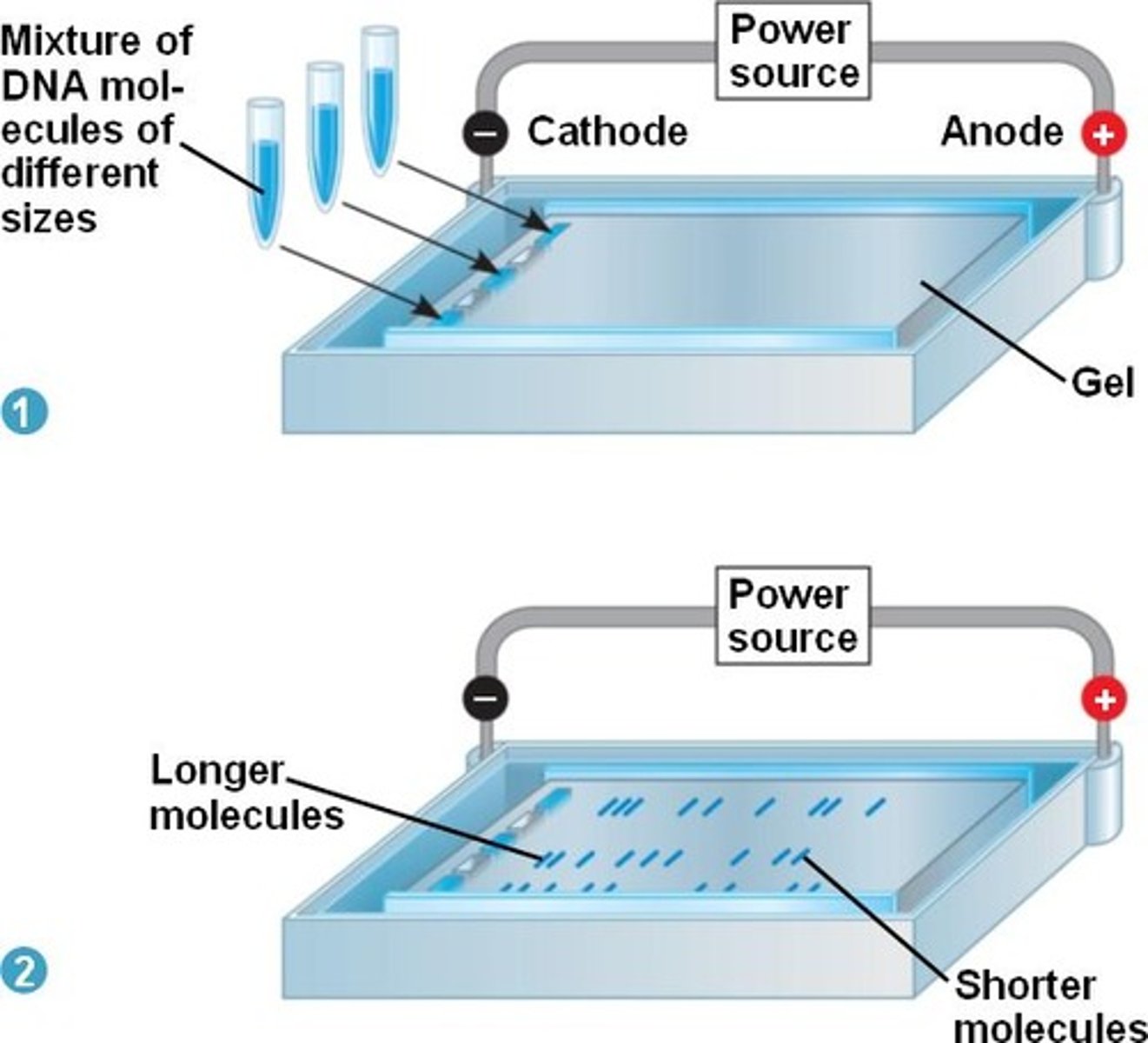
components of an agarose gel electrophoresis
an agarose gel, a gel box (also known as an electrophoresis chamber), a power supply, loading buffer, and a visualization system. These components work together to separate DNA or RNA fragments based on their size and charge
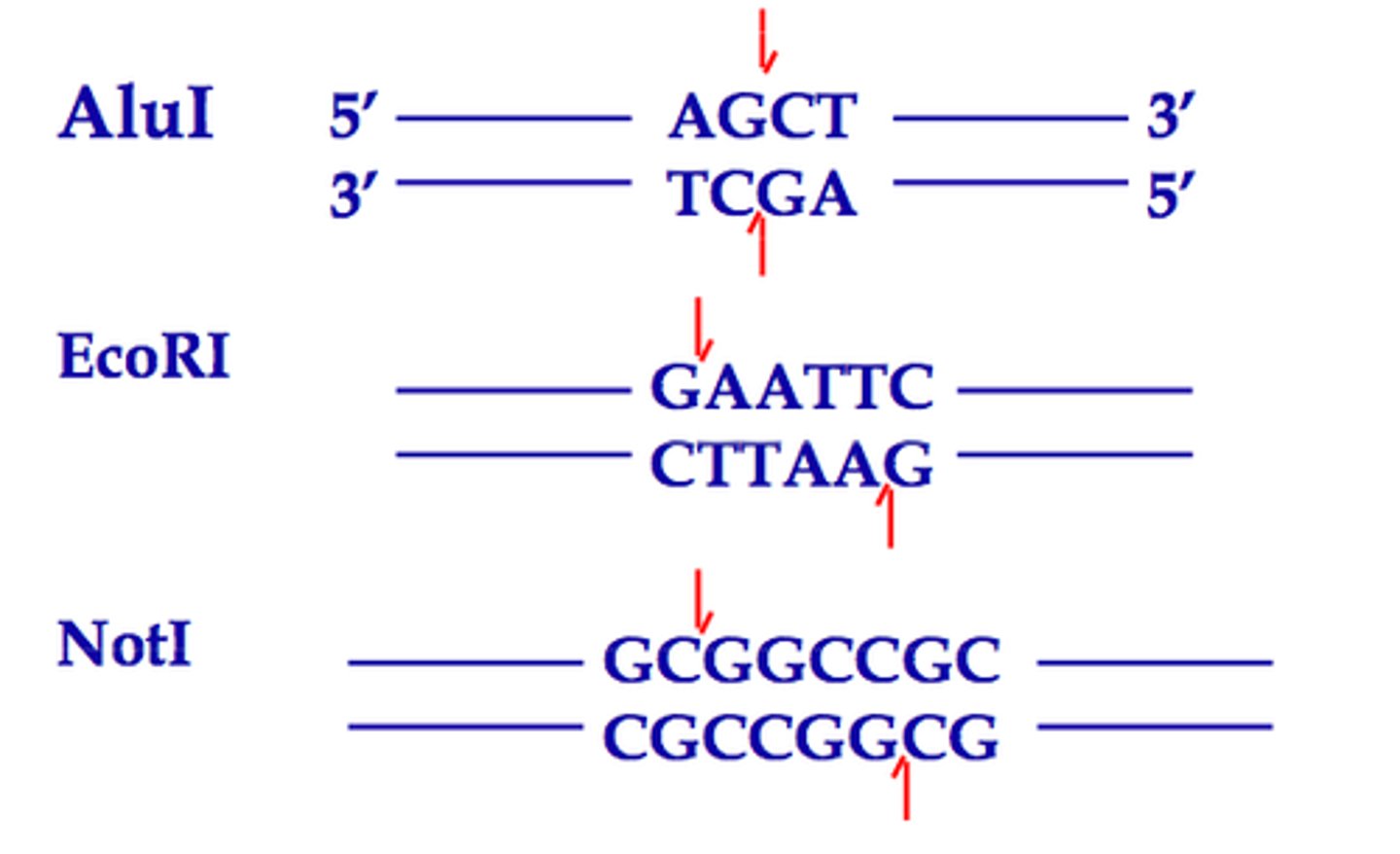
what do restriction enzymes regogonize
specific DNA sequences called recognition sites or restriction sites. These sites are typically short, specific sequences of nucleotides (usually 4-8 base pairs long) in the DNA double helix. Each restriction enzyme recognizes a unique sequence, and when it finds its target, it cuts the DNA molecule at a specific location within or near that site.
restriction sites
Specific site where a restriction enzyme cuts the DNA
how are restriction sites related to palindromes
cut double stranded DNA molecules by cleaving phosphodiester bonds at palindromic sequences
is argose gel solid or porus
In essence, agarose gel is a solid material with a permeable, porous structure, allowing for the passage of molecules through its matrix.
what does it mean if a gel is porous
A gel is considered porous when it contains interconnected voids or pores within its structure, allowing for the passage of fluids or other substances
Agarose gel electrophoresis separates DNA fragments _______-
primarily based on their size (or molecular weight)
the enzyme that cuts or chemically separates the DNA moleulce at a site
restriction site
when were restriction enzymes first isolated
from bacteria that could not be infected with bacteriophages
what 3 restriction enzymes did we use
EcoRI- isolated from Escherichia coli
Hinddll- Haemophilus influenze
Pstl - providencia stuartii
what does each cut between
EcoRI-
Hinddll-
Pstl -
EcoRI- between G and A
Hinddll- As
Pstl - C and T