Unit 3 Biology
1/111
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
112 Terms
Gene Expression in Prokaryotic Cells
Circular chromosome; occurs at the same time
RNA vs DNA
Single stranded, contains a ribose instead of a deoxyribose, and can contain more complex structures
rRNTp vs dNTP
Differences: 2’ —OH vs 2’ —H
Similarities: 3 Phosphates, added to 3’ -OH
Types of Prokaryotic RNA
mRNA and ncRNA
mRNA (how long?)
Messanger RNA; Long (average 3000 nucleotides), codes for proteins. Coding region is 1200 BPs
Contains a promoter sequence, regulatory sequence, coding sequence, and transcription terminator
Types of sequences in mRNA
Promoter: Determines where transcription starts
Regulatory Sequence: Determines when, where, how much mRNA is made
Coding Sequence: ORF, opening region frame
Transcription Terminator: Where transcription ends
ncRNA
Includes tRNA (transfer RNA), rRNA (ribosomal RNA), and sRNA (small RNA)
Gene
Unit of heredity, carries the genetic information for a protein (polypeptide) or RNA
Opening Reading Frame
Stretches of DNA that lack a stop codon; every 3 nucleotides encode for amino acid during translation of mRNA to form a proteins; there are 3 reading frames possible per DNA strand
Basic Structure of a Bacterial Protein-Coding Gene (DRAW)
Promoter: From -10 to -35 site, where RNA polymerase binds. Determines transcription start
Transcription Start Site (TSS): From +1
Opening Reading Frame (ORF): Protein coding sequence
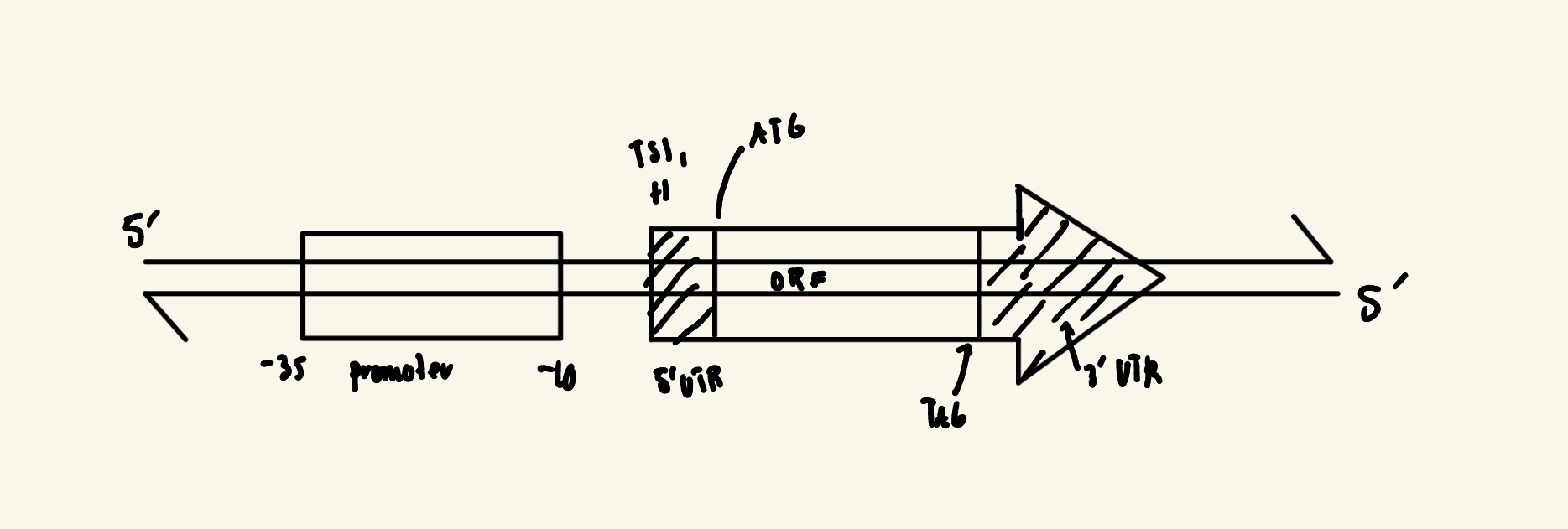
Start and Stop Codons
Start; ATG
Stop: TAG, TGG, TGA
Template vs Coding Strand
Template strand is use for mRNA synthesis
Coding strand is identical to mRNA strand, except A is replaced with U
Transcription Overview
RNA binds to single stranded DNA template at the promoter
DNA helix unwinds
RNA is synthesized from rNTPS from 5’ to 3’ direction
5’ end of transcript is displaced from template as polymerase moves
How to know which strand is template strand for a gene? Draw
3’ to 5’
RNA vs DNA Polymerase
Doesn’t require a primer, only one nucleotide polymer is built, no proofreading
How many total reading frames are possibe?
6
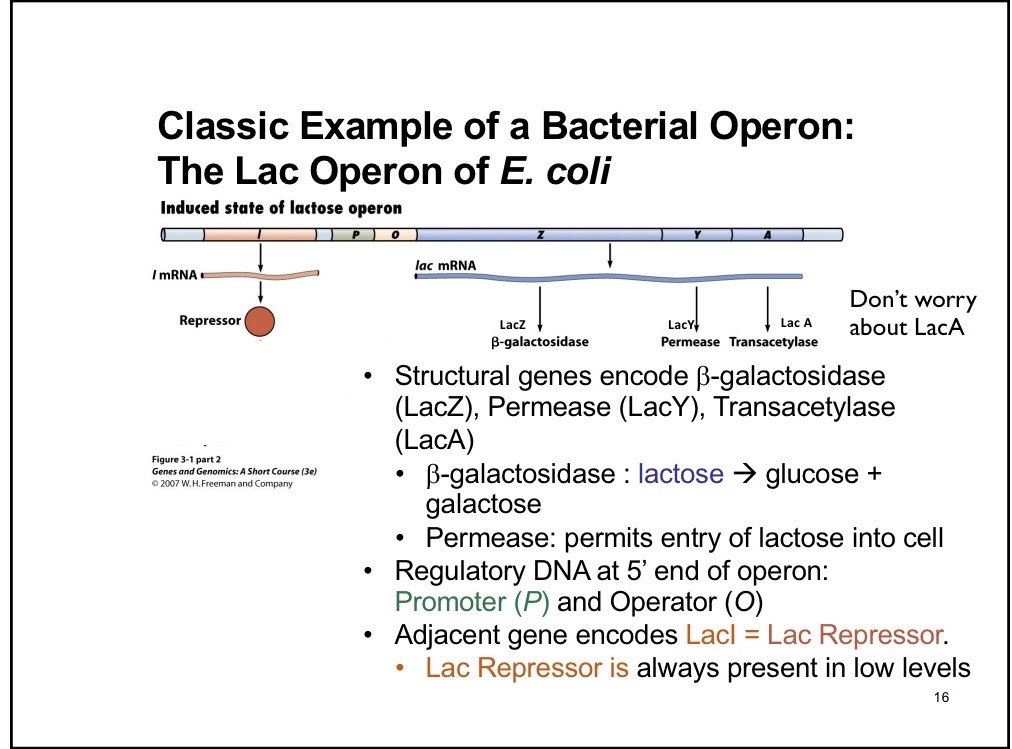
Operons
Are in prokaryotic cells. Multiple ORF in one sequence, each has its own ATG and STOP sequence.
RNA Polymerase Structure
6 subunits, one leaves
Initiation
Consensus sequence upstream of TSS in the promoter
Sigma subunits binds to the -10 and -35 sequences to correctly position the enzyme
After initation, the sigma subunit disassociate and RNA polymerase can continue elongation
Elongation
RNA polymerase synthesizes. 17 bp are unwound in the transcription bubble. Moves at 50 nucleotides per second and the error rate is 10^4 or 10^5
Intrinsic Termination
G-C rich stem-loop structure of mRNA followed by a rich string of As. RNA polymerase dissasociates when it encounters the G-C loop.
Rho Dependent Termination
Rho binds to rut sequence upstream of termination site and acts as helicase to pause the RNA-DNA hybrid and facilitates the release of mRNA
Helix Turn Helix
Recognition of specific binding sites
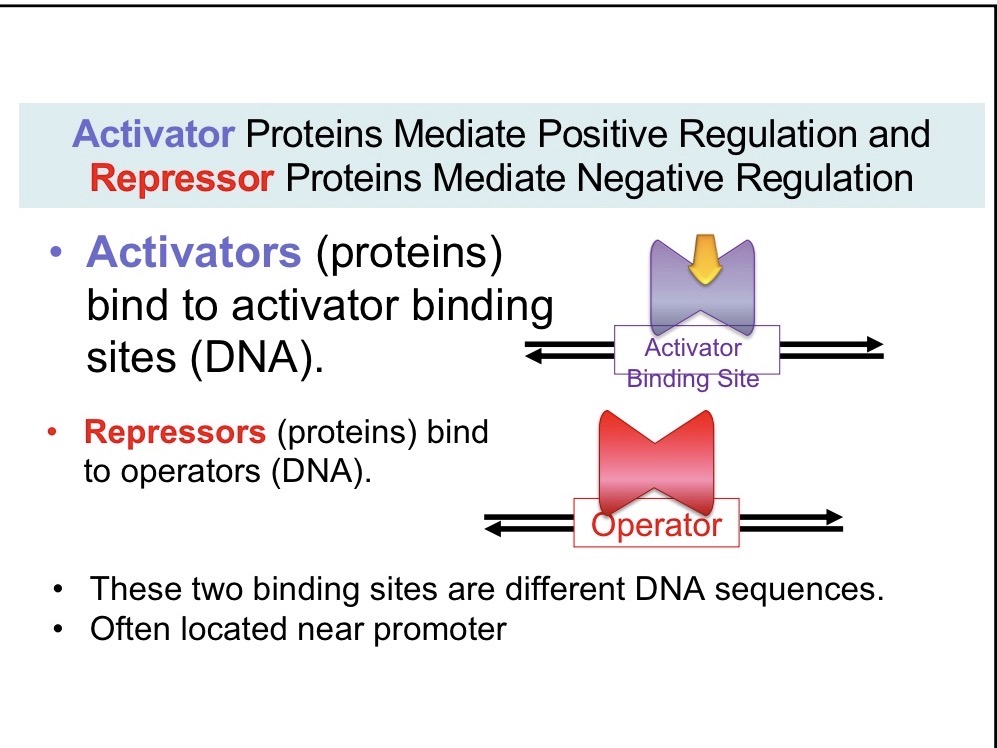
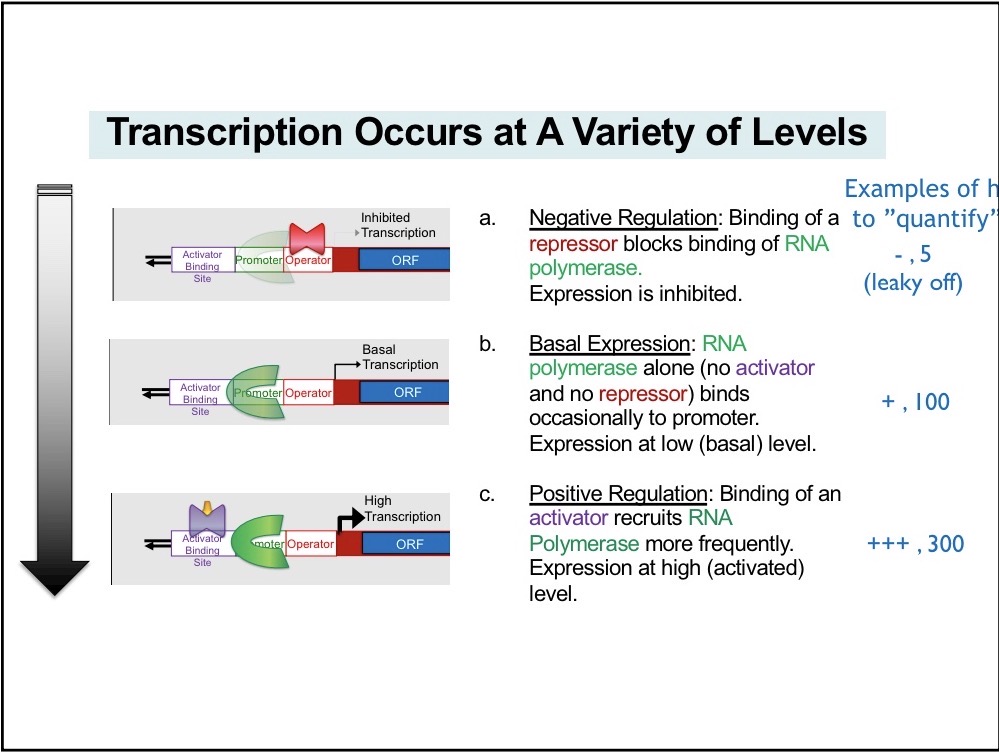
Activator
Mediate positive regulation and bind to DNA
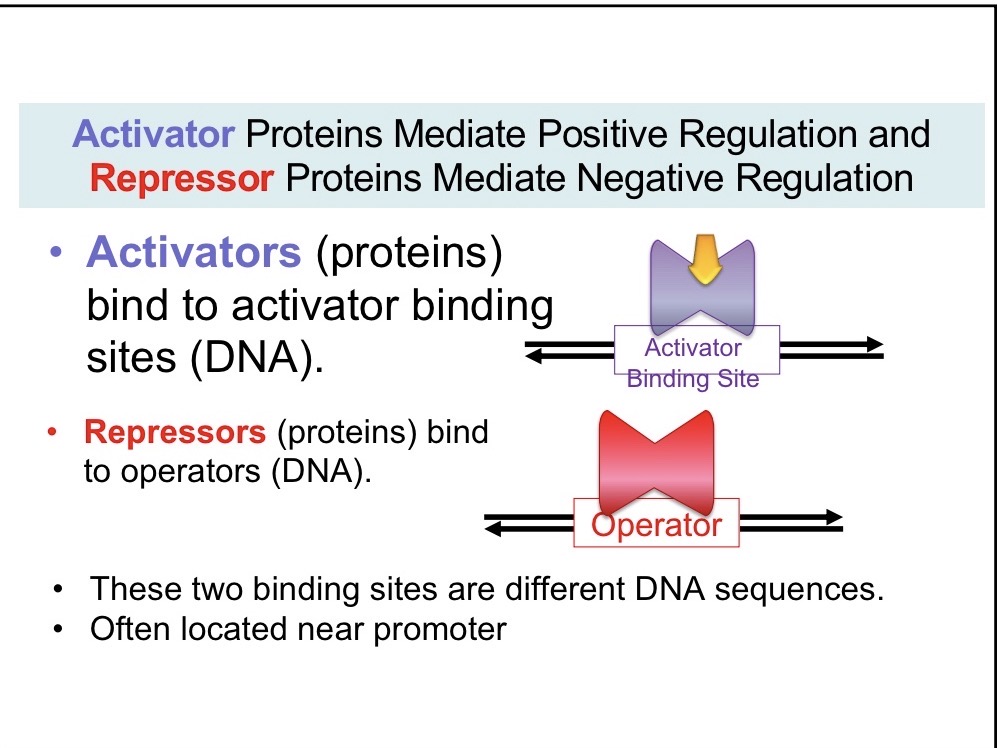
Repressors
Mediate negative regulation and bind to operators, block -10 part of consensus sequence
Inducers
Change the conformation of activators and repressors
Activator: Requires inducer to bind to active site
Repressor: Leaves operator in presence of inducer
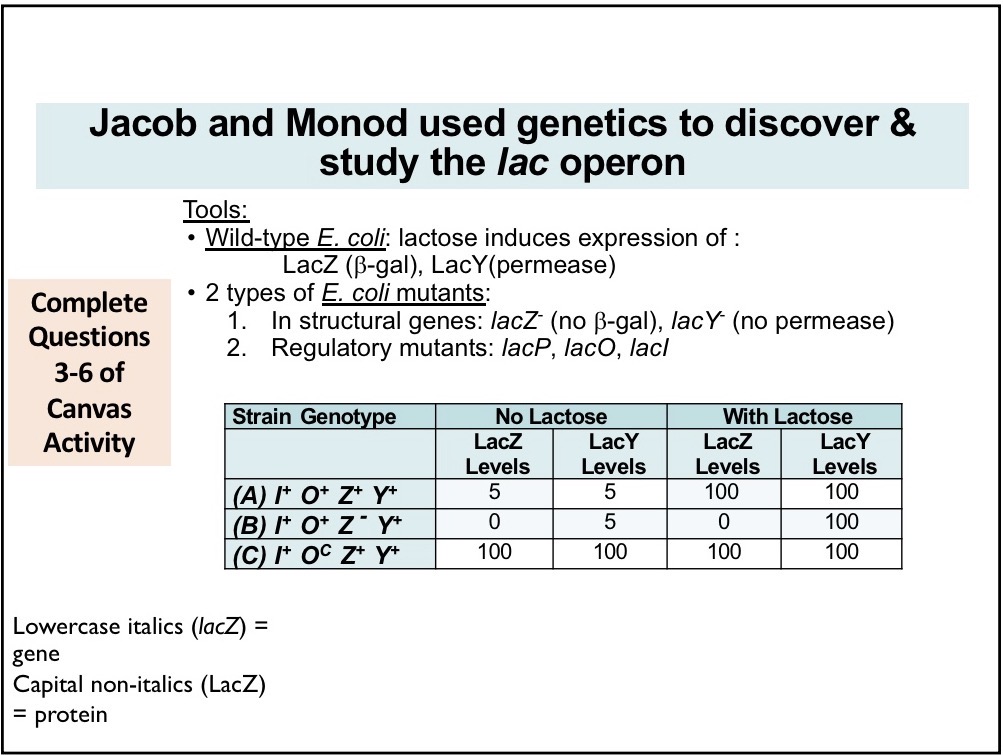
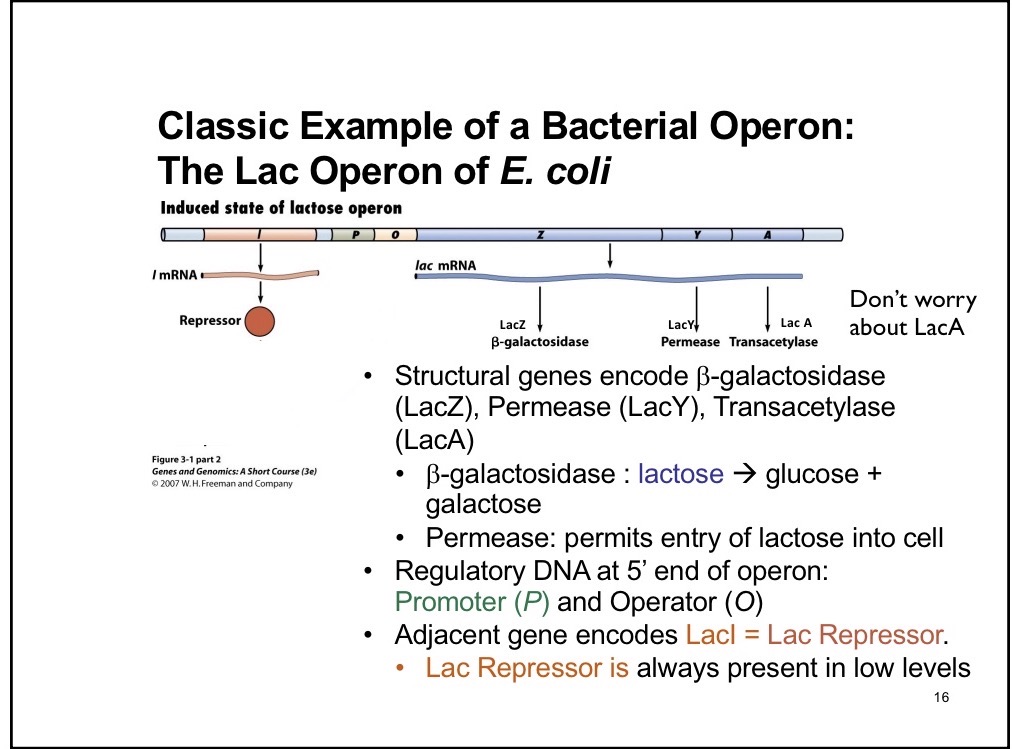
LacZ
B-galactosidase enzyme that breaks down into glucose + galactose
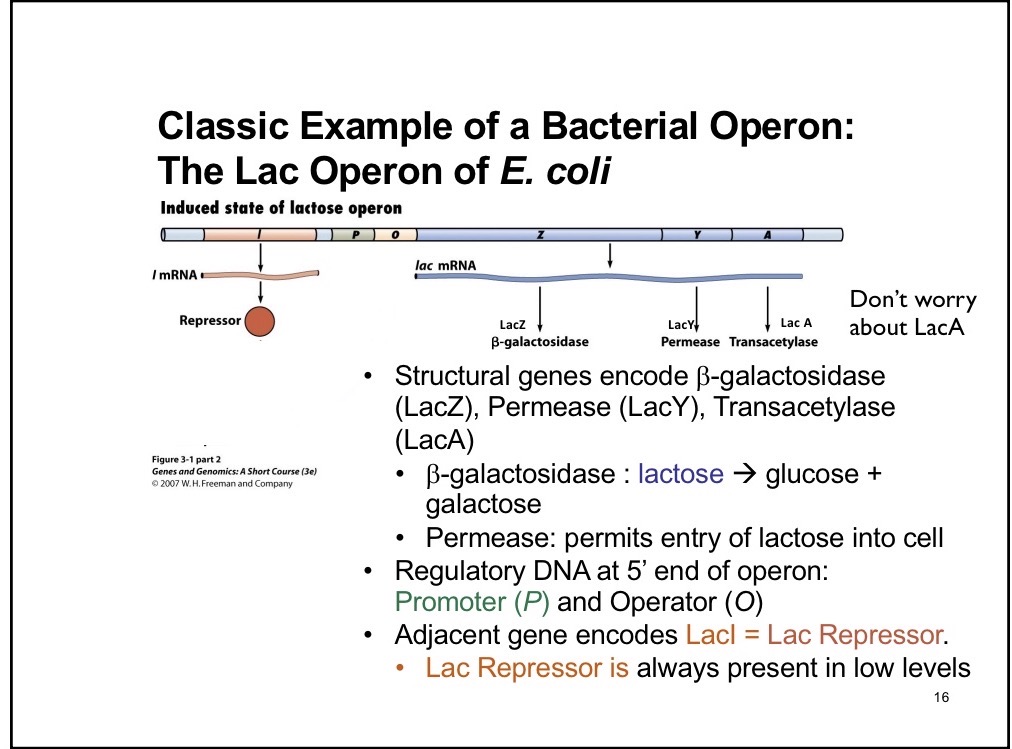
LacY
Permease; permits entry of lactose into cell
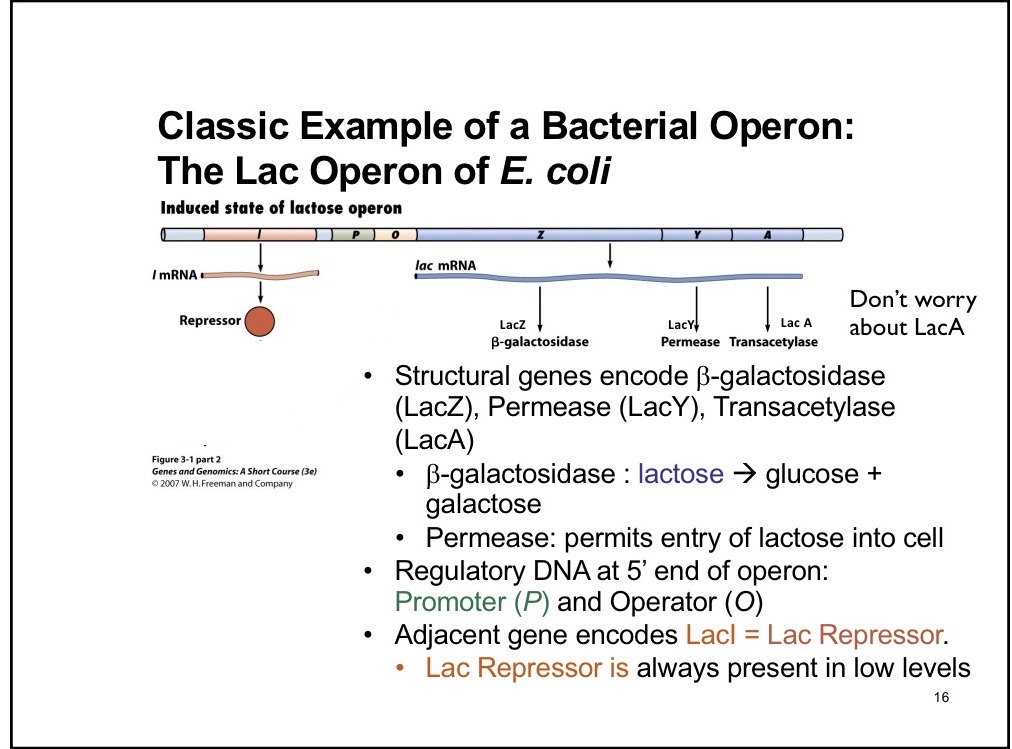
LacA (don’t need to know)
Transacetylase
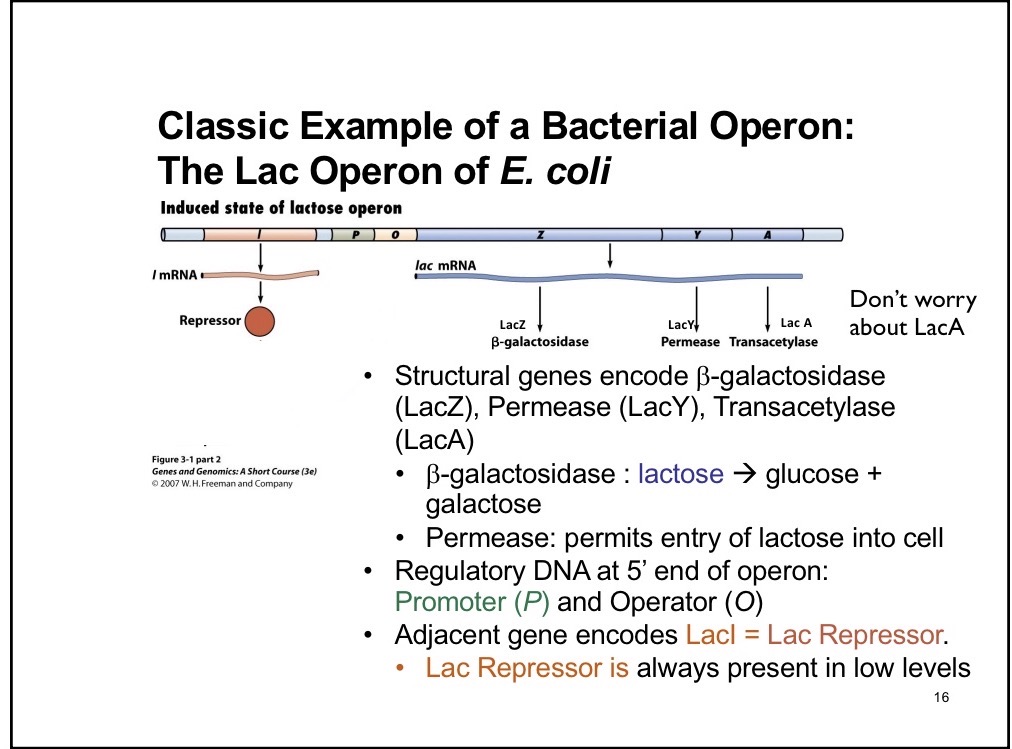
I component of Lac Operon
Creates repressor
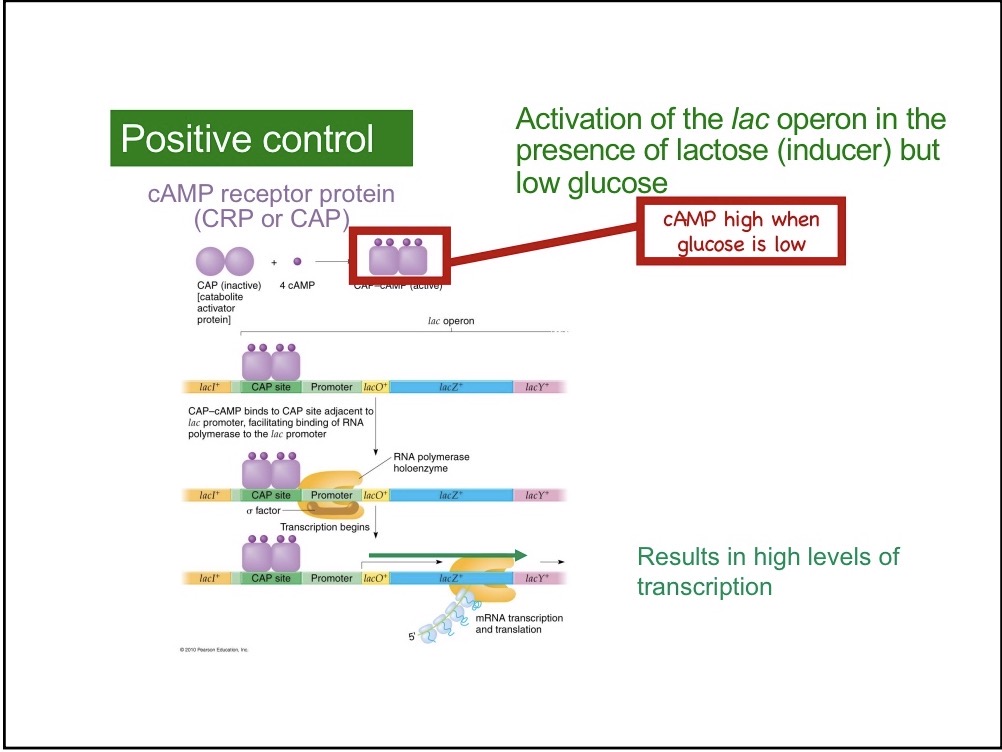
Glucose and cAMP relationship
Low glucose leads to high levels of cAMP
Highest Transcription Levels
High lactose and low glucose
CAP
cAMP binds to CAP, an activator protein. CAP protein recruits RNA polymerase by binding to the C-terminal domain of the alpha subunit
Basal Expression
No repression and no activator
Lac Repressor Binding
Binds to DNA as a dimer, is symmetrical, helix-turn-helix
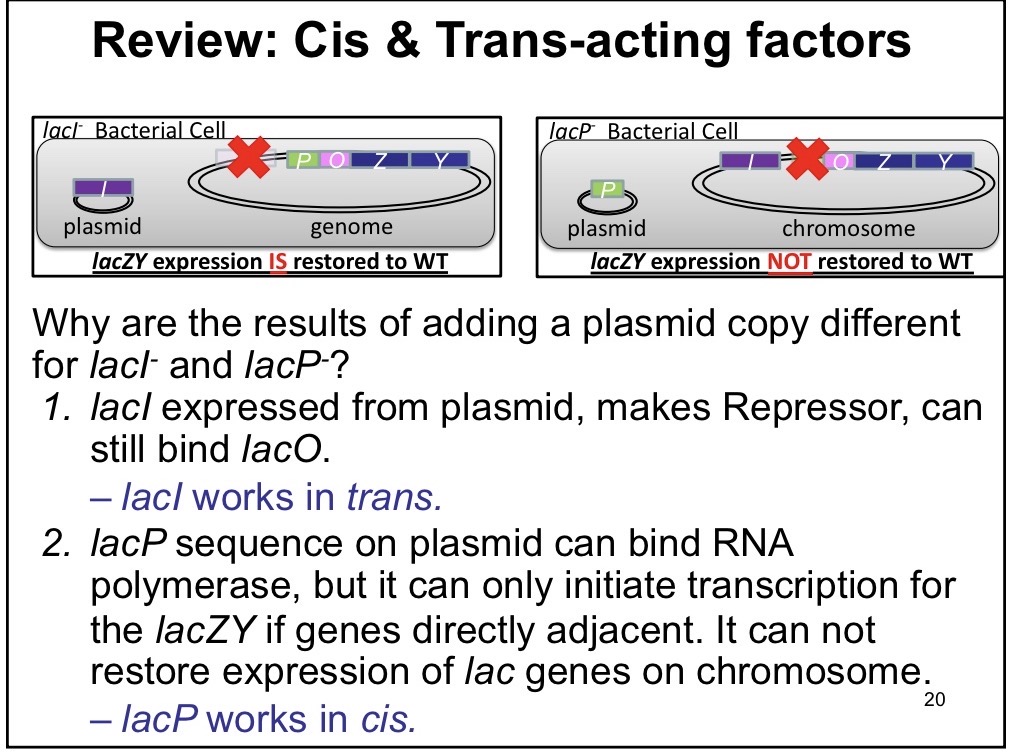
Jacob and Monod: Cis
Position and sequence are both important, DNA binding sites
Ex: Promoter
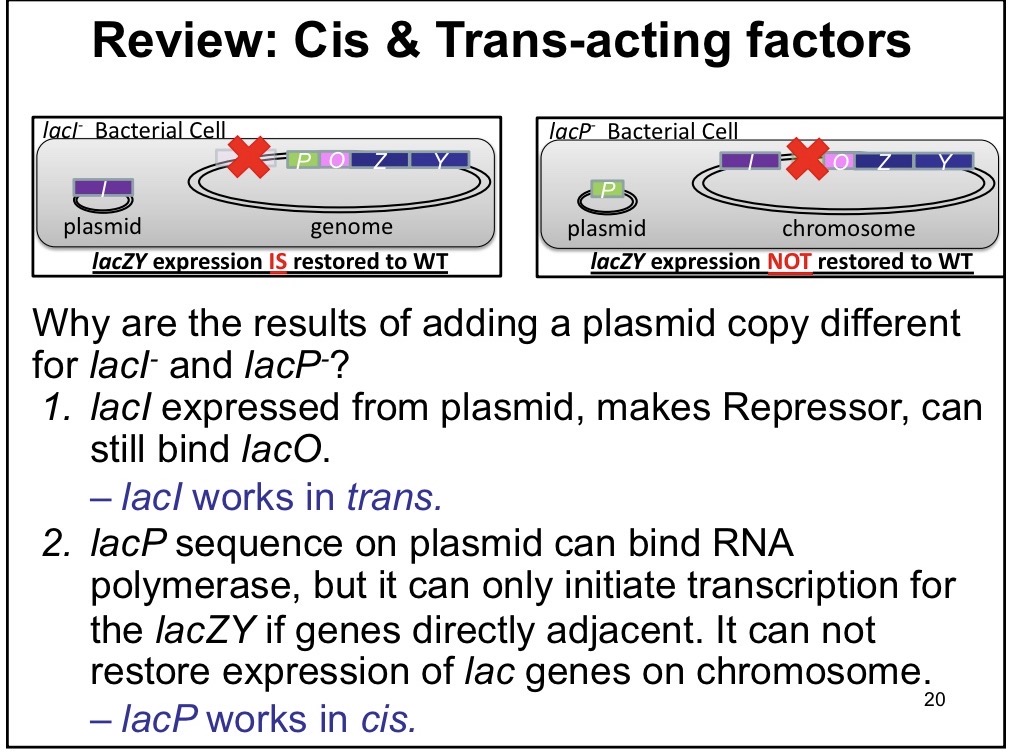
Trans
The protein made from the gene sequence is important, can diffuse
Ex: Regulatory proteins —> LacI acts in trans since it can be generated on the plasmid and still express wild type levels even if it can’t be generated on the chrommosome
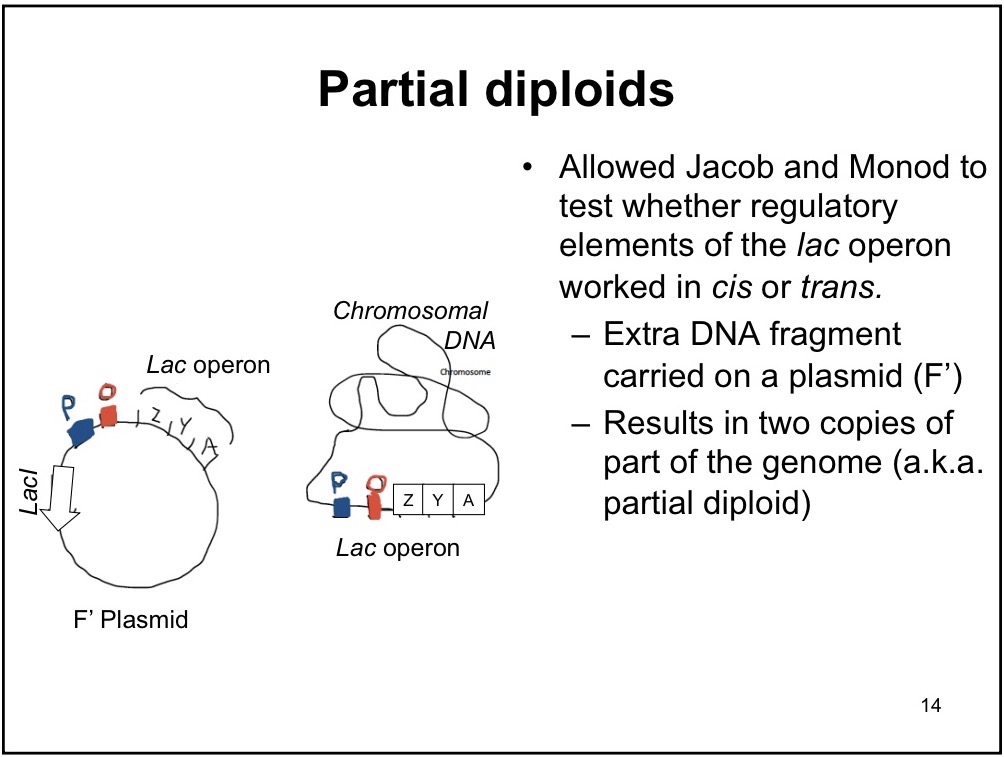
Partial Diploid
Adding gene to the bacteria via plasmids; one chromosomal DNA and one bacterial DNA
Genetic Code Redundancy
Triplet Code: 64 different 3 nucleotide “codons”
Not 64 tRNAs
Ex: For alanine there are 4 codons but not 4 tRNAs
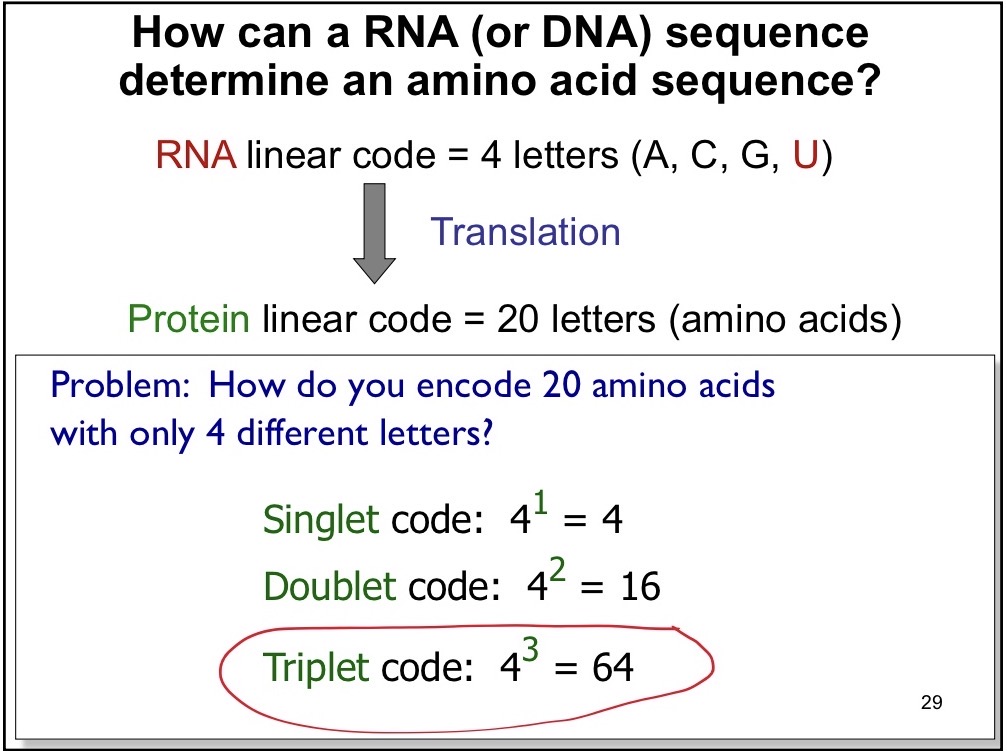

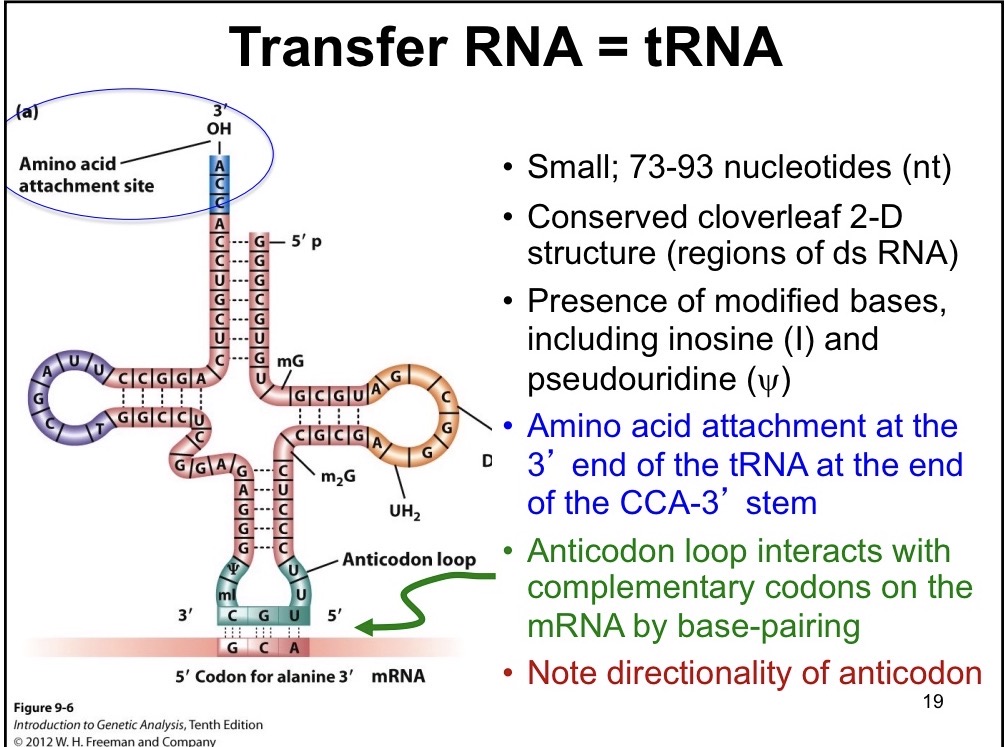
tRNA
Reads base pairs on mRNA
Small 73-93 nucleotides
Presence of modified bases such as inosine (I) and pseudouridine
Contains STEM loops
Amino acid attachment to 3’ CCA STEM
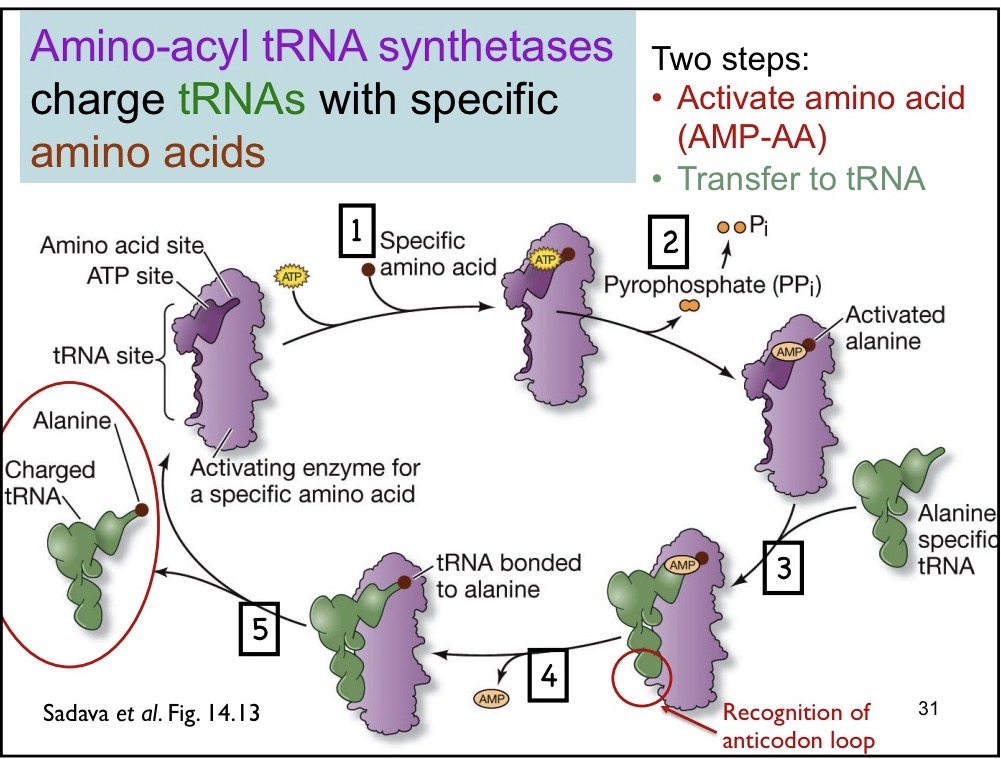
Aminoacyl-tRNA syntheases Draw
Charge tRNAs with specific amino acids
1) ATP and specific amino acid binds to tRNA
2) Amino acid specific tRNA binds to the syntheases
3) Recognizes the anticodon loop and binds to the amino acid (hydrogen bonding, van der Waals, hyrophoic)
4) Is released
Inosine
Deamination product of A, base-pairs with A, U, or C in the third (or wobble) position, on 5’ end of anticodon
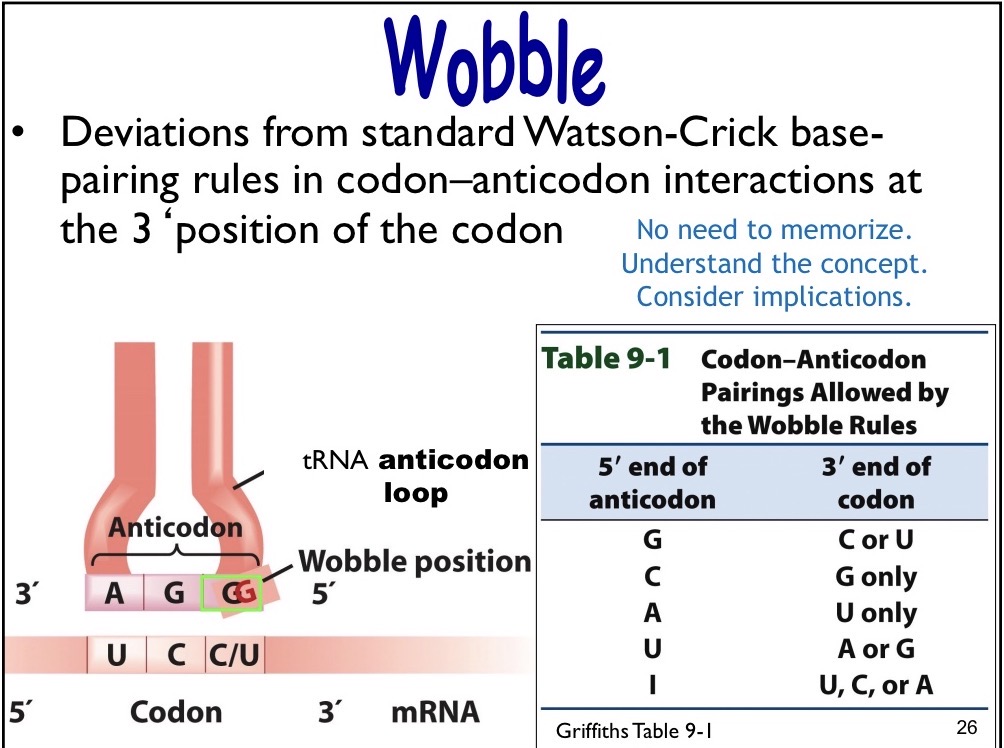
Aminoacyl t-RNA linkage
Carboxyl group of amino acid is linked to 3’ OH of tRNA
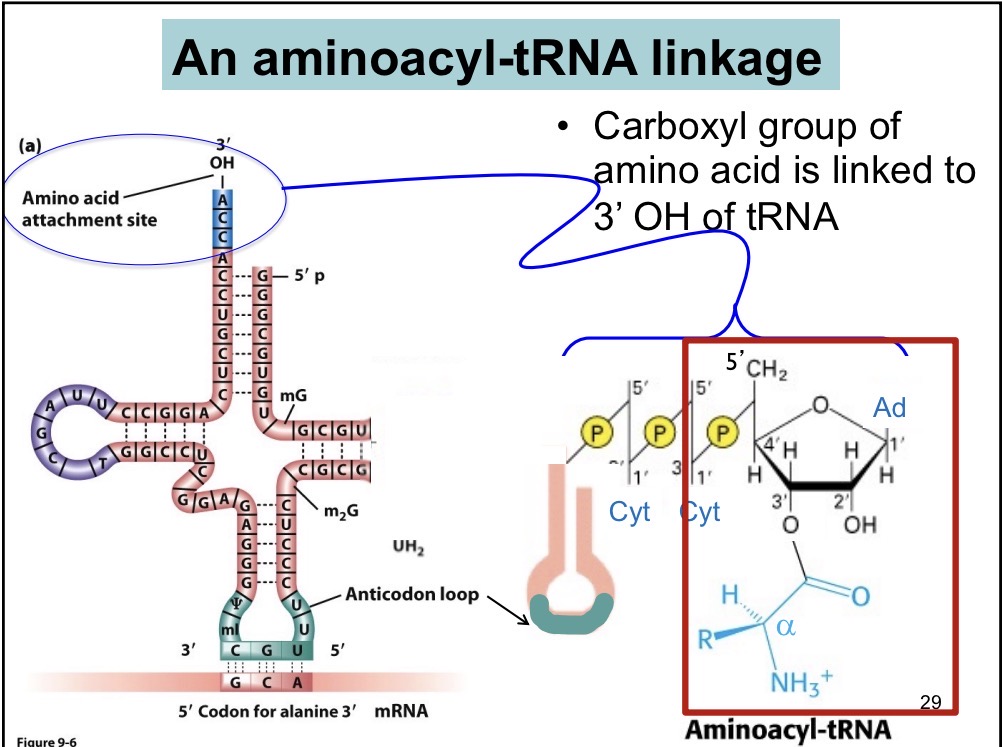
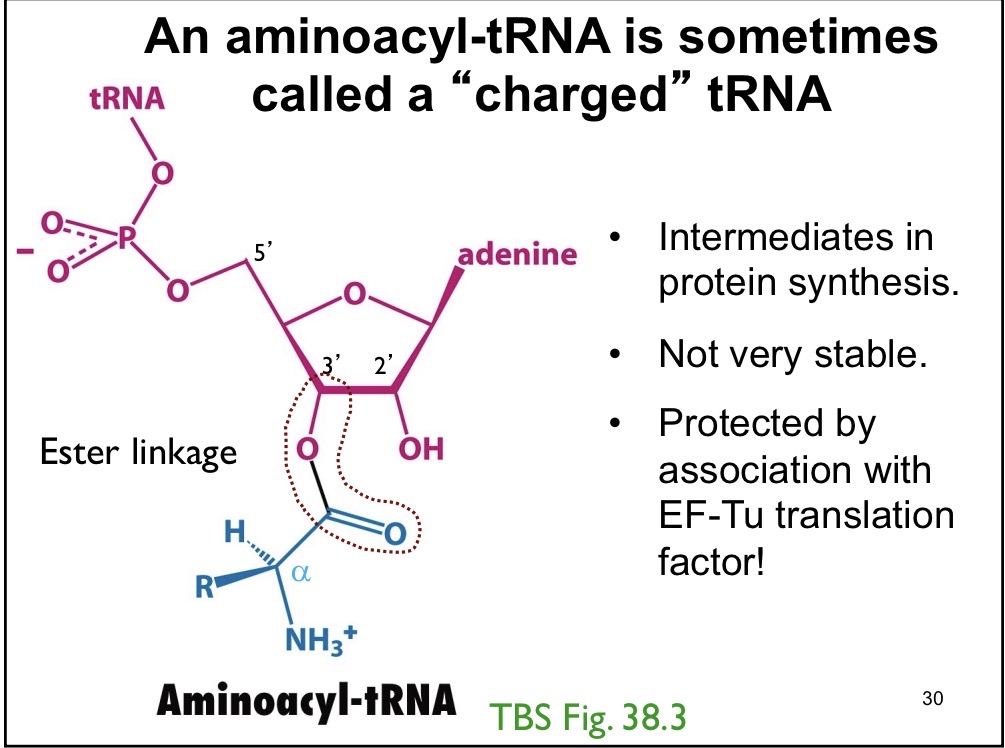
Aminoacyl t-RNA
Intermediates in protein synthesis, not very stable, protected by EF-TU translation factor
Ribosome
Bind mRNA and tRNA
3 Binding Sites of mRNA
A: Aminoacyl/Acceptor, new tRNA molecule comes in
P: Peptidyl, Growing peptide chain
E: Exit, Site where tRNA will leave
Direction of mRNA and Amino Acid Chain
mRNA is read 5’ —> 3’, amino acid chain is addd to the COO- end
Ribosome Composition
Large Subunit: 50s = 23S rRNA +5sRNA +31 proteins
Small Subunit: 30s = 16S rRNA +21 proteins
Need to know: Made of RNA and protein, protein is more structural
IF2: Initiation of Translation in Prokaryotes
1) mRNA binds to small 30S ribosomal subunit
2) Initiation Factor 2 (IF2) delivers a special tRNA charged with formylmethionine (fMET) to the P site (AUG)
3) Once tRNA-fMet is in place, the 50S subunit of ribosome can bind, GtP bound to IF2 is hydrolyzed and released
Shine-Dalgarno Sequence
Ribosome binding site on mRNA that 16s rRNA binds to. Positions the mRNA at the correct spot so that AUG is at the P site
Formyl Groups
Blocks the reactive N atom of the amine group to help with stability and folding (not used in eukaryotes)
G protein Translation Factors
IF2, EF-TU and EF-G
Ef-TU
GTP-bound forms a complexed with charged tRNA = tRNA-AA(amino acid), if codon/anticodon interaction between tRNA and A site is correct, GTP on EF-Tu is hydrolyzed and Ef-Tu is released (A site conformational change)
Peptide Bond Formation
Amino acid end of tRNAs in P and A sites are in close proximity and a peptide bond is formed, catalyzed by rRNA
Between C//O in the amino acid on tRNA at the P site and N on the amino acid in the A site
EF-G
Hydrolyzes GTP to GDP for translocation of ribosome toward 3’ end
Ribozyme
rRNA catalyzes peptide bond, not protein
Termination
Mediated by Release Factor proteins that interact with stop codons in the A site
Molecular mimicry
Release factor mimics tRNA molecule
Polysome
Couple transcription and translation in prokaryotes
Transcription in Eukaryotes
1) Separation of transcription and translation in eukaryotes with
2) More complex transcription regulation.
Three RNA polymerases
General Transcription Factors GTFs
cis acting elements are more varied
3) Extensive processing of mRNA
Nucleosomes (+Histone Types)
DNA (approx 160BP) is wrapped around a protein core of 8 histone molecules (4 different types H2A, H2B, H3, H4)
Levels of Chromatin Packing
DNA —> Nucleosome (10:1) —> Chromatin(50:1) —> Scaffold-associated chromatin (250:1)—> Condensed heterochromatin (5000:1)—> Compacted chromosome (8000:1)
Histone Structure
Histones interact with DNA minor groove; A/T base pairs are favored. Differ in N terminus tails
N-terminal tail
Targets of extensive modification such as acetylation and methylation (mostly at amino acids arginine/R and lysine)
Protein Classes and Histone Modifications
Writers: Covalently modify histone amino acids. Ex: Methyl transferases, acetyl transferases
Erasers: Restore modified to unmodified. Ex: Demthylases, deacetylases
Readers: Bind modified histone amino acids. Ex: Methyl readers and acetyl readers
Mammalian Gene Structure
Introns (leave) and Exons (stay)
Promoter: -25 TATA box, -200 nearby elements.
Enhancers: Distal (-10kb), proximal (-500kb), and downstream
Chromatin vs Non Chromatin DNA
Promoter Core: TATA and TSS Start site. Sequence needed for general transcription factors and RNA Polymerase to non-chromatin DNA
Promoter: Core promoter plus nearby elements. Minimal sequence required to recruit RNA polymerase to TSS in a cell (DNA packaged into DNA)
Enhancers
Work from far away to increase transcription, often involved in cell-type specific expression
Bacteria vs Eukaryotes
Bacteria: Ground state is on and promoters are DNA
Eukaryotes; Ground state is off and promoters are really protein-DNA complexes embedded in a chromatin environment
General Transcription Factors (GTFs)
GTFs and RNA polymerase assemble at the core promoter to form a Pre-Initiation Complex (PIC)
Need to Know: TFIIB, TFIID, TFIIH, pol II
Assembly of Pre-Initiation Complex
1) TATA-box Binding Protein (TBP) as part of TFIID; the anchor
2) TFIIB
3) RNA Polymerase II (binds to TFIIB)
4) TFIIH
TFIIH
1) Acts as a helicase
2) Phosphorylation of the carboxy terminal (CTD) RNA POLY II by a TFIIH kinase
Histone lysine acetylation
Acetylation neutralizes the positive charge of DNA, loosening the interaction between negatively charged DNA
3 Elements Required for In Vivo Transcription
1) Chromatin Remodeler to Reposition Nucleosomes
2) Formation of transcription initiation complex
3) Complexes that mediate these interactions
Nucleosome Remodeling
Gene activator protein binds to TATA box, bringing in histone acetylase to loosen the structure and chromatin remodeling complex to move histones
Chromatin Remodeler
Requires ATP
Mediator
1) Is required for transcription of most genes in cells
2) Interacts with RNA Polymerase, other GTFs and activator proteins bound to far-away enhancers
Activators and Repressors (Eukaryotic)
Bind to enhancers or nearby elements
Activators
two domains: DNA binding and activation domain. Targets of activation domains are recruited to the promoter by protein-protein interactions
Co-Activators
1) General transcription factors
2) Histone Code Writers
3) Chromatin remodoling complexes
Yeast Case Study
GAL 7, GAL 10, and GAL 1 are all coordinately regulated by a transcriptional activator Gal4.
Yeast Case Study: ± galactose
In the absence of galactose Gal80 binds to Gal4. In the presence of galactose, galactose binds to Gal3 which binds to Gal80 in the cytoplasm. This prevents Gal80 from getting into the nucleus, Gal 4 activation domain, binds a histone acetyltransferase coactivator complex (SAGA) and a mediator which recruit GTFs and RNA promoter.
Combinations of Activators
Different genes require different combinations of activators, can work cooperatively. Activation of the same gene at different times and places also involves different combinations of activators
DNA Looping
Transcriptional regulator proteins that bind to far-away enhancers are brought into the proximity of the promoter by DNA looping and interactions with the Meidator
C-Terminal Domain
CTD provides a binding site for factors involved in processing the transcript and regulating the activity of transcribed chromatin
Histone Deacetylases
Repressors can recruit co-repressors with histone deacetylase (HDAC) activity, similarly to how co-activators regruit Histone Acetyltransferase (HAT)
Mig1
Glucose availability induces the binding of a repressor called Mig 1 (galactose preferred source). Mig1 binds to repressor and recruits a co-repressor, HDAC
Heterochromatin
Constitutive heterochromatin form at all telomeres and sequences that flanked centromeres.
Fly Eye Experiment
White gene with no mutations: red eye color
White gene with mutations: white eye
Chromatin breaks off and rearranges so that gene is near heterochromatin which doesn’t get transcribed
HP1
HP1 is a protein that binds to methylated histone tails and recruits a histone methyltransferase
Barrier
Between heterochromatin and euchromatin, competing between whether they are methylated or acetylated
X Chromosome
Two versions of a gene on each chromosome, choice of X chromosome of X for inactivation occurs early in development
Genetics
DNA changed but transcription still occurs: ex mutation
Epigenetics
DNA unchanged but transcription doesn’t occur
Epigenetics
DNA unchanged but transcription doesn’t occur
Chromatin Modifications Associated with Heterochromatin and Epigenetic Silencing
Methylation of H3K9 (lysine at position 9) by HMT
Methylation at C5 of Cytosines by DNMT
mRNA processing
1) mRNA capping
2) Cleavage and polyadenylation (“poly A”) addition to generate the 3’ end
3) Intron splicing (removal of intron sequences)
Where is mRNA processed
Nucleus
5’ mrNA cap
RNA Pol II has a “stall factor” shortly after initiation to add the 5’ cap
1) Removal of the terminal Pi from the 5’ end
2) A guanine cap is attached to the 5; end using GTP (5’ to 5’ linkage)
3) Methylation of guanine