BIO 81 Exam 4 Review
1/117
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
118 Terms
A virus achieves host specificity through _____ on the surface of the capsid or envelope, which bind to specific receptor molecules on the surface of the host cell.
Proteins
A virus identifies its target host cell through a "lock and key" mechanism. The "key" consists of specific surface proteins (often called spikes or glycoproteins) located on the viral capsid or envelope.
Recognition: These viral proteins are shaped to fit perfectly into specific receptor molecules (which are also typically proteins) on the host cell's membrane.
Specificity: Because this binding is so precise, a virus can usually only infect a specific range of hosts or specific cell types within a host (such as how the flu virus targets respiratory cells).
Other components: While envelopes contain lipids and some proteins are glycosylated (have sugars attached), it is the specific amino acid structure of the proteins that dictates the binding affinity and host specificity. Nucleic acids are tucked safely inside the virus and do not participate in the initial attachment process.
What causes antigenic drift to occur in viral infections?
gradual changes in the viral genome due to replication errors
Why arent viruses considered to be alive?
Viruses cannot carry out metabolic reactions on their own.
Define an operon and its three main components.
An operon is a cluster of functionally related genes under coordinated control by a single "on-off switch". It includes:
Operator: The DNA segment that acts as the regulatory switch, usually within the promoter.
Promoter: Where RNA polymerase binds.
Genes: The actual DNA sequence for the enzymes being controlled.
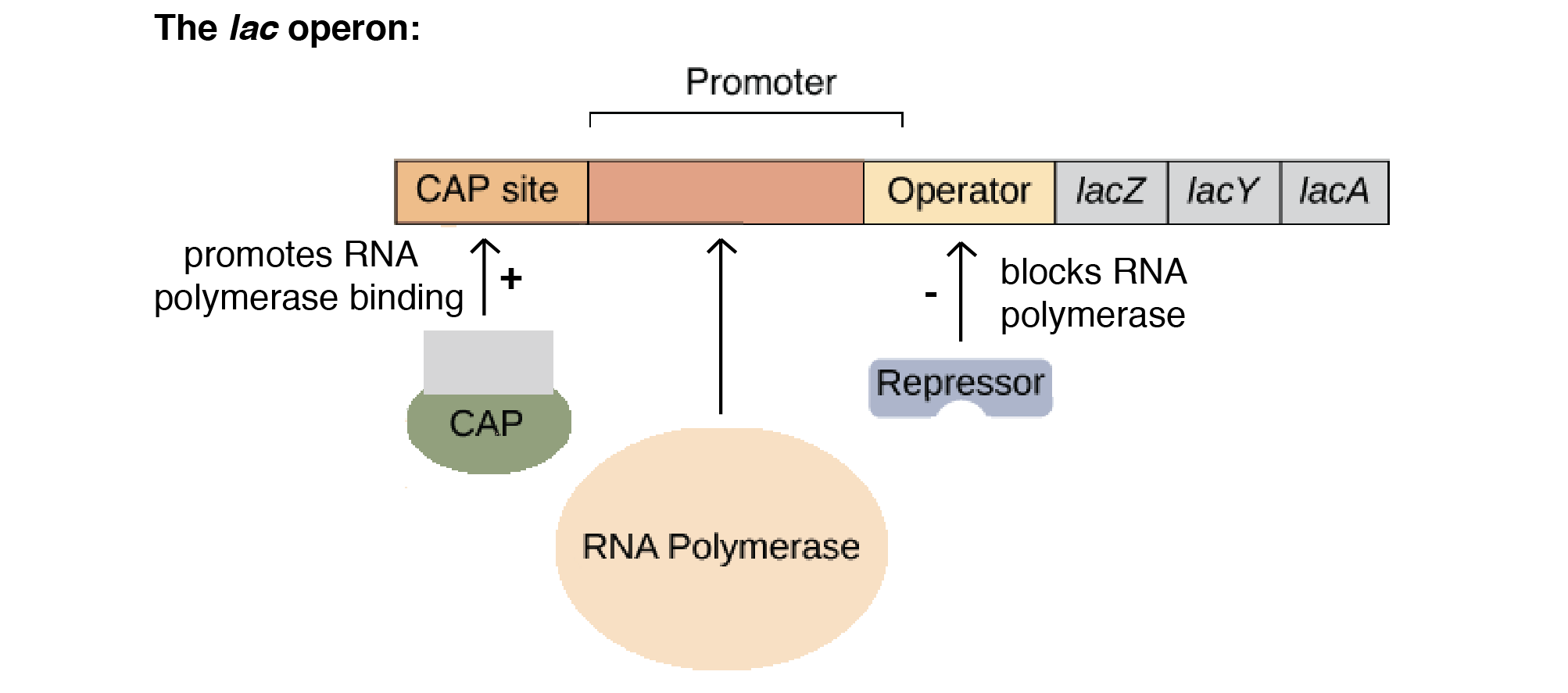
Distinguish between a repressible operon and an inducible operon.
Repressible (e.g., trp operon): Usually ON; binding of a repressor shuts off transcription. Functions in anabolic (synthetic) pathways.
Inducible (e.g., lac operon): Usually OFF; an inducer molecule inactivates the repressor to turn transcription on. Functions in catabolic (breakdown) pathways.
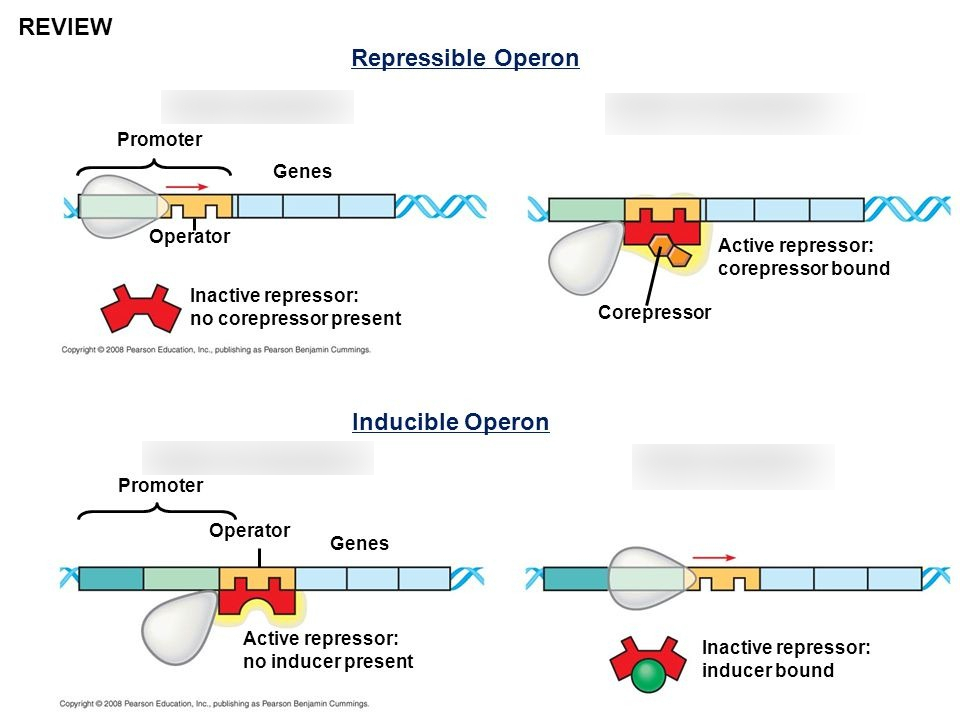
How does E. coli prioritize glucose over lactose?
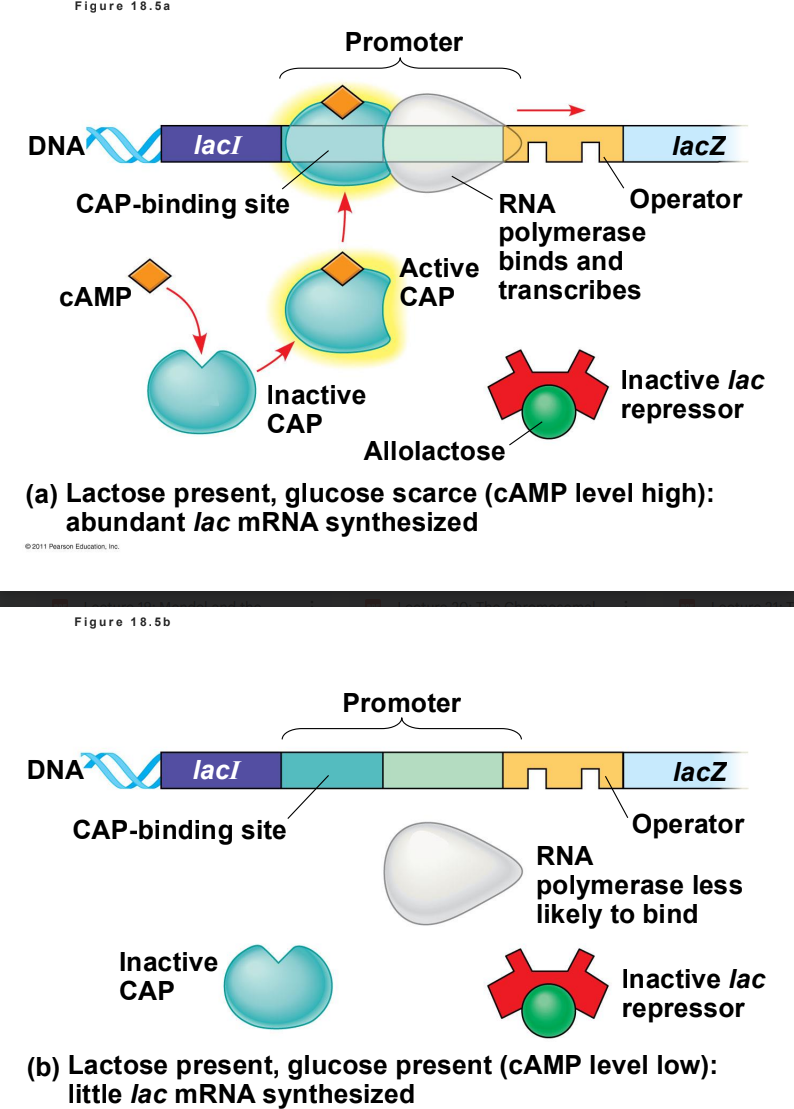
Through Positive Gene Regulation using Catabolite Activator Protein (CAP). When glucose is scarce, cAMP levels rise and bind to CAP. Activated CAP attaches to the lac promoter, increasing RNA polymerase affinity and accelerating transcription.
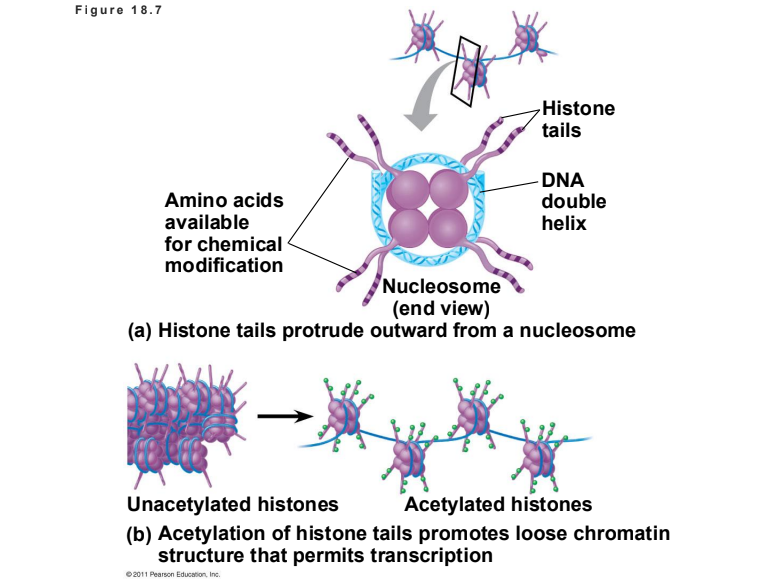
Explain how Histone Acetylation and DNA Methylation affect gene expression.
Histone Acetylation: Acetyl groups attach to lysines in histone tails, loosening chromatin structure and promoting transcription.
DNA Methylation: Addition of methyl groups to DNA bases (usually cytosine) is associated with reduced transcription and long-term gene inactivation.
What is epigenetic inheritance?
The inheritance of traits transmitted by mechanisms that do not involve the nucleotide sequence itself (e.g., chromatin modifications passed to future generations).
What are distal control elements and their role in transcription?
Distal control elements, called enhancers, are DNA sequences far from the gene. Activator proteins bind to these enhancers and interact with mediator proteins and general transcription factors to form the transcription initiation complex.
How can one gene produce multiple different proteins?
Through Alternative RNA Splicing, where different mRNA molecules are produced from the same primary transcript depending on which segments are treated as exons and which as introns.
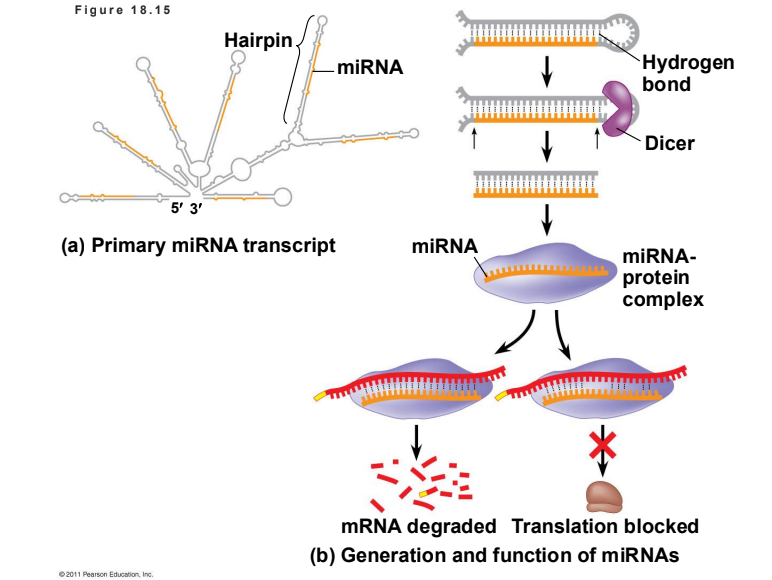
What is the role of MicroRNAs (miRNAs) in gene regulation?
miRNAs are small single-stranded RNA molecules that bind to complementary sequences in mRNA. They can either degrade the target mRNA or block its translation. This phenomenon is called RNA interference (RNAi).
What is the difference between cell determination and differentiation?
Determination: The process that commits a cell to its final fate (e.g., a cell becoming a myoblast).
Differentiation: The actual structural and functional specialization marked by the production of tissue-specific proteins.
Explain the Morphogen Gradient Hypothesis using the Bicoid gene.
Morphogens are substances whose gradients establish an embryo's axes. In Drosophila, the Bicoid protein is concentrated at the anterior end; this gradient determines the "head" end of the fly.
Define Proto-oncogenes and how they become Oncogenes.
Proto-oncogenes: Normal genes responsible for cell growth and division.
Oncogenes: Cancer-causing genes.
Conversion: Can occur via DNA translocation, gene amplification, or point mutations that lead to overexpressed or hyperactive proteins.
What is the "Guardian of the Genome" and why is it important?
The p53 gene is a tumor-suppressor gene that inhibits the cell cycle if DNA is damaged. Mutations in p53 prevent suppression of the cell cycle, allowing mutations to accumulate and leading to cancer.
Why does the incidence of cancer increase with age?
Full-fledged cancer generally requires multiple mutations. Usually, this involves at least one active oncogene and the mutation of several tumor-suppressor genes (like APC, ras, and p53).
Define feedback inhibition vs. gene regulation in the context of tryptophan. What is the corepressor?
Feedback Inhibition: Tryptophan (the product) inhibits the activity of the first enzyme in the pathway for immediate response.
Gene Regulation: Tryptophan (the corepressor) binds to the inactive repressor to activate it, shutting down the production of new enzymes.
Name the inducer and the three enzymes produced by the lac operon .
Inducer: Allolactose (an isomer of lactose).
Enzymes: beta-Galactosidase (breaks down lactose), Permease (transports lactose into the cell), and Transacetylase.
Beyond acetylation, what other two chemical modifications affect chromatin, and what is the Histone Code Hypothesis?
Phosphorylation: Adding phosphate groups next to a methylated amino acid loosens chromatin.
Methylation: Can condense chromatin (except when paired with phosphorylation).
Histone Code Hypothesis: Proposes that specific combinations and the order of modifications determine chromatin configuration and transcription levels.
Describe the role of DNA-bending proteins and Mediator proteins .
Activators bind to distal enhancers; a DNA-bending protein brings the bound activators closer to the promoter. Mediator proteins then interact with these activators and general transcription factors to help RNA polymerase II bind to the TATA box
How does a cell "mark" a protein for destruction?
The cell attaches a small protein called Ubiquitin to the target protein. Giant protein complexes called Proteasomes recognize the ubiquitin tag, unfold the protein, and degrade it into fragments
Distinguish between miRNAs, siRNAs, and piRNAs
miRNAs/siRNAs: Both lead to RNA interference (RNAi) by degrading mRNA or blocking translation. miRNA uses the Dicer enzyme for processing .
piRNAs (Piwi-associated): Induce heterochromatin to block "parasitic" DNA elements called transposons.
Explain why MyoD is considered a "master regulatory gene"
It is a transcription factor that commits an embryonic precursor cell to becoming a myoblast (determination) . It binds to enhancers of other muscle-specific genes and stimulates its own transcription to maintain the "determined" state
List the three ways a proto-oncogene becomes an oncogene
Translocation/Transposition: Moving the gene near a highly active promoter.
Gene Amplification: Increasing the number of copies of the gene.
Point Mutation: Creating a hyperactive or degradation-resistant protein
What are the specific genetic steps (in order) for colon cancer development?
Loss of tumor-suppressor gene APC (leads to a small polyp).
Activation of ras oncogene (leads to a larger adenoma).
Loss of tumor-suppressor gene DCC.
Loss of tumor-suppressor gene p53 (leads to a malignant carcinoma).
Draw positie gene regulation in both eukaryotes and prokaryotes.
Positive gene regulation is a mechanism where regulatory proteins, known as activators, bind to specific DNA sequences to promote and increase the transcription of a gene into messenger RNA (mRNA). Without this active protein, the target genes typically remain unexpressed or transcribed at very low basal levels
1. Prokaryotic Activators (e.g., The Lac Operon)
In bacteria like E.coli, when glucose is scarce but lactose is available, the cell needs to break down lactose for energy.
The Process: Low glucose levels lead to an accumulation of cyclic AMP (cAMP). cAMP binds to the Catabolite Activator Protein (CAP), activating it.
The Result: The cAMP-CAP complex binds to the promoter region of DNA, physically helping RNA polymerase attach. This positively regulates the operon, causing the cell to rapidly produce the enzymes needed to digest lactose.
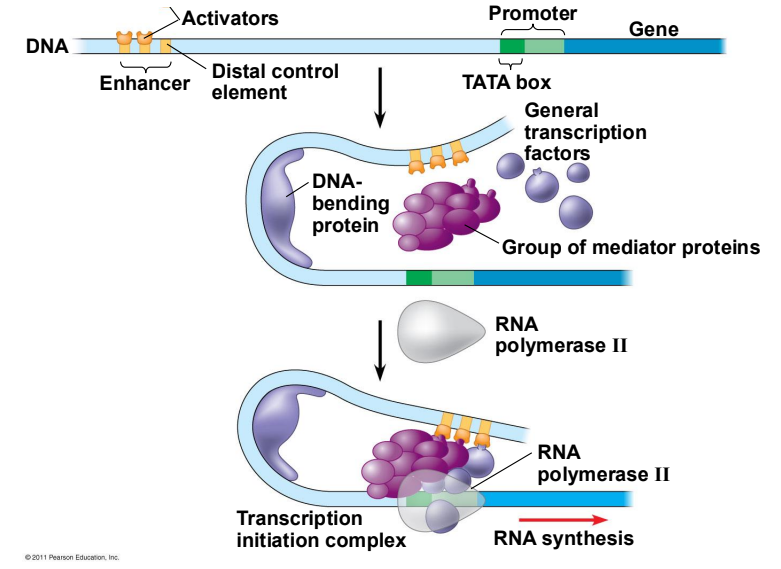
2. Eukaryotic Transcription Factors
In multicellular organisms, positive regulation is the dominant form of control, as the default state of most genes is "off".
Activators & Enhancers: Activator proteins bind to specific DNA regions called enhancers.
Mediator Proteins: These activators interact with mediator proteins and general transcription factors to assemble a functional transcription complex.
The Result: This coordinated binding pulls RNA polymerase to the promoter, significantly upregulating the rate of transcription.
Why It Matters
Positive regulation acts much like a volume knob that turns up gene production. It allows organisms to adapt to stress, grow, and differentiate by only producing specific proteins exactly when they are required.
Draw and explain transcription initiation.
The first and primary regulatory step of gene expression. It begins when RNA polymerase and accessory proteins bind to a specific DNA sequence called a promoter, allowing the enzyme to unzip the double helix and prepare to synthesize a complementary RNA strand
1. Promoter Recognition
Transcription starts upstream of a gene at a specific DNA region known as the promoter. Prokaryotes: The main transcription enzyme, RNA polymerase, binds directly to specific consensus sequences (like the -35 and -10 regions) using a helper subunit called a sigma factor.
Eukaryotes: The process requires a more complex assembly. General transcription factors must first bind to promoter elements, such as the TATA box, to recognize the start site.
2. Formation of the Open Complex
Once bound to the promoter, the enzyme must separate the two DNA strands to expose the template strand.
The RNA polymerase causes the DNA double helix to locally unwind or "melt".
This creates a small open region called a transcription bubble.
The open complex gives RNA polymerase direct access to read the single-stranded DNA template.
3. Initial RNA Synthesis
Once the transcription bubble is formed, the enzyme begins to build a complementary RNA chain.
RNA polymerase synthesizes RNA in the \(5'\) to \(3'\) direction.
It matches free-floating RNA nucleotides to the complementary DNA strand (e.g., matching Adenine with Uracil, and Cytosine with Guanine).
4. Promoter Clearance
During the initial stages of synthesis, the RNA polymerase may release the short RNA transcript and attempt the process again—a phase known as abortive initiation. Once the RNA chain reaches a sufficient length (usually around 10-12 nucleotides), the polymerase successfully breaks its initial contacts with the promoter and clears the starting region. This marks the end of initiation and the start of elongation
Fertilization
The formation of a diploid zygote from a haploid egg and sperm
Polyspermy
The entry of multiple sperm nuclei into a single egg, which is prevented by changes at the egg surface to ensure normal genomic numbers
Acrosomal Reaction
Triggered when sperm meets the egg; the acrosome at the sperm's tip releases hydrolytic enzymes that digest the egg’s surrounding protective layer
Fast Block to Polyspermy
Immediately following gamete contact/fusion, the egg cell membrane depolarizes, preventing additional sperm from fusing
Cortical Reaction & Slow Block to Polyspermy
A calcium-mediated reaction where cortical granules beneath the egg membrane release contents to lift the vitelline layer, forming a hardened fertilization envelope
Egg Activation
A sharp rise in cytosolic Ca2+ concentration triggers a rapid increase in cellular respiration and protein synthesis, activating the egg's metabolism using pre-existing maternal mRNAs and proteins
Mammalian Fertilization Characteristics
Internal fertilization where sperm must cross a layer of follicle cells and the zona pellucida (extracellular matrix). The overall process is slow; the first cleavage occurs 12–36 hours after sperm binding
Cleavage
A period of rapid cell division without cell growth that follows fertilization, partitioning the huge single-celled zygote cytoplasm into many smaller cells called blastomeres
Blastula & Blastocoel
Blastula: A hollow ball of cells produced by cleavage (termed blastocyst in mammals).
Blastocoel: The fluid-filled inner cavity of the blastula
Asymmetric Cleavage in Frogs
Cleavage is uneven due to heavy yolk concentration at the vegetal pole (more yolk, divides slower/larger cells) compared to the animal pole (less yolk, divides faster/smaller cells)
Regulation of Early Cleavage
Initial metabolic control and structural setup are fully programmed by maternal RNA and proteins deposited into the egg during oogenesis
Morphogenesis
The overarching process by which an organism takes shape and cells occupy their appropriate physical locations, primarily driven by cell movement, shape changes, and survival changes
Gastrulation
The morphogenetic process that rearranges the single-layered blastula into a three-layered embryo called a gastrula
Ectoderm Derivatives
Forms the outer layer of the embryo: epidermis of the skin (and derivatives), nervous and sensory systems, pituitary gland, adrenal medulla, jaws, teeth, and germ cells
Mesoderm Derivatives
Fills the middle space: skeletal and muscular systems, circulatory and lymphatic systems, excretory and reproductive systems (except germ cells), dermis of skin, and the adrenal cortex.
Endoderm Derivatives
Lines the innermost areas: epithelial lining of the digestive tract and its associated organs (liver, pancreas), lining of respiratory, excretory, and reproductive tracts, plus the thymus, thyroid, and parathyroid glands
Human Blastocyst Anatomy
Inner Cell Mass: The inner cluster of cells that will actually develop into the embryo.
Trophoblast: The outer epithelial layer that initiates uterine implantation (~7 days post-fertilization) and forms extraembryonic membranes rather than the embryo itself
Extraembryonic Membranes of Amniotes
Four specialized membranes that protect the embryo from drying out, enabling reproduction on dry land:
Amnion: Encloses the protective amniotic fluid.
Chorion: Functions primarily in gas exchange.
Yolk Sac: Encloses nutrient yolk.
Allantois: Disposes of nitrogenous waste and assists gas exchange
Organogenesis
The stage of development where various regions of the three embryonic germ layers specialize into rudimentary organs
Induction
A key developmental process where a group of cells or a tissue secretes signaling molecules that cause a structural or fate change in nearby target cells
Neurulation Process
The notochord (formed from dorsal mesoderm) secretes inductive signals causing the overlying ectoderm to form the neural plate, which curves inward to become the neural tube (future central nervous system: brain and spinal cord)
Neural Crest Cells
Unique cells that develop along the border of the closing neural tube and migrate long distances through the embryo to form diverse structures like nerves, parts of teeth, and skull bones
Somites
Blocks of mesoderm lateral to the notochord that separate into migrating mesenchyme cells, helping form vertebrae, ribs, and associated skeletal muscle
Cytoskeleton Roles in Morphogenesis
Microtubules lengthen cells to change their shape (e.g., during neural plate formation), while microfilaments (actin) contract at one end to wedge/bend cells into tubes
Convergent Extension
A morphogenetic movement where a sheet of cells rearranges itself by elongating and crawling between each other, resulting in a tissue structure that becomes significantly longer and narrower
Cell Adhesion Molecules (CAMs) & ECM
Transmembrane glycoproteins (CAMs) and the extracellular matrix (ECM) provide the physical tracking, anchors, and chemical signals necessary for directed cell migration
Apoptosis in Development
Programmed cell death used to remove redundant or temporary structures (e.g., eliminating extra embryonic neurons or absorbing a tadpole’s tail during metamorphosis)
Determination vs. Differentiation
Determination: The process by which a cell becomes irreversibly committed to a specific developmental pathway.
Differentiation: The subsequent physical specialization in cellular structure and function.
Frog Axis Determination
Anterior-Posterior (A-P): Set during oogenesis by animal-vegetal cytoplasm polarity.
Dorsal-Ventral (D-V): Established at fertilization; sperm entry triggers cortical rotation, moving vegetal cortex molecules to expose the gray crescent, marking the future dorsal side
Totipotency
The developmental capability of a cell to give rise to all possible distinct cell types of an organism. In mammals, embryonic cells remain totipotent until the 8-cell stage.
Spemann and Mangold's Organizer Experiment and why his expeiment was so revolutionary.
They transplanted the dorsal lip of the blastopore from one early gastrula into a host embryo. The dorsal lip acted as an "organizer," inducing surrounding host tissues to clear a second gastrulation and build a full, mirror-image secondary body plan
It proved that specific tissues dictate the directions and patterns of surrounding tissues through induction.
Limb Buds & Positional Information
Vertebrate limbs start as limb buds that read chemical coordinate systems along three axes: proximal-distal, anterior-posterior, and dorsal-ventral.
AER vs. ZPA
Apical Ectodermal Ridge (AER): Thickened ectoderm at the tip of the limb bud regulating outgrowth along the proximal-distal axis.
Zone of Polarizing Activity (ZPA): Mesodermal tissue at the posterior base of the bud that secretes Sonic hedgehog protein to pattern the anterior-posterior axis (e.g., digit arrangement)
Chronological path from fertilization
Fertilization → Fast/Slow Blocks → Cleavage → Gastrulation → Organogenesis/Neurulation
Explain how the bicoid determines the head structures and the significance of that research
Describe the acrosomal and cortical reactions
Describe two different ways to prevent polyspermy
Describe the process of neurulation in detail
Define protoconcogenes, and tumor suppressor genes and how they can function to create tumors
How does the rising calcium in the cytosol affect the egg activation rates?
Compare and contrast the lytic and lysogenic cycles.
How does a vaccine work to help you fight off viruses?
What is the difference between embryonic and adult stems cells?
Explain how size and charge of the particle (DNA or protein) affects the results of gel electrophoresis
Explain the process of a PCR
Describe pre-transcriptional, post-transcriptional and post-translational mechanisms of gene regulation
List three noncoding RNAs and their function
What is epigenetic inheritance? Give an example
Explain the process of CRISPR
Describe how DNA sequencing works
What are forensic applications of biotechnology. Explain
What is the basic definition of a virus?
An infectious particle consisting of genes (nucleic acid) packaged in a protein coat
Why are viruses described as leading a "borrowed life"?
They cannot reproduce or carry out metabolism outside of a host cell, existing in a "shady area" between life-forms and chemicals
How did Wendell Stanley confirm the existence of viruses?
Crystallizing the tobacco mosaic virus (TMV)
In 1935, biochemist Wendell M. Stanley confirmed the existence of viruses as distinct, tangible, and manipulatable particles by isolating the tobacco mosaic virus (TMV) in pure, crystalline form. He demonstrated that these crystals, despite behaving like inanimate chemicals, retained the ability to infect plants, effectively blurring the line between living organisms and non-living molecules.
Key Aspects of Stanley's Confirmation:
Isolation and Crystallization: Using refined chemical techniques, Stanley processed tons of infected tobacco leaves to isolate the virus, producing needle-shaped crystals.
Demonstrated Infectivity: He showed that these crystals, when dissolved and applied to healthy plants, caused the disease, proving they were not just a chemical byproduct but the active, infectious agent.
Proof of Re-infection: Even after repeating the crystallization process 15 times, the substance remained highly infectious, demonstrating it was a stable, pure particle rather than a contaminant.
Composition Analysis: Further studies determined the particles were composed of protein and ribonucleic acid (RNA), revealing the molecular nature of viruses
What does it mean that viruses are "obligate intracellular parasites"?
They can only replicate within a host cell.
What is a "host range"?
The limited number of host species or cell types a particular virus can infect
What are the two essential components of all viruses?
A viral genome (DNA or RNA) and a protein capsid
What are the protein subunits that build a capsid?
Capsomeres
What is a viral envelope and where does it come from?
A membranous outer layer derived from the host cell's plasma membrane; it contains both viral and host molecules
How are viral genomes classified?
They can be double-stranded DNA, single-stranded DNA, double-stranded RNA, or single-stranded RNA
What is the specific structure of a bacteriophage (phage)?
They typically have an elongated capsid head enclosing DNA and a protein tail piece used to inject DNA into the host
What is the main difference between the lytic and lysogenic cycles?
The lytic cycle culminates in the death of the host cell by lysing (breaking open) its wall to release new viruses.
The lysogenic cycle replicates the phage genome without destroying the host by incorporating viral DNA into the host chromosome.
What is a "prophage"?
Integrated viral DNA within a bacterial chromosome during the lysogenic cycle
What is a "provirus"?
Viral DNA that is permanently integrated into an animal host's genome (e.g., HIV); unlike a prophage, it never leaves the host genome
How do retroviruses like HIV replicate?
They use the enzyme reverse transcriptase to copy their RNA genome into DNA
How do restriction enzymes protect bacteria?
They identify and cut up foreign (phage) DNA
What is the CRISPR-Cas system?
A bacterial immune system where "spacer" sequences from previous phage infections are used by Cas proteins to recognize and cut invading phage DNA
How can viruses cause cancer?
Some carry oncogenes that promote cell division, or they may integrate into a host's proto-oncogene, disrupting its normal function and leading to tumor growth
Distinguish between horizontal and vertical transmission in plants.
Horizontal transmission occurs when a virus enters through damaged cell walls; vertical transmission is when a plant inherits a virus from a parent
What are prions?
Infectious, incorrectly folded proteins that cause degenerative brain diseases (e.g., mad cow disease) by converting normal proteins into the misfolded version.
Why are antibiotics ineffective against viral infections?
Antibiotics target bacterial processes; viral infections are best prevented by vaccines or managed with antiviral drugs that inhibit viral synthesis/assembly
What is the difference between an epidemic and a pandemic?
An epidemic is a localized general outbreak; a pandemic is a global epidemic.
What are the three main causes for the emergence of new viral diseases?
High mutation rates in RNA viruses, dissemination from isolated populations, and the spread of viruses from animals to humans