MAME Exam 2
1/86
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
87 Terms
Microbial Diversity
The variety of microorganisms present in a specific environment, including bacteria, archaea, fungi, and viruses.
Biogeochemical Cycles
Natural processes that recycle nutrients in various chemical forms from the environment to living organisms and back.
Nutrient Cycling
The movement and exchange of organic and inorganic matter back into the production of living matter.
Decomposition
The process by which organic substances are broken down into simpler organic or inorganic matter, often facilitated by microorganisms.
Microbial Interactions
The various ways in which microorganisms interact with each other and their environment, including competition, predation, and symbiosis.
Symbiosis
A close and often long-term interaction between two different biological species, which can be mutualistic, commensalistic, or parasitic.
Nitrogen Fixation
The conversion of atmospheric nitrogen into a form usable by living organisms, primarily carried out by certain bacteria.
Phytoplankton
Microscopic plants that live in aquatic environments and are a crucial part of the food web, contributing to primary production.
Microbial Mats
Layered microbial communities that can be found in various environments, often consisting of different types of microorganisms.
Biofilms
Complex communities of microorganisms that adhere to surfaces and are embedded in a self-produced matrix of extracellular polymeric substances.
Metagenomics
The study of genetic material recovered directly from environmental samples, allowing for the analysis of microbial communities without the need for culturing.
Ecosystem Services
The benefits provided by ecosystems to humans, including nutrient cycling, waste decomposition, and climate regulation.
Trophic Levels
The hierarchical levels in an ecosystem, defined by the position of organisms in the food chain, from producers to various levels of consumers.
Carbon Cycle
The series of processes by which carbon compounds are interconverted in the environment, involving photosynthesis, respiration, and decomposition.
Anaerobic Processes
Biological processes that occur in the absence of oxygen, often involving fermentation or sulfate reduction.
Microbial Ecology
The study of the interactions between microorganisms and their environment, including their roles in nutrient cycling and ecosystem functioning.
gram staining
A process by which components of bacterial cell walls are bound to Gram's stain. Depending on the amount of peptidoglycan in their cell walls, bacteria stain differently and are classified as Gram-negative or Gram-positive.
Barcode of Life Project
uses sequences of the COI gene and other genes found in mitochondria to identify and classify invertebrates and vertebrates
what is the most commonly used gene for prokaryotes?
16S rRNA
what is the most commonly used gene for eukaryotic microbes?
18S rRNA
cultivation-independent methods
approaches that do not rely on cultivation
tag
refers to an added oligonucleotide sequence used to label and later separate sequences from different samples that are analyzed together
what is the current read length of today's sequencers?
few hundred base pairs
how many base pairs in a complete 16S rRNA gene?
1500
species richness
number of OTUs in a community
clade
closely related group of organisms descended from a common acestor, meaning it is monophyletic
strain
word used to describe different isolates or new cultures of the same microbe
eveness
takes in account the number of individuals per OTU
alpha diversity
diversity within a habitat
gamma diversity
total diversity over a landscape with several habitats
beta diversity
thought of as "species turnover" or the differences among habitats over a landscape
microbial communities are very diverse but they are not very ____.
even
OTUs
cluster of DNA sequences that share 97% or more similarity
one way to "examine" new microbes is to subject to microscopy alone or a biochemical test followed by microscopy. Which group did he say was examined using Gram staining?
bacteria and archaea
what is an "amplicon"?
DNA produced by PCR (in vitro)
which gene is the phylogenetic marker gene that is the target of PCR for determining bacterial community structure?
16S (SSU) rRNA gene
which gene is a protein-encoding gene, whereas the others in the list are simply part of the tRNA, rRNA operon?
mtCO1
what is the proper "cutoff" for DNA sequence similarity to group two organisms' sequences into the same "species" or better, "OTU"?
97%
when we do amplicon library sequencing, what method is used regardless if the subseqent sequencing method is done?
PCR
why is "species" a bad word to use when describing different taxonomic groups of bacteria or archaea?
the classical definiton of species is a collection of individuals that can mate and reproduce, so this word doesn't mean anything with asexually reproducing organisms (most microbes)
what is PCR?
makes a lot of copies of particular fragment of DNA that you're interested in
how many base pairs on average does a gene have?
1000
denaturation
separates two strands of DNA at 96C
what is the proper cutoff for clumping DNA sequences into an "OTU" designated as an ASV/ESV taxonomic unit?
100%
selection is a deterministic process in that the success of a species is set strictly by how its biochemical, physiological, and ecological capabilities match up with the habitat. In contrast with selection, _____ are both stochastic processes.
diversification and drift
in the "kill the winner" hypothesis ______ directs the "kill" part of the hypothesis
viruses that infect bacteria
why 16S?
the gene is present in DNA of organisms in all 3 domains and because you can use universal primers to match well to hybridize to the CONSERVED regions to amplify the variable regions
who would have an advantage in the deep ocean where sunlight is limiting/absent?
chemolithoautotrophs that can oxidize ammonia for energy
ribosomes are used in which cellular process?
protein production (translation)
Referring to asexually reproducing microbes, which terms given below are possible synonyms of species?
phylotype, ribotype, operational taxonomic unit (OTU), and Amplicon Sequence Variant ASV/ESV
design of "universal" PCR primers is necessary to do amplification from phylogenetic marker genes among all species present in the environmental DNA sample. Universal PCR primers match well to _____ regions of sequence in the phylogenetic marker gene.
conserved
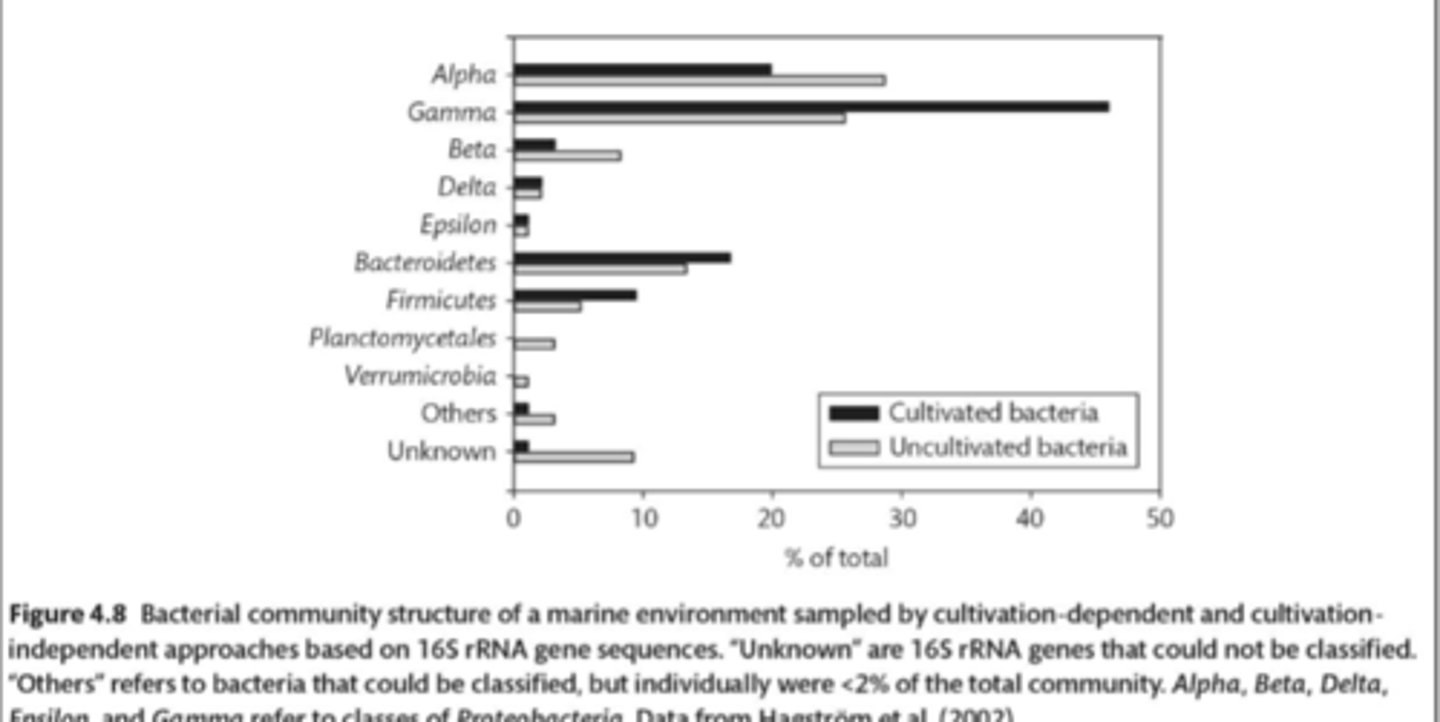
T/F: Relative abundances of marine bacteria as determined by culturing are similar to relative abundances we obtain by sequencing of 16S rRNA genes from environmental DNA.
false
the graph from the book shows bacterial community structure, inferred from data obtained from both culture-dependent and culture-independent methods. based on these data, which type of bacteria listed blow is easiest to culture?
gamma-proteobacteria
the approximate genome size of a member of the SAR11 clade is _____
1.3 Mbp
Proteohodopsin is a protein found in SAR11, a globally dominant member of oceanic bacterioplankton communities. Which of the following best describes the molecular structure of proteohodospin?
a folded chain of amino acids
One proteohodospin gene sequence was discovered by sequencing a segment of genomic DNA from a marine bacterium. The gene sequence was posted in the NCBI database and is 837 bp long. How long is the protein encoded by this gene?
279 amino acids
metagenomics
method you would use to find out about the presence, taxonomic composition, and functional gene sequences from DNA isolated from a mixture of organisms or from a sample within your lower colon
detritus
non-living particulate organic material
rarefraction curves
constructed by calculating the number of OTU's that would be found in a sample with a particular number of individuals
T/F: the number of bacterial taxa in a community is positively correlated with the abundance of bacterial individuals in the ecosystem
true
which clade of bacteria are both ubiquitous and abundant in most oceanic waters?
SAR11 (includes Pelagibacter genus)
bacteria are in less relative abundances in waters below ~100 meters, but some groups of Archaea are more dominant in the deep, because they _____
are chemolithoautotrophs and can oxidize ammonia to obtain energy
The relationship between diversity and organic carbon decomposition is complicated by the presence of many "redundant" taxa with similar metabolic capabilities. However, in some limited instances, for example, an oil spill, the hydrocarbon-degrading bacteria within the rare biosphere community can flourish. This fact is supportive of the _______ hypothesis
seed bank hypothesis
uniquitous
The subclade of the SAR11 clade related to HTCC1062 is found globally in euphotic zones of all oceans
Of the 12 clones they sequenced, how many clones were assigned to the SAR11 clade, based on sequences of the clones?
8 clones, 66.7% of the clones
What would be the # of amplicons with a SAR11-like sequence from a 50,000 read library if you assume SAR11 clade bacteria truly do make up 66.7% of the bacteria in the ocean water?
33,335
If the "average" size of a typicalprotein-encoding gene inprokaryotes is ~1,000 bp, thenhow many protein-encodinggenes would be present in aSAR11 genome of 1,300,000 bp?
1,300 functional genes
list ecological advantages for SAR11 bacteria in maintaining streamlined genomes compared to other marine heterotrophic bacteria
-Selection acts to reduce genome size because of the metabolic burden of replicating DNA with no adaptive value
-Since bacteria in this clade are often dominant in oligotrophic environments, but grow slowly, we can presume a single rRNA operon is sufficient to maintain their moderate but steady rate of growth
-DNA replication requires not only carbon, nitrogen, and phosphorus assimilation, but substantial amounts of energy. A larger chromosome could compromise the careful balance of metabolic efficiency and growth
pangenome
all genomes available of a species
why do obligate intracellular bacteria and Pelagibacter ubique both have streamlined genomes, but for different evolutionary reasons?
Both types get nutrients and organic matter from their environments to supply many metabolites, but obligate intracellular bacteria obtain them directly from their host cells, while P. Ubique and its relatives have adapted to nutrient-poor seawater by eliminating non-essential genes
bacteriophages
simply, viruses that can infect bacteria
why is low % G's and C's better in oligotrophic environment?
Low G + C content reduces the number of nitrogen atoms needed for nucleotide biosynthesis, conserving a limiting nutrient in the ocean
the field of ______ uses data about entire genomes of organisms
genomics
the study of metabolites or the low molecular weight compounds in cells is _________
metabolomics
a _________ is a stretch of DNA encoding a protein
functional gene
when we aim to sequence the genome of a single bacterial strain that we have growing in culture, we are going to do _________
genomics
a _______ study can tell us about the expression patterns of the consortium of bacteria in the ocean without having to culture any of those bacteria
metatranscriptomics
open pangenome
small core genome, large accessory genomes
closed pangenome
large core genome, small accessory genome
what was the example given in the book to describe the concepts of open and closed pangenomes and core genomes?
E. Coli K12 strain
supergenome
sequencing of all genomes of a species
The model has been used to explain genome reduction in Prochlorococcus, a photoautotroph that reaches population sizes in the oceans that are similar to those of Pelagibacter. Briefly, this explanation states that cells in some ecosystems are subject to powerful selection to minimize the material costs of cellular replication. Selection acts to reduce genome size because of the metabolic burden of replicating DNA with no adaptive value
streamlining hypothesis
What is a factor that correlates with prokaryotic genome sizes and also correlates well with maximum growth rates? Hint, think of an important piece of cellular machinery necessary for all protein production (translation)
number of copies of rRNA operon
Codon usage bias can provide clues about possible trophic strategies of the bacteria from which the genomes were derived. What is a factor that correlates with prokaryotic genome sizes and also correlates well with maximum growth rates? Hint, think of an important piece of cellular machinery necessary for all protein production (translation)
Less N is required for DNA synthesis and chromosome replication
metabolomics
study of metabolites or the low molecular weight compound in cells
operon
group of genes with a single promoter (transcribed as a single mRNA). This gene organization structure is very common in prokaryotes, but not present in eukaryotic genomes.
assembly
Bioinformaticians line up overlapping sequences to create in silico contiguous longer pieces of DNA sequence. The DNA fragment sequence reconstructed in the computer, the "contig," is presumed to be the sequence that originated from a single organism, as it was in the original genome