Genetics Final
1/37
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
38 Terms
Define a Mutation and heritability.
An inherited change in DNA sequence. It can be passed to daughter cells (somatic) or offspring (germline)
What is the difference between Forward and Reverse Genetics?
Forward: Starts with a Phenotype —> seeks the mutation.
Reverse: Starts with a Mutation —> observes the resulting phenotype.
Contrast Monogenic vs. Polygenic disorders
Monogenic: Mutation in a single gene.
Polygenic: Mutations across multiple genes.
How does CRISPR/Gene Editing handle "Knock-outs" vs. "Knock-ins"?
Knock-out: Uses Non-homologous End Joining (NHEJ) to break/disable a gene.
Knock-in: Uses Homologous Recombination to insert a specific marker or donor DNA.
Compare Somatic vs. Germline mutations
Somatic: In non-reproductive cells; passed via mitosis to a "clone" of cells; NOT passed to offspring.
Germline: In gamete-producing cells; passed via meiosis to offspring; every cell in offspring carries it.
What is the difference between a Transition and a Transversion?
Transition: Purine to Purine (A→G) or Pyrimidine to Pyrimidine (C→T)
Transversion: Purine to Pyrimidine (A→C or G→T)
Define Frameshift vs. In-frame Indels.
Frameshift: Insertion/deletion not in a multiple of 3; alters the entire reading frame.
In-frame: Insertion/deletion of 3 (or multiples of 3); does not disrupt the reading frame.
What causes expanding nucleotide repeats? What is an example?
The Mechanism: Strand Slippage
How: During replication, the newly synthesized strand "slips" out and forms a hairpin loop (secondary structure).
The Error: DNA Polymerase loses its place and re-copies the repeat sequence.
The Result: The number of repeats increases in the next generation (Anticipation).
The Example: Fragile X Syndrome
Gene: FMR1.
Repeat: CGG.
Threshold: >200 repeats triggers methylation, which silences (shuts down) the gene.
Define Missense, Nonsense, and Silent mutations
Missense: Results in a different amino acid.
Nonsense: Creates a premature STOP codon (shortens protein).
Silent: Changes codon but results in the same amino acid.
What is a Neutral mutation?
A missense mutation that changes the amino acid but does not change the protein's actual function.
Contrast Loss-of-Function vs. Gain-of-Function mutations
Loss: Absence of normal protein function.
Gain: Protein gains a new or abnormal function.
What is the difference between Intragenic and Intergenic suppressors?
Intragenic: Second mutation in the same gene restores function.
Intergenic: Second mutation in a different gene (like a tRNA gene) restores function.
What is the Difference Between a Spontaneous Mutation and a Mutagen?
Spontaneous mutations occur under normal conditions
Mutagens chemically induce mutations
Explain Strand Slippage results
Slippage on New strand → Insertion.
Slippage on Old (Template) strand → Deletion.
Define Depurination and Deamination
Depurination: The "Missing Base"
The Damage: Loss of a Purine (A or G).
The Site: Leaves an AP site (a "pothole" where the base used to be).
The Risk: During replication, DNA polymerase "guesses" what was there, usually leading to mutations.
Deamination: The "Identity Change"
The Damage: Loss of an amino group (NH2).
The Classic Example: Cytosine → Uracil.
The Risk: Changes the code. Since Uracil pairs with Adenine, a C-G pair becomes a T-A pair in the next generation.
What is the primary purpose of the Ames Test?
To identify if a chemical is a mutagen (and therefore a potential carcinogen).
What are the three layers of defense the cell uses to ensure DNA accuracy?
High-fidelity polymerases: Low initial error rate (1 per 100,000).
Proofreading: Immediate 3' → 5' exonuclease activity that fixes 99% of errors.
Post-replication repair: Multiple pathways (MMR, BER, NER, etc.) that fix damage after synthesis is complete.
Why is "redundancy" a key theme in DNA repair?
Many types of DNA damage (like pyrimidine dimers) can be corrected by more than one repair system (e.g., Direct Repair, BER, and NER).
In E. coli, how does the MMR complex distinguish the "old" template from the "new" strand?
The old strand is marked by methylation at GATC sequences. The new strand is temporarily unmethylated, which is where the repair occurs.
What is the step-by-step mechanism of MMR?
The Loop (The "Identify"): The protein complex finds the error and the "Old Strand" tag (methylated GATC), pulling them together into a loop.
The Cut (The "Delete"): Exonucleases move in and strip away the incorrect section on the new strand.
The Re-Type (The "Fill"): DNA polymerase lands on the gap and fills it with the correct nucleotides.
The Seal (The "Save"): DNA ligase seals the final break (the nick) to finish the job.
Compare the genetics and clinical presentation of Lynch Syndrome vs. CMMRD.
Lynch Syndrome (The "1-Hit" Adult Version)
Genetics: Autosomal dominant (inherit 1 bad gene).
Science: MMR "instructions" are broken.
Results: High risk for colorectal and endometrial cancer.
Timing: Usually happens before age 50.
CMMRD (The "2-Hit" Childhood Version)
Genetics: Inherited from both parents (2 bad genes).
Science: Zero working MMR tools from birth.
Results: Early childhood brain, blood, and colon cancers.
Visual: Café-au-lait spots (brown birthmarks).
Timing: ~90% cancer risk by age 18.
What is the primary chemical difference between Direct Repair and Excision Repair?
Direct Repair (The "Fixer")
The Action: Restores nucleotides to their original, correct structure.
The Key: Does not replace any nucleotides.
Analogy: Like popping a dent out of a car door—the original part stays.
Excision Repair (The "Replace-er")
The Action: Removes (excises) the damaged base or a whole chunk of the strand.
The Key: Uses a template strand to fill the gap with new nucleotides.
Analogy: Like cutting out a rusted panel and welding in a brand-new piece.
Identify the two specific direct repair examples
1. Photolyase (The UV Fixer)
Damage Repaired: Pyrimidine dimers (usually T-T) caused by UV light.
The Catch: Found in bacteria, insects, and fish—but NOT mammals.
Logic: Directly breaks the covalent bonds between dimers to restore the original structure.
2. MGMT (The Methyl Remover)
Damage Repaired: O6-Methylguanine (Guanine with an added methyl group).
The Action: O6-Methylguanine-DNA methyltransferase (MGMT) physically removes the methyl group.
The Result: Reverts the base back to a normal Guanine, preventing a G-C → T-A transversion mutation.
What is the defining mechanism of Base-Excision Repair (BER)?
Unlike Mismatch Repair (MMR), which chews up a long stretch of DNA, BER is a highly precise, two-step surgery on a single modified base.
Step 1: The Plucker (DNA Glycosylase): This enzyme finds the specific "weird" base (like Uracil) and cleaves only the base itself.
The Result: It leaves the "skeleton" (backbone) behind, creating a "pothole" called an AP site.
Step 2: The Jackhammer (AP Endonuclease): Because the backbone is still there, this enzyme must nick the phosphodiester bond to remove the empty sugar-phosphate.
The Completion: DNA Polymerase fills the one-base gap, and DNA Ligase seals the nick.
Describe the two detection pathways for Nucleotide-Excision Repair (NER).
1. Global Excision Repair (The "Whole-Genome Scan")
Target: Scans the entire genome for damage.
The Logic: It is independent of transcription. This pathway is constantly surveying "silent" parts of your DNA that aren't currently being read.
Key Proteins: Relies on specific sensors like XPC and DDB to bind to the distortion.
2. Transcription-Coupled Repair (The "Priority Fix")
Target: Only identifies damage in genes that are actively being transcribed into RNA.
The Logic: It initiates specifically when RNA Polymerase stalls because it hits a "bulky" lesion it can't get past.
Key Proteins: Does not require XPC or DDB because the stalled RNA Polymerase itself acts as the "alarm".
What are the clinical signs of Xeroderma pigmentosum (XP)?
Autosomal recessive mutation in NER enzymes. Causes extreme sun sensitivity, severe sunburns, freckling, and high cancer risk. 20-30% have nervous system problems.
Compare Homologous Recombination (HR) and NHEJ.
Homologous Recombination (HR) – The Perfect Fix
Accuracy: Error-free restoration of the original sequence.
Mechanism: Uses a homologous template (sister chromatid) to guide new DNA synthesis.
Key Feature: Involves strand invasion and the formation of Holliday junctions.
Timing: Primarily active in S and G2 phases when a sister chromatid is available.
Non-Homologous End Joining (NHEJ) – The Quick Fix
Accuracy: Error-prone; often causes deletions or insertions.
Mechanism: No template is used; specialized proteins simply rejoin the broken ends.
Key Feature: Recognized by Ku70/Ku80 proteins which recruit processing enzymes.
Timing: Active throughout the cycle, but is the primary repair in G1.
When is each repair system most active?
All Phases: Direct Repair
Logic: Constant 24/7 vigilance.
Activity: Stays "always on" to fix simple chemical changes.
G1 Phase: Daily Maintenance
Logic: Living and growing; no backup copy of DNA is available yet.
Systems: Highest BER and NER activity.
DSB Repair: Uses NHEJ as the primary "quick-fix" for breaks.
S / G2 Phase: Copying Cleanup
Logic: DNA has just been copied, providing a fresh "blueprint".
Systems: Mismatch Repair (following the replication fork).
DSB Repair: Homologous Recombination (uses the new sister chromatid for an error-free fix).
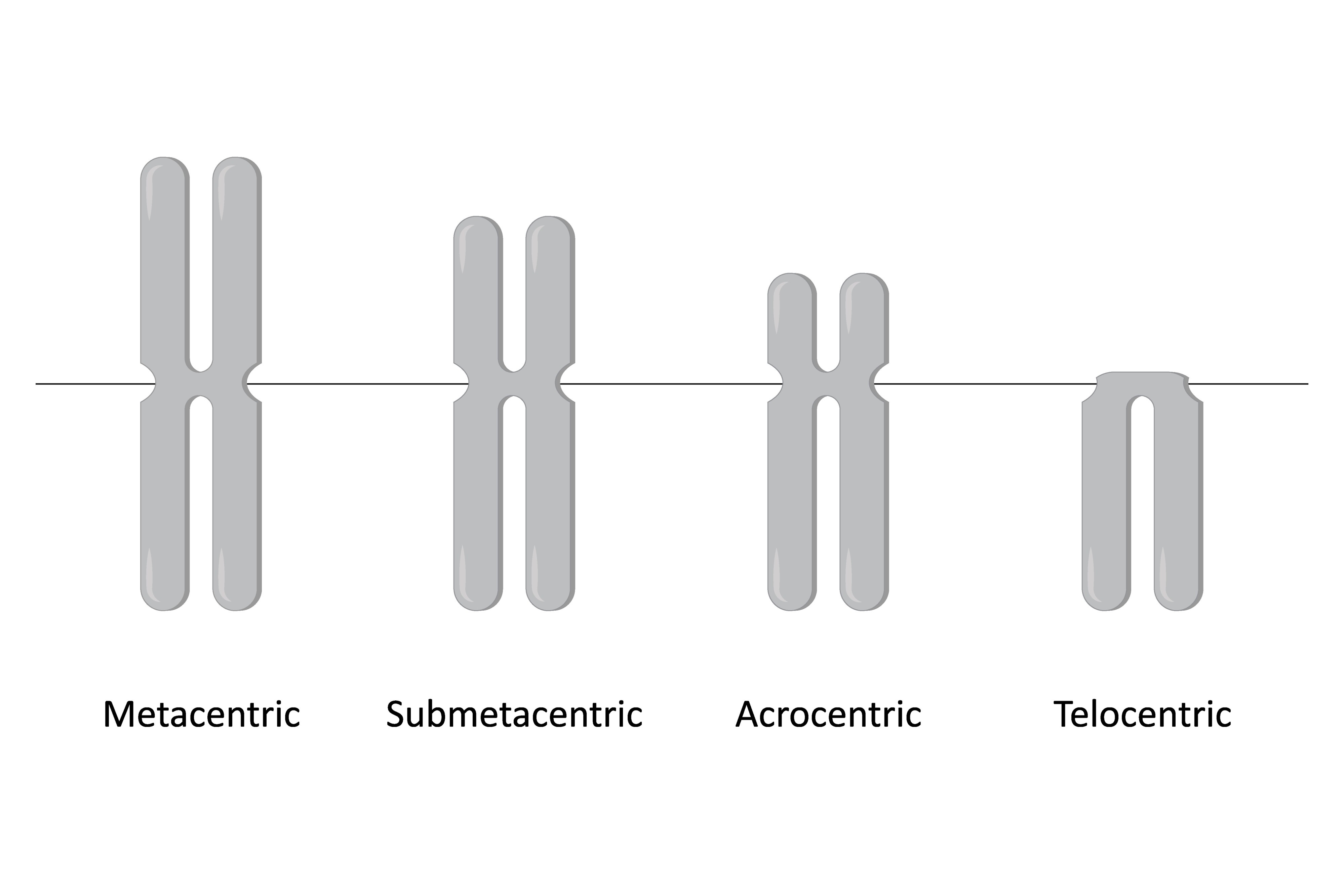
What are the four types of chromosomes based on centromere position, and which one is not found in humans?
Metacentric: Middle.
Submetacentric: Displaced toward one end.
Acrocentric: Near the end.
Telocentric: Absolute end (Not in humans).
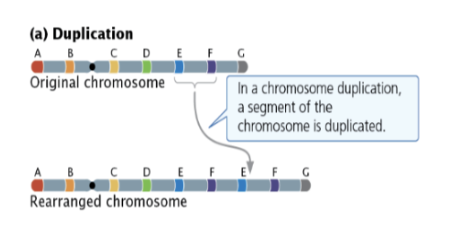
Define Chromosome Duplication and identify its primary cause and effect on phenotype
A segment of the chromosome is doubled.
Cause: Unequal crossing over due to misalignment.
Effect: Unbalanced gene dosage; extra copies disrupt developmental processes that require specific protein ratios.
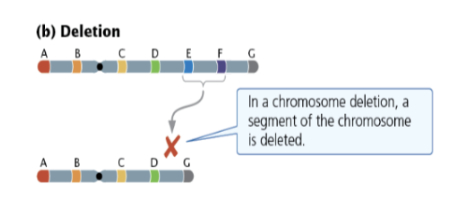
Define Chromosome Deletion and explain the terms Pseudodominance and Haploinsufficiency.
Definition: A segment of the chromosome is lost.
Pseudodominance: Expression of a normally recessive allele because the wild-type homolog was deleted.
Haploinsufficiency: When a single remaining copy of a gene is not enough to produce a normal phenotype.
Lethality: If one centromere is deleted, the entire chromosome is lost during cell division
Examples: DiGeorge Syndrome (22q deletion)
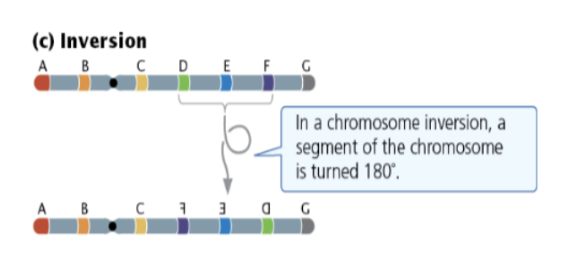
Contrast Paracentric and Pericentric inversions and describe their behavior in meiosis.
Paracentric: Does not involve the centromere.
Pericentric: Involves the centromere.
Meiosis Alignment: Both can only pair with the formation of an inversion loop.
Outcome: Crossing over within the loop results in abnormal gametes, leading to no recombinant progeny.
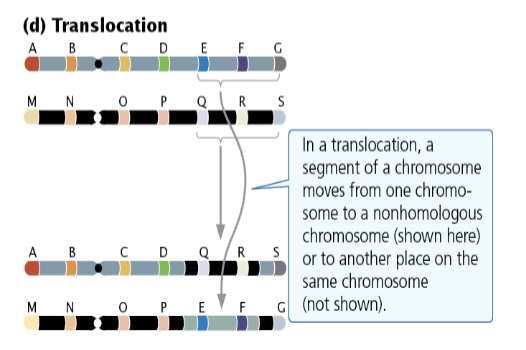
Define the different types of Translocations and identify which causes Familial Down Syndrome.
Reciprocal: Two-way exchange between nonhomologous chromosomes.
Nonreciprocal: One-way movement.
Robertsonian Translocation: Long arms of two acrocentric chromosomes join at a centromere; causes Familial Down Syndrome.
Define Aneuploidy and state the formulas for Nullisomy, Monosomy, Trisomy, and Tetrasomy.
Nullisomy: (2n - 2)
Monosomy: (2n - 1)
Trisomy: (2n + 1)
Tetrasomy: (2n + 2)
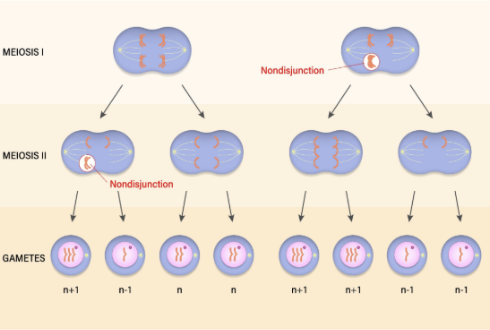
Why are sex-chromosome aneuploidies (e.g., Klinefelter XXY) more common than autosomal ones?
Due to X-inactivation, extra X chromosomes are better tolerated as their gene products are largely silenced.
Distinguish between Autopolyploidy and Allopolyploidy based on the source of chromosome sets.
Autopolyploidy: Sets from the same species (via nondisjunction).
Allopolyploidy: Sets from different species (via hybridization and doubling).
Why is polyploidy significant in plant evolution, and why is it usually lethal in animals?
It increases cell size and vigor in plants (e.g., wheat, potatoes); in mammals, it disrupts developmental gene dosage.
A diploid organism has (2n = 36) chromosomes. How many are in a trisomic member versus an autotetraploid member?
Trisomic: 37 (2n + 1)
Autotetraploid: 72 (4n = 2 * 36)