Hardy-Weinberg Equilibrium & Allele Frequencies
1/29
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
30 Terms
Allele Frequency
Refers to how frequently a particular allele is detected (or observed) in a population
Calculating Allele Frequency: Two Methods
Assumptions:
We are dealing with a single locus in a large and random mating population
We are dealing with diploid species
Can calculate allele frequency by counting or proportions
Calculating Allele Frequency by Counting
Simply add the number of alleles in each genotype
Ex: in the A1A1 genotype, there’s 78 individuals, so the count of the A1 allele in that genotype is 78 + 78 = 156
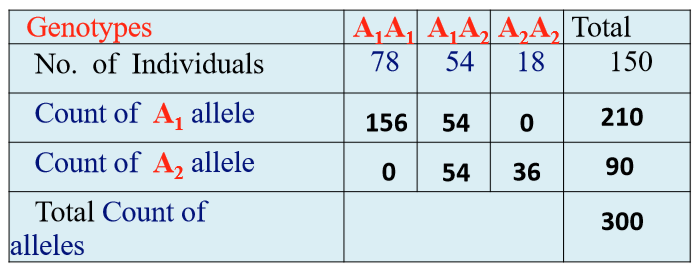
Notations for Allele Frequency
In the context of genotypes and allele frequency: the dominant allele is the most common allele in that population
Most common allele frequency is p
In this case, A1
Minor allele frequency (MAF) is q
In this case, A2
Finding frequency (see image):
f(A1) = 210/300 = 0.7 (this is p)
f(A2) = 90/300 = 0.3 (this is q)
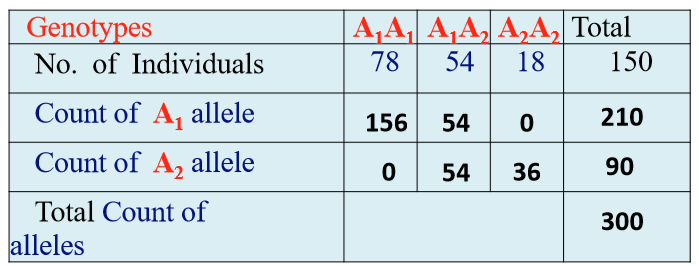
Calculating Allele Frequencies by Proportion
Genotype frequencies: P, H, and Q
P is the frequency of homozygous A1 (A1A1)
Q is the frequency of homozygous A2 (A2A2)
H is the frequency of heterozygous genotype (A1A2)
When calculating, simply divide the number of individuals of that genotype out of the total number of individuals
Ex: P = f(A1A1) = 78/300 = 0.52
To find the frequency of A1 (p) or A2 (q), you use:
p = P + ½ H
q = Q + ½ H
Rules & Assumptions for Calculating Allele Frequencies
p = P + ½ H
q = Q + ½ H
P + H + Q = 1
P = 1 - (H + Q)
Q = 1 - (H + P)
p + q = 1
p = 1 - q
q = 1 - p
Assumptions: large number and random mating (AKA hard-weinberg)
Hardy-Weinberg Equilibrium
Occurs in a large and random mating population in the absence of migration, mutation, and selection
Allele frequencies and genotypic frequencies remain constant
Genotypic frequencies are determined by allele frequencies
Comparing Frequencies Between Generations
If you compare the genotypic and allelic frequencies in g0 and g1 and they are not constant, that means the g0 population is not in HWE
Only one generation of random mating is required for g0 population to reach HWE
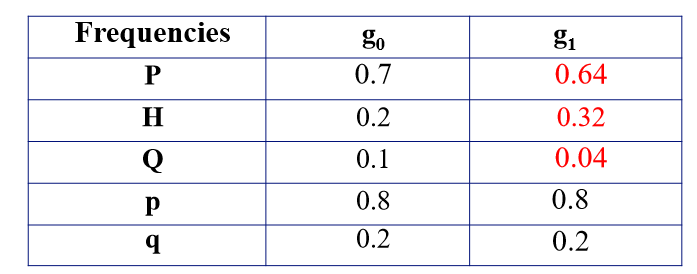
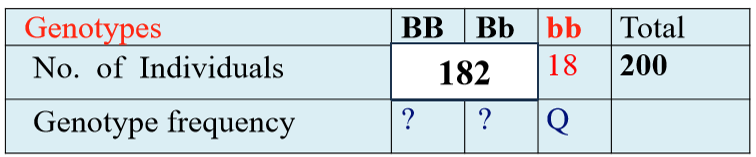
Estimating Allele Frequencies Using Recessive Condition Genotypes
ONLY use f(bb) to estimate f(b) if f(Bb) is not available. Since we’re ignoring the contribution of heterozygotes to the q calculation, we’re assuming the population is in HWE
P = p2
H = 2pq
Q = q2
We know there’s 18 recessive individuals (bb), so we can find Q
Q = f(bb) → 18/200 = 0.09
If Q = q2, then take the square root of Q (AKA q2) to get q: √0.09 = 0.3
Now we can find p: 1 - q = p → 1 - 0.3 = 0.7
Example: Est. Allele Frequencies Using Recessive Condition Genotypes
If you are told 6.0% of the population suffers from a recessive condition, what is the frequency of the dominant allele (rounded to three decimal places)? What is the assumption to calculate the recessive allele frequency?
f(aa) = 0.06
Assume HWE
q = √Q = √q2 → √0.06 = 0.24
p = 1 - 0.24 = 0.76
Genotype Frequency in the Next Generation
f(BB) = p2
f(Bb) = 2pq
This is the carrier
f(bb) = q2
The percentage (frequency) of the individual progeny (the next generation of individuals) showing the genetic condition
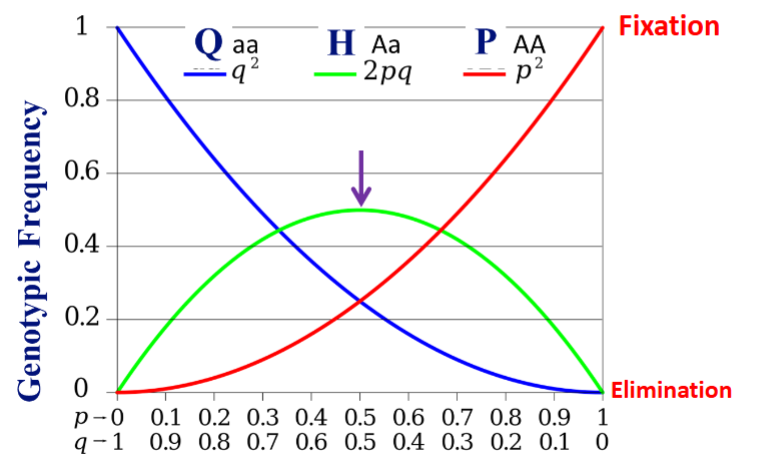
Properties of Equilibrium Populations
Frequency of heterozygote: H = 2pq
The biggest H (Hmax) can be is 0.5
H = 2√PQ or H/√PQ = 2
Provides a quick test for determine HWE without needing to use allele frequencies
If it is equal to 2, then the population is in HWE
Detection of Carriers
Carriers are individuals of any species that carries a single allele of a genetic variant (i.e. heterozygote) that is associated with a specific genetic disorder. Can detect carriers by:
DNA Testing Technology
SNP chip that tests for several genetic conditions or disorders
Relatively cheap and provides lots of services
Test mating
Use of HWE in Detection of Carriers
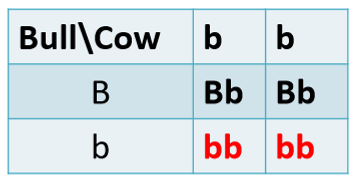
Example: we have a locus controlling coat color with genotypes: BB, Bb, and bb
What’s the probability of being a carrier? (Bb)
P(Bb) = the probability of being carrier or having a red calf
Level of confidence: there is an X% chance that we’d detect the bull is a carrier
Use of HWE in Detection of Carriers: Test Mating
P(detect Bb) = 1 - P(detect Bb after n)
General rule: P(detect Bb after n) = 1 - [1*f(BB) + ¾ f(Bb) + ½ f(bb)]n
When n is the number of progeny
The 1, ¾ and ½ represent the respective probability of not seeing a red calf
Example: Detection of Carriers
P(detect Bb after n) = 1 - [1*f(BB) + ¾ f(Bb) + ½ f(bb)]n
If all 1000 cows are BB, f(BB) = 1
Level of confidence = P(detect Bb) → 1-11000 = 1 - 1 = 0
100% of the cows will be black
If mated to 10 cows, ½ are Bb and ½ are bb
P(detect Bb) = 1 - [ ¾ (½) + ½(½)]10 = 0.9909 or 99%
We’re 99% confident that a cow is not a carrier
We know none of these cows are BB, so we don’t include that factor in the equation
Application of HWE
Genotypes Quality Control (QC) for genomic studies
If genotypes aren’t in HWE, they’re excluded before being used in further analyses as they’re considered genotyping errors
Assumption: loci not in HWE can be lethal or causing disease
Use HWE for deception of lethal alleles is hard because of the need to use a large number of individuals to detect rare variants and standard QC eliminates these genotypes
HWE in Testings
Parentage testing (humane + livestock)
Forensics (wildlife + crime)
In these test panels, the markers (SNPs) should be in HWE and segregate in most of the breeds
These tests are very sensitive to genotyping errors
HWE in the Real World
Most populations in the real world may not be in HWE, but HWE serves as a baseline model for studying the effects of many evolutionary and demographic factors on the genetic structure of the population
I.e. the departure from HWE
Genetic and genomic improvement have been done using selection or crossbreeding, so if we maintain all conditions for HWE, no genetic improvement would be achieved
Conditions of HWE
A random mating population
A large population
No migration
No mutation
No selection
If any of these aren’t met, HWE is violated
Ex: no random mating would be assortative mating and autosomal and sex linkage
Ex: migration would be crossing/importing and exporting semen and embryos
Random Mating
A mating system in which all matings are equally likely
To achieve HWE from a non-equilibrium population, one generation of random mating is required
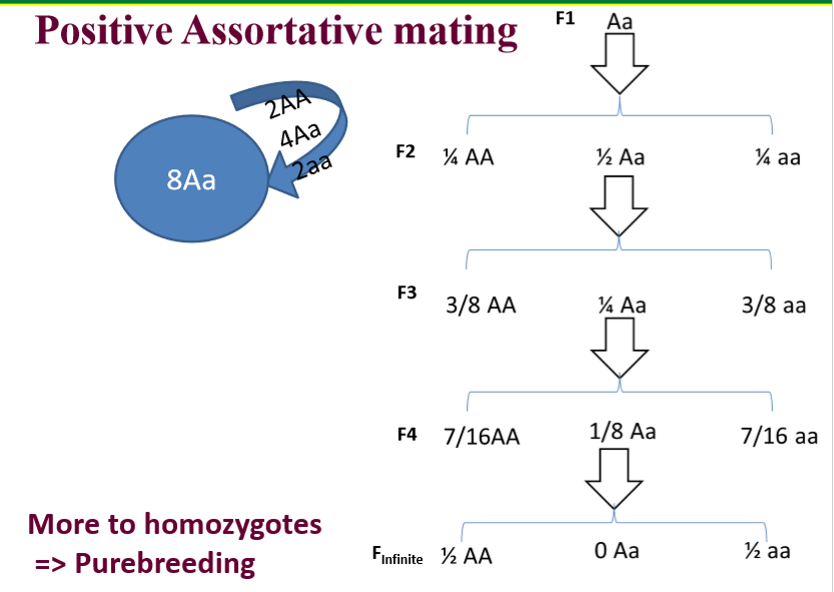
Violating Random Mating: Positive Assortative Mating
Individuals with similar phenotypes or genotypes mating with each other
Mating like to like
Based on pedigree likeness or individual likeness such as performance success, disposition, body shape, etc.
Leads to more homozygotes
Ex: purebreeding
Violating Random Mating: Negative Assortative Mating
Unlike mates to unlike (i.e. different genotypes or phenotypes mating together)
Leads to more heterozygotes
Ex: crossbreeding
(+) or (-): change in genotype frequency but not allele frequency
Recap of Assortative Matings
Assortative matings with no selection lead to change in genotype frequencies not allele frequencies (no removal; we aren’t doing selection)
(+) assortative matings: more to homozygotes like purebreeding
(-) assortative matings: more to heterozygotes like crossbreeding
Factors Affecting HWE: Selection
Goal of selection in a breeding program:
To remove deleterious alleles
To increase frequencies of favorable genes (alleles) in a population
Selection for or against a particular allele changes in both genotypic and allele frequencies
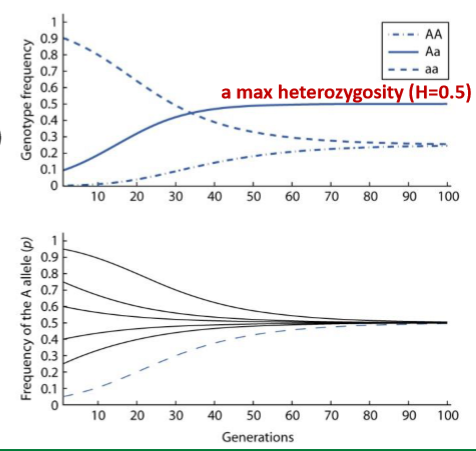
Selection Under Heterozygote Advantage
There’s an overdominance for fitness (i.e. selection favors Aa genotype
For any initial allele frequency, the population converges on a max heterozygosity (H = 0.5)
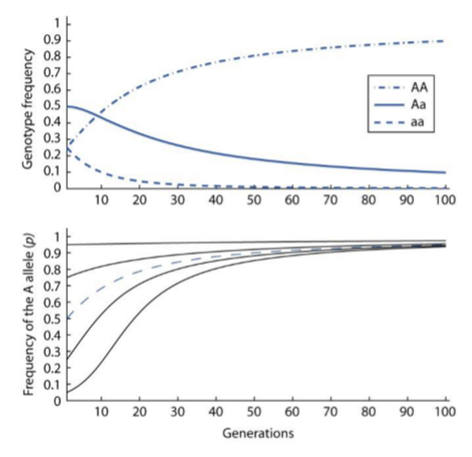
Selection Under Heterozygote Disadvantage
There’s an underdominance for fitness (i.e. selection acts against Aa genotype)
Starting p < 0.5 pops head toward loss
Starting p > 0.5 pops head towards fixation
Populations converge at minimum heterozygosity
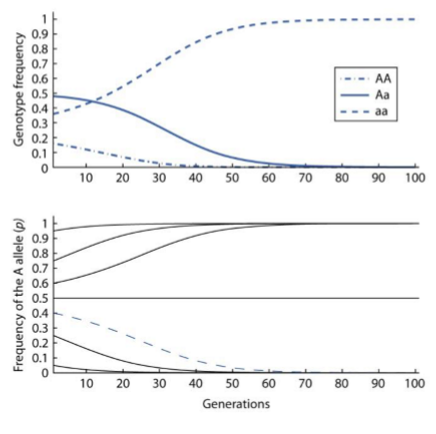
Selection Against a Recessive Homozygote
Selections acts against the aa genotype
Frequency of dominant allele (A) rapidly approaches fixation from any initial allele frequency
Factors Affecting HWE: Migration
Migration is movement of alleles from one population to another
Animals don’t necessarily need to move but if their genetic materials moved using reproductive technology (e.g. frozen embryos, semen)
If allele frequencies are the same between the two populations, then the migration has no effect on allele frequency in the new population
If allele frequencies are different, then migration alter the allele frequency for the new pop.
Factors Affecting HWE: Small Population
Sampling from a small population will lead to random genetic drift
Genetic drift is the random change in allele frequencies due to sampling a finite number of parents