Cell Biology- Final Lab Practical
1/103
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
104 Terms
What general type of centrifuge rotor did you use for the isolation of mitochondria?
Fixed rotor
In the mitochondria isolation lab, name TWO components that you could find in the pellet following the first centrifuge step (low speed for a short amount of time).
Nuclei, cells
Why did you add sand to the homogenization step for the isolation of mitochondria?
to break open the plant cell wall
For the affinity chromatography lab, what protein did you purify using the column?
albumin
What type of biological interaction was used in the affinity column to bind the protein to the column?
antibody-antigen
Clearly explain how the bound protein in the affinity chromatography lab was eluted from the column.
The albumin was eluted from the column using a salt concentration gradient. The increasing salt concentration created an ionic shift in the column and disrupted the antibody-albumin interaction, causing the albumin to elute out.
T or F: For the mitochondrial enzyme assay lab, an increase in enzyme activity corresponded to a low absorbance reading.
True
In the enzyme assay of isolated mitochondria, what enzyme was used to determine the purity of your organelle isolation?
succinate dehydrogenase
Where is succinate dehydrogenase located in the cell/organelle in the lab it was used in?
used in mitochondria isolation lab
found in inner mitochondrial membrane
What chemical was added to the enzyme assay of isolated mitochondria to prevent the final transfer of electrons to oxygen?
sodium azide
What chemical/solution was used as an artificial electron acceptor in the enzyme assay in the mitochondria isolation lab?
DCIP
Clearly explain how you used a color assay and the spectrophotometer to determine enzyme activity in the mitochondria isolation lab.
The DCIP was a blue color but as it was reduced, the color decreased. We used samples with the mitochondrial solution and pellet and added the succinate to evaluate the level of enzyme activity in the samples. We used the spectrophotometer to measure the absorbance at 5 minute intervals. A lower absorbance corresponded to higher enzyme activity
Clearly explain how you used the spectrophotometers to detect the presence of isolated protein from the affinity chromatography experiment.
We eluted out the albumin into fractions using a salt concentration gradient. We then used the spectrophotometer to measure the absorbance of each fraction. We were able to do this because Red660 was added to the samples, which activated the fluorescent marker on albumin. A brighter color = higher absorbance = higher albumin concentration
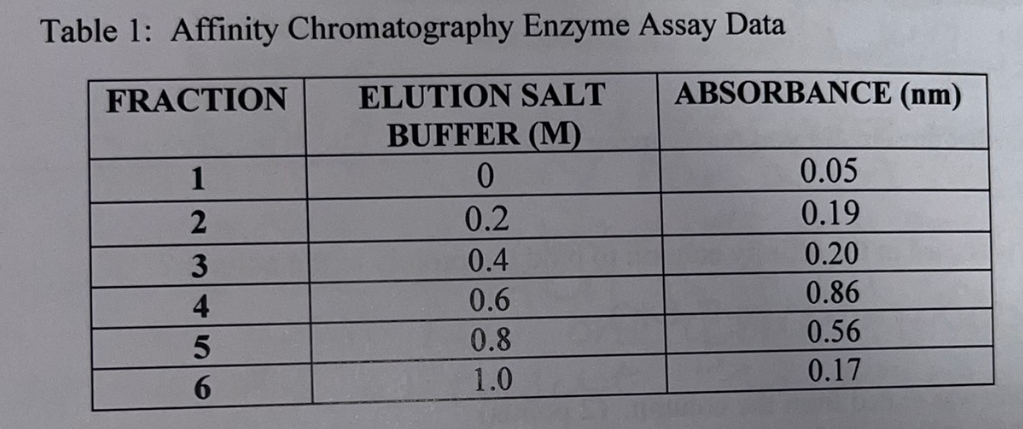
Based on the table, what sample number do you think represents the column flow through? Explain.
1 because salt buffer has not yet been added, which means it was just a flow through containing unbound Albumin
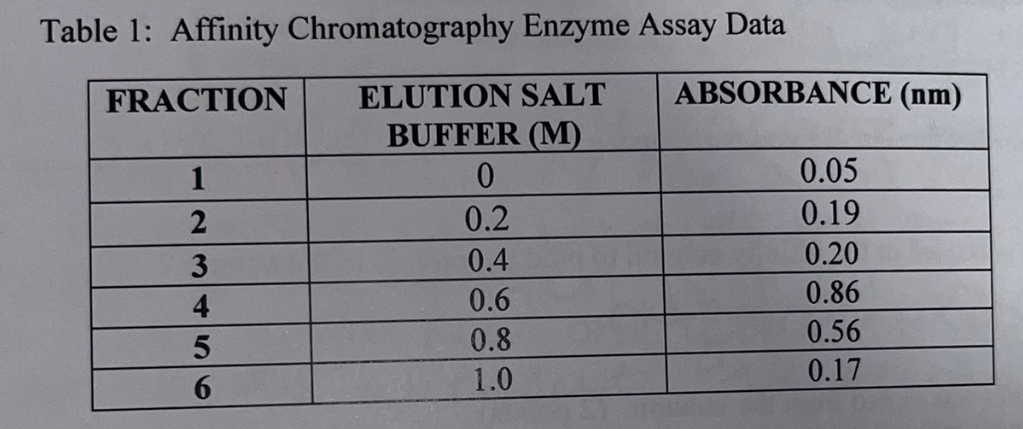
Compare the absorbance values among the fractions and how those values relate to the corresponding salt concentrations. Explain why fraction #4 has the highest absorbance value yet the elution buffer concentration is only 0.6 M.
Fraction #4 has the highest concentration because it is the concentration that disrupted the most antigen-albumin interactions (was the optimal salt concentration). The concentration created the greatest ionic shift. Following elution #4, the majority of albumin has been eluted leading to decreasing absorbances in fractions 5 and 6.
You can test the purity of your mitochondria sample by staining the isolated mitochondria with Janus green. Do you think this staining can be used to predict the outcome of your mitochondrial enzyme assay (i.e. if the assay will work or not)? Explain.
No because janus green only stains mitochondria, it doesn’t tell us if the mitochondria are functional.
What plant species did you use when setting up the experiments in the cytoskeleton lab?
Elodea
Based on the following data, what row(s) would you expect to show a positive ELISA result? Explain.
You have an ELISA plate with four rows and three columns
You have added antigen into all the wells in rows 1, 2, 3, and 4, primary antibody into rows 2, 3, and 4, and secondary antibody to rows 1, 2, 3, and 4.
You then added a substrate specific for the enzyme marker linked to the secondary antibody. The substrate is added to all the wells.
Rows 2, 3 and 4 because they were the only rows to have all the components required for a positive result (antigen, primary and secondary antibodies). The only way for the antibody to bind to the antigen is if both antibodies are used.
If one of the patient samples gave a negative ELISA result for smallpox virus, does this mean that the patient does not have the disease? Give one reason for a negative result when an individual has the disease.
they may not have developed the antibodies yet
In the food borne illness lab, you determined the presence of Shigella sonnei in food by DNA gel electrophoresis. According to the case study, of the six people who were sick only 5 ate the contaminated food. How is this possible?
the person who didn’t eat it could have contracted the bacteria from someone who did since it can spread person to person
Name one cell biology technique (that we did this semester in lab) that you could use to confirm the presence of Shigella sonnei in a food sample.
ELISA
Why would the addition of either 0.5% Janus green or 0.5% neutral red have no effect on cytoplasmic streaming in a plant cell?
Because they are vital stains which are meant to stain living cells
In the cytoskeleton lab, what drug was used to inhibit the function of microtubules?
Taxol
In the cytoskeleton lab, what was the purpose of testing for cytoplasmic streaming in the presence of DMSO?
Both Taxol and cytochalasin D were mixed with DMSO so we had to ensure that DMSO did not affect streaming because if it did we would be changing 2 variables and wouldn’t be able to gauge the true effects of the disruptors.
In the cell surface receptor lab exercise, what receptor was localized in epithelial cells?
Con A
What sugar forms the side chain on the con A receptor?
Mannose
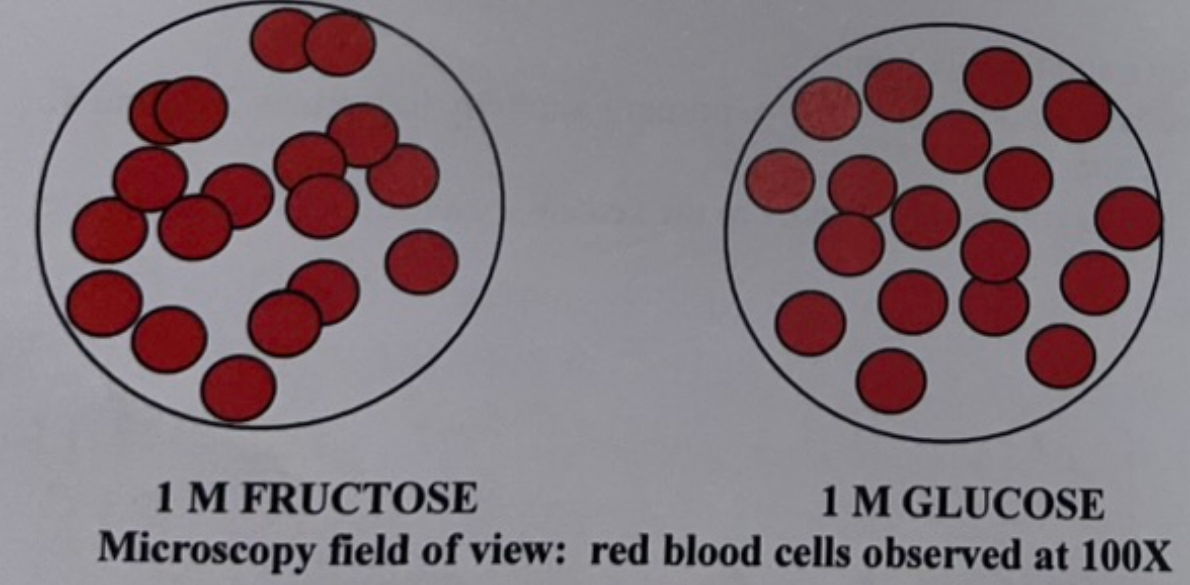
Based on the data shown in the image, what sugar would form the side chain on the receptor involved in red blood cell agglutination?
Glucose because it inhibited the agglutination of the red blood cells. The lectin bound to the excess glucose because it has a high affinity for it, meaning there was not enough lectin to bind to the receptors. Fructose cannot be the side chain because there is agglutination, meaning the lectin preferred the receptor over the excess sugar.
Based on your data from the cytoskeleton lab, what filament(s) are required for cytoplasmic streaming? Explain.
Actin because when we added cytochalasin D, which disrupts actin, we observed no streaming. However, when we added taxol which prevents microtubule disassembly, it did not affect streaming.
What cytoskeletal motor protein might be required for cytoplasmic streaming in the cells we observed in lab?
Myosin because the cells are actin-associated
Would you be able to use a standard ELISA to detect the presence of nucleic acids in a sample? Explain.
No because nucleic acids all have the same structure, unlike antibodies.
For the acridine orange staining protocol that you used in lab. what solution was used to permeabilize the cells?
acetic alcohol
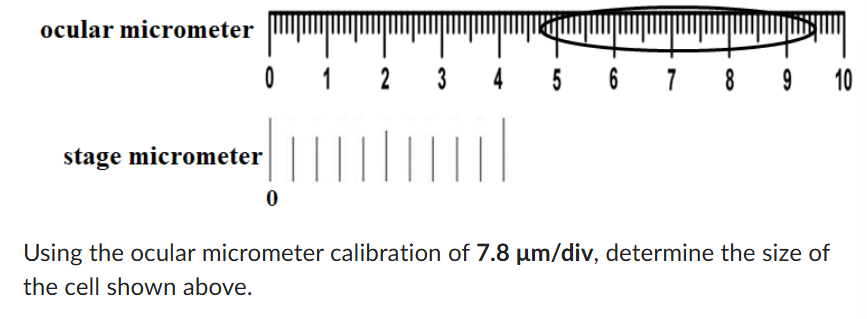
374.4 um
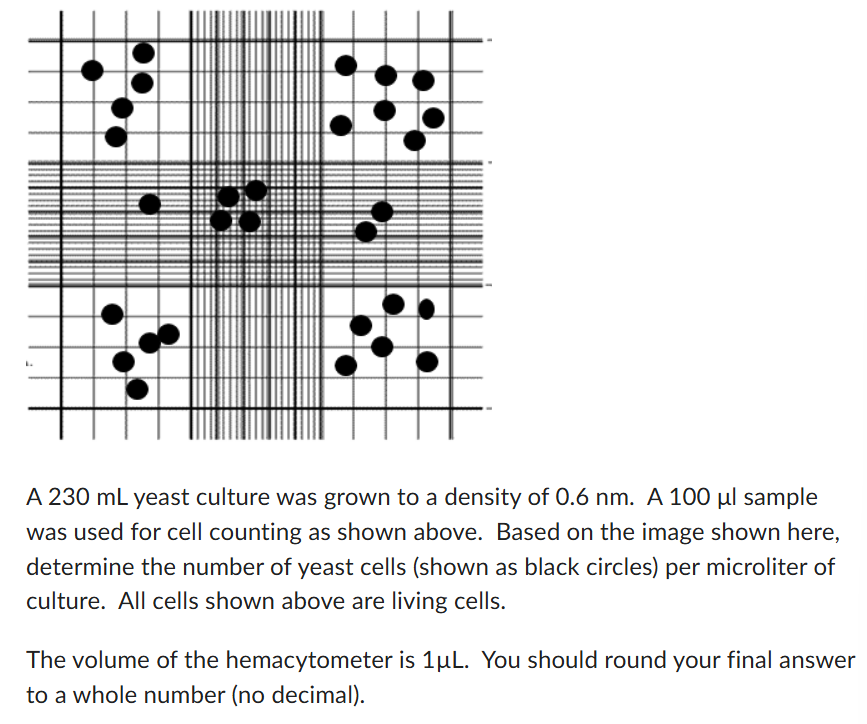
1 cell/uL
For the fractionation step of an isolation procedure to isolate cell nuclei, would you use a high speed or low speed centrifuge spin?
low
What type of microscope would you use to visualize the nucleus in a living epithelial cheek cell?
phase contrast
What type of microscope would you use to visualize double stranded DNA in a epithelial cheek cell?
fluorescence
In the acridine orange staining protocol, what solution(s) can act as a cell fixative?
ethanol and acetic alcohol
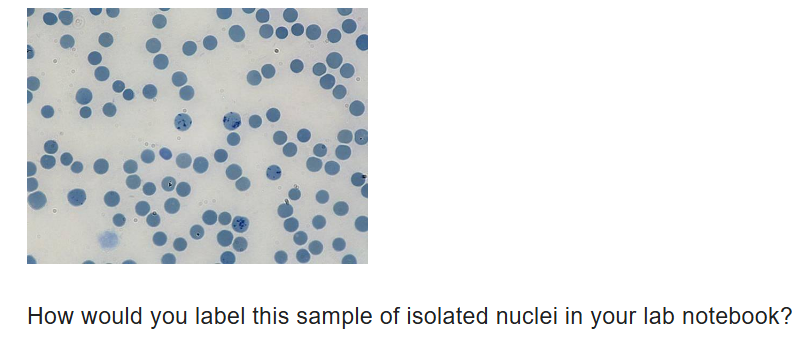
stained positive control slide
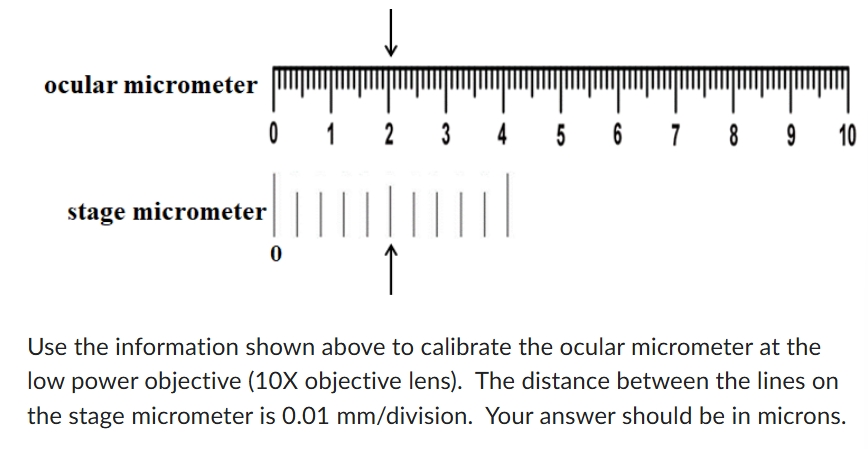
2.5 um
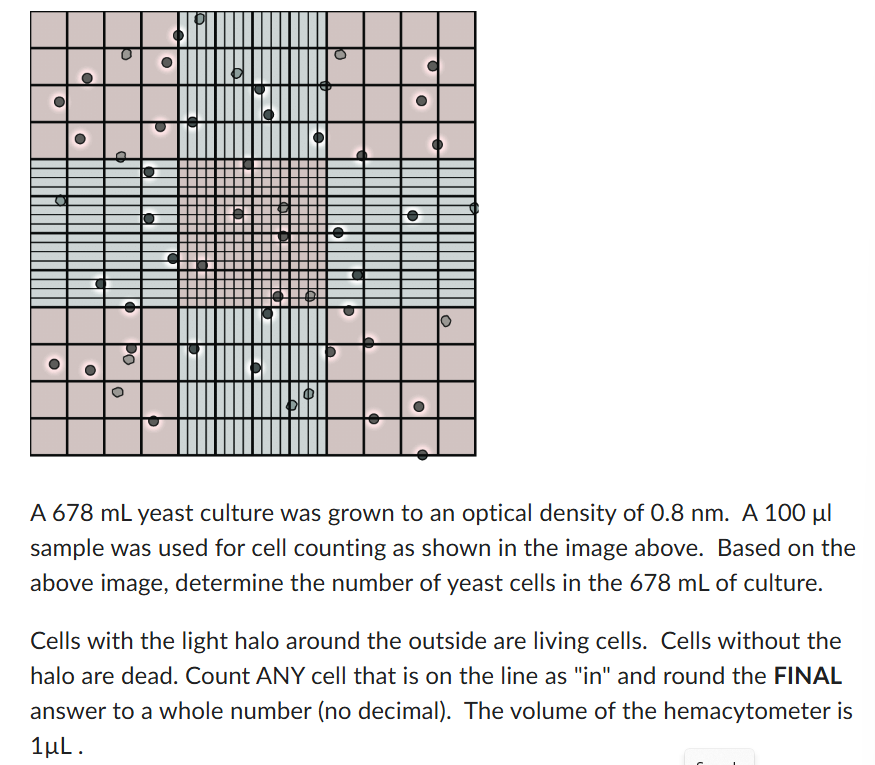

890,000
What non-fluorescence dye or stain can be used to determine cell viability?
trypan blue
Would the number of intact nuclei visible on a slide increase or decrease after treating the nuclei sample with 0.5 M NaCl and staining with methylene blue?
decrease
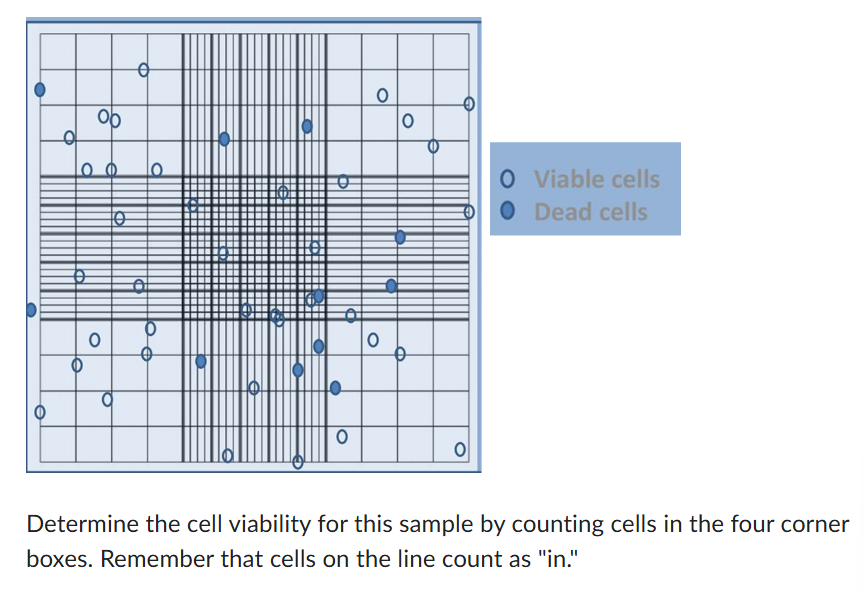
91.3%
What type of microscope would you use to visualize a nucleus in an epithelial cheek cell without the addition of a stain?
phase contrast
What does ELISA stand for?
enzyme linked immunosorbent assay
What is the purpose of an ELISA? What kind of samples is this performed on?
uses antibodies to detect proteins (antigens; antibodies; hormones)
What are the overall steps of an ELISA?
binding of the antigen to the wells of the microtiter plate
binding of the antibody to the antigen
detection of the antigen
When performing an ELISA, how does the antigen bind to the wells of the microtiter plate?
plates are made of polystyrene which binds proteins by hydrophobic interactions
When performing an ELISA, how does the antibody bind to the antigen?
generally use 2 antibodies: primary and secondary (indirect)
primary antibody is specific to the antigen
secondary antibody is linked to HRP (horseradish peroxidase)
When performing an ELISA, how is the antigen detected?
adds a colorless substrate that is broken down into colored products by HRP
At the conclusion of the ELISA lab, how did you know which samples contained smallpox?
the light blue wells meant smallpox were present
What are two medical applications of an ELISA?
pregnancy tests
sandwich ELISA
antibody used to detect human chorionic gonadotropin (HCG), the commonly measured protein (in blood or urine) that indicates pregnancy
disease detection in humans, animals, and plants
used to detect disease causing organisms (ex: viral proteins of smallpox virus)
used to detect antibodies produced in response to infection (HIV)
What is the advantage to performing an ELISA?
highly sensitive and specific
What are the disadvantages of performing an ELISA?
occurrence of false (+) and false (-) result
in many cases it is followed up by a confirmatory test
false (+) may occur if you have an underlying condition (autoimmune diseases) such as lupus or rheumatoid disease
false (-) result doesn’t always mean there is no infection
titer
some antibodies are not produced immediately following infection
What is cytoplasmic streaming? What is the function?
flowing of the cytoplasm in eukaryotic cells
occurs in both plant and animal cells
actin or microtubule-based process
function:
creates cytoplasmic reorganization during cell production, aids in the delivery of nutrients and metabolites to all parts of the cell
Describe cytoplasmic streaming in Elodea
organelles (such as chloroplasts) move along the inner cell wall
rate of motion is affected by light exposure, temperature, and pH levels
chloroplasts are moved in the cytoplasm stream possibly to a position of optimum light absorption for photosynthesis
motor proteins propel the movement of chloroplasts
What vital stains did we apply to the elodea leaf? What result would you expect for these stains to have on cytoplasmic streaming?
vital stains (0.05%):
janus green: stains mitochondria
neutral red: reacts with charged molecules in cytoplasm
neither stain should affect streaming since vital stains are made to stain living cells
What temperature treatment did we apply on the Elodea leaf? What result would you expect on cytoplasmic streaming?
temperature (4 C)
placed on ice because fridge light goes out when door is closed so 2 variables would change
you would slower cytoplasmic streaming when coming off the cold but speeds up as the leaf warms up
How did we test the effect of light on cytoplasmic streaming? What result would you expect?
we covered the slide with foil for 30 minutes
when we took the foil off, we saw faster streaming because dark → light
How did we test the effect of pH on cytoplasmic streaming? What result would you expect?
water buffered to pH 8.3
still should see streaming
water buffered to pH 5.8
could affect the structure of the motor protein
How did we test the effect of anesthetics on cytoplasmic streaming? What result would we expect?
used 2% ethanol
still saw streaming because the concentration of ethanol that we used was low
higher concentration would have stopped streaming
Which cytoskeletal disrupters did we use in the cytoplasmic streaming experiment? Why? How were they prepared?
wanted to determine which cytoskeletal filament is needed for streaming of chloroplasts in Elodea
cytoskeletal disruptors:
cytochalasin D: disrupts actin filaments
taxol: prevents microtubule disassembly
both disruptors are insoluble in water
both solutions were made in DMSO (dimethyl sulfoxide)
first test streaming in presence of DMSO alone- why is this control necessary?
to make sure streaming can still happen in DMSO by itself
Which cytoskeletal filament is needed for streaming of chloroplasts in Elodea? How do you know based off the experiment?
actin filaments
when we applied cytochalasin D, streaming stopped
when we applied taxol, streaming was not interrupted
What are lectins?
glycoproteins that cause cell agglutination
glycoprotein: proteins with a covalently bound oligosaccharide (sugar) side chain
side chains can be composed of glucose, mannose, galactose, fructose, galactosamine, or sialic acid
lectin (ligand) will bind lectin receptors on the surfaces of neighboring cells (agglutination)
How does lectin bind to a lectin receptor?
by interacting with the sugar side chain on the receptor (different lectins selectively bind to different sugar side chains
Describe the theory behind localizing receptors for Con A and determining the sugar.
used epithelial cheek cells
Con A couples to horseradish peroxidase (HRP) which reacts with a peroxidase substrate to generate colored products
receptor is on surface of cheek cell so that’s where there should be color change
How does sugar competition work with Con A binding to receptors?
add excess sugar to the samples
the sugar that inhibits binding of Con A to the receptor (no color on the side) is the sugar that forms the side chain
if Con A can recognize and bind to the excess free sugar, there is not enough Con A to then bind to the receptor on the surface of the cell
What sugar forms the side chain on the Con A receptor?
the sugar that inhibits agglutination is the sugar that forms the side chain on the Con A receptor
What tool did we use in the foodborne outbreak lab?
mini-one DNA electrophoresis system
What is PCR? What are the steps?
PCR = polymerase chain reaction
use extracted DNA to produce DNA products of known size
process:
1. denaturing
2. annealing
3. extension
Describe agarose gel electrophoresis.
used to separate DNA fragments
DNA fragments migrate through the agarose gel towards the (+) electrode (DNA has a negative charge)
DNA separation is based on size (larger fragments don’t migrate as far in the gel as smaller fragments)
once the agarose gel is prepared, the gel is placed in the electrophoresis chamber and covered with buffer (TBE)
TBE contains tris, borate, and EDTA
DNA gel loading dye contains 2 different dyed (bromophenol blue and xylene cyanol) for visual tracking of DNA
the presence of glycerol ensures that the DNA in the sample forms a layer at the bottom of the well
the agarose contains gel red which allows us to visualize the DNA after running
What is the biological interaction used for ELISA?
antibody antigen
Why are chloroplasts moved around by cytoplasmic streaming?
to get to a position of optimum light absorption for photosynthesis
What does it mean if cells agglutinate?
stick together to form clumps
What are the receptors for lectins?
cell-surface glycoproteins
Pea lectin has a high affinity for…
mannose and will interact selectively with mannose chains on membrane glycoproteins. The addition for excess mannose to a cell culture will inhibit the binding of the pea lectin to the cell surface
How do red blood cells stick together?
a single lectin molecule contains two or more binding sites for receptor interaction. as a result, lectins will form bridges between receptor-containing cells which causes the cells to stick together to form clumps. In part B, you will perform a Con A induced agglutination reaction with sheep red blood cells to identify the sugar residues of the con A receptor that are involved in Con A binding
How were we able to tell how much Con A bound to the receptor?
the intensity of the purple color
In the Con A lab, why was the mouth rinsed several times with tap water?
bacteria and mucus will interfere with the binding assay
How were the epithelial cheek cells fixed in the Con A lab? Why did we fix them?
first by heat
second by ethyl alcohol
the fixation makes the cells permeable to macromolecules and causes them to adhere to the surface of the slide so they won’t be washed away in subsequent steps
Why did we was the slides with fresh con A buffer after adding the solutions to the slides?
this stem removed Con A-peroxidase that was not bound to the epithelial cells
How do you calculate percent agglutination?
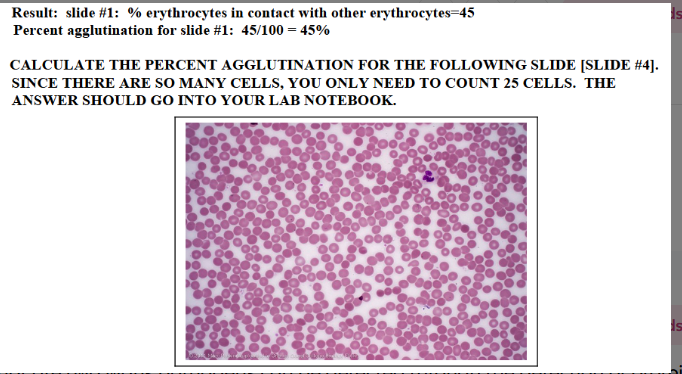
What are two ways pathogens can be detected through the detection of proteins?
western blot and ELISA
What was the pathogen we were trying to detect in the foodborne illness lab?
Shigella sonnei by the presence of DNA
What are the basic steps of organelle isolation?
homogenization: disruption of cells to create a cell homogenate
fractionation: separation of cellular molecules/organelles using centrifugation
Describe homogenization in the mitochondria isolation lab.
creates a homogenate
uses buffers containing detergents (NP-40 or Triton X) and other components (proteases and chelating agent)
uses mechanical action to break apart cells in tissue layer and to break open cells (mortar and pestle)
addition of sand
During the homogenization sand of the mitochondria lab, we added sand- why?
because we were using plant tissue and the sand was abrasive and broke open the plant cell wall
Describe fractionation in the mitochondria isolation lab.
uses centrifugation
differential velocity centrifugation
this process separates organelles based upon their size and density
vary time and speed of the centrifugation to isolate organelles
usually use a fixed angle rotor
when one centrifugation step is done, you remove the supernatant and centrifuge again
Describe differential velocity centrifugation in the mitochondria isolation lab.
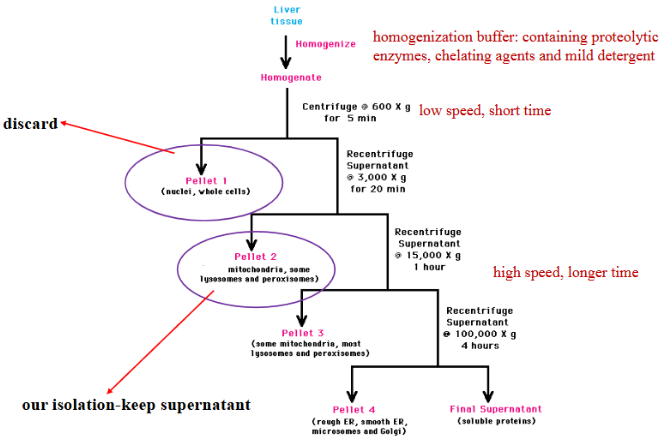
What are the methods of determining the purity of a sample?
staining
janus green: vital stain, stains mitochondria
downside: doesn’t tell us if the organelle is functional
enzyme assay
determines purity of sample
determines if mitochondria (or other organelle being studied) is functional (cell free system)
What is a cell free system?
A cell-free system (CFS) is defined as an in vitro technology that allows for the synthesis of proteins and the study of cellular processes outside of living cell
What role does succinate dehydrogenase play in mitochondria? What is the substrate?
bound to inner mitochondrial membrane (with its coenzyme FAD [flavin adenine dinucleotide])
catalyzes the oxidation of succinate to fumarate (generates 2 protons and 2 electrons)
FADH2 carries the protons and electrons to the electron transport chain
accepted by ubiquinone
subsequently passed down the chain to oxygen
substrate of succinate dehydrogenase: succinate
Overall, how was the enzyme assay performed in the mitochondria isolation lab?
studied the activity of succinate dehydrogenase by replacing ubiquinone with an artificial electron acceptor: DCIP (blue dye)
oxidized form of DCIP will be reduced by accepting electrons from FADH2
changes the dye from blue to colorless; color change is measured by spectrophotometer and a decrease in absorbance
THEREFORE, a lower A600 means there is higher enzyme activity
sodium azide (poison!!) is added to the reaction to block the final transfer of electrons to oxygen
ensures that the electrons are passed to the dye and not funneled into the electron transport chain
What should the graphs have looked like after the mitochondria enzyme assay?
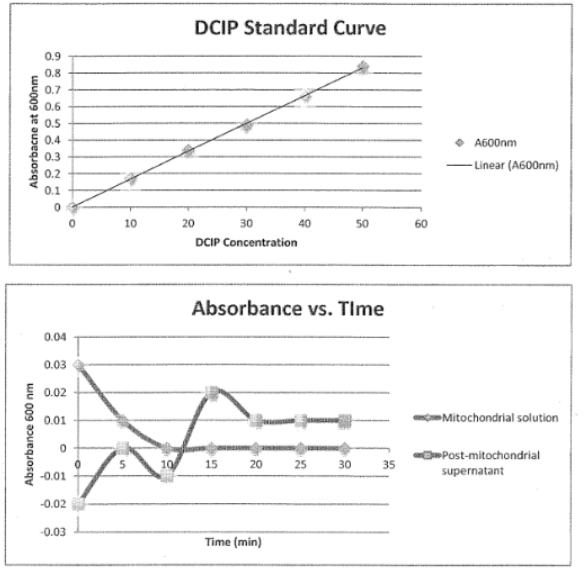
How did we purify albumin in lab?
affinity chromatography
What is albumin?
a protein made by the liver and found in vertebrate blood
maintains fluid balance in the blood and tissues
How does affinity chromatography work (specifically in the albumin purification lab)?
biological interactions used: antibody-antigen (target protein: albumin)
loading the column/binding of albumin:
the column contained beads bound to an antibody for albumin
albumin can bind to the antibody so the albumin will bind to the antibody bound to the beads in the column
washing the column
column is washed with an affinity binding buffer to remove proteins that did not bind
elution of albumin
bound albumin will be eluted from the column by adding varying salt concentrations
salt solutions can disrupt the antibody-antigen interactions by creating an ionic strength shift between the bound molecules
the shift in ionic strength will elute the protein from the column