Homology Modeling
1/9
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
10 Terms
Homology Modeling
a computational technique to predict a protein’s secondary and tertiary structure based on sequence similarity to a known structure (template)
Homologous Proteins
homology: common evolutionary ancestry
we infer homology when two sequences share more similarity than expected by chance (excess similarity)
however homologous protein do not always share significant sequence similarity
homology is used for structural modeling
Key Assumption
proteins w/ similar sequences have similar structures
from structural studies, protein structures is more highly conserved throughout evolution than the protein’s sequence
in some protein families, fewer than 5% identical residues are present, while 50% of the related protein’s structure is highly conserved
below a certain threshold, may or may not be homologous
need to have at least 30% sequence identity to find a homolog
if we can’t, forget about homolog
Homology Modeling Pipeline
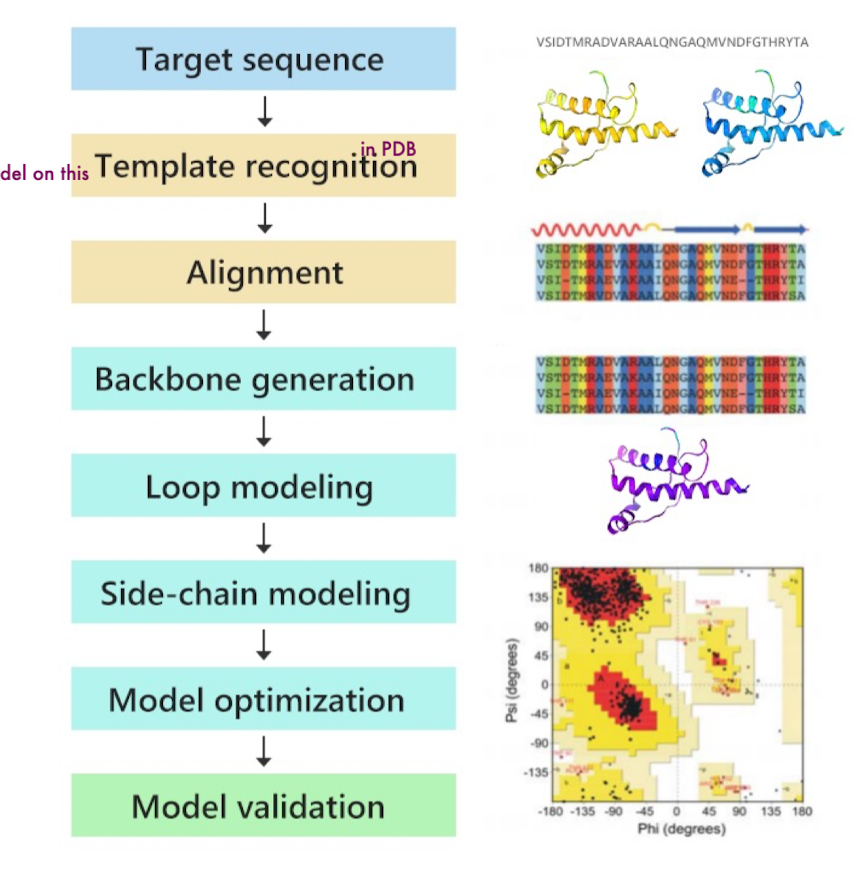
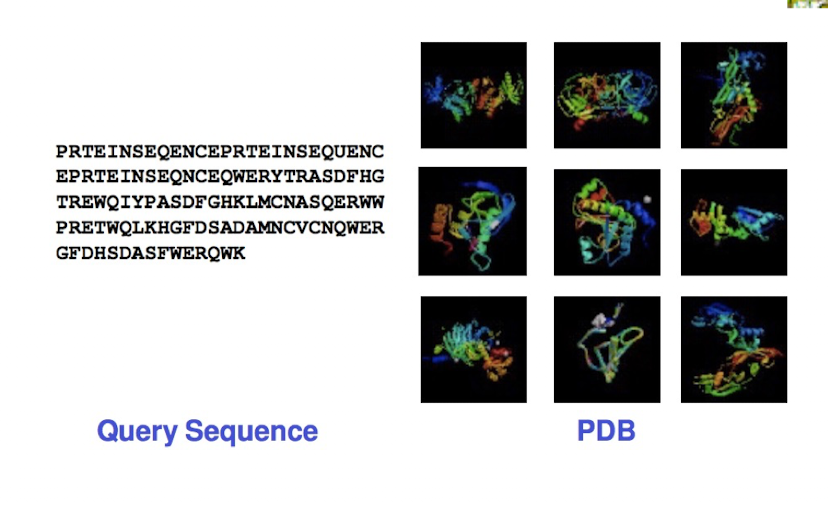
Homology Modeling Pipeline: Template Identifcation
tools: BLAST, PSI-BLAST
databases: protein data bank (PDB)
selection criteria:
sequence identity >30% for good results
structural resolution of template
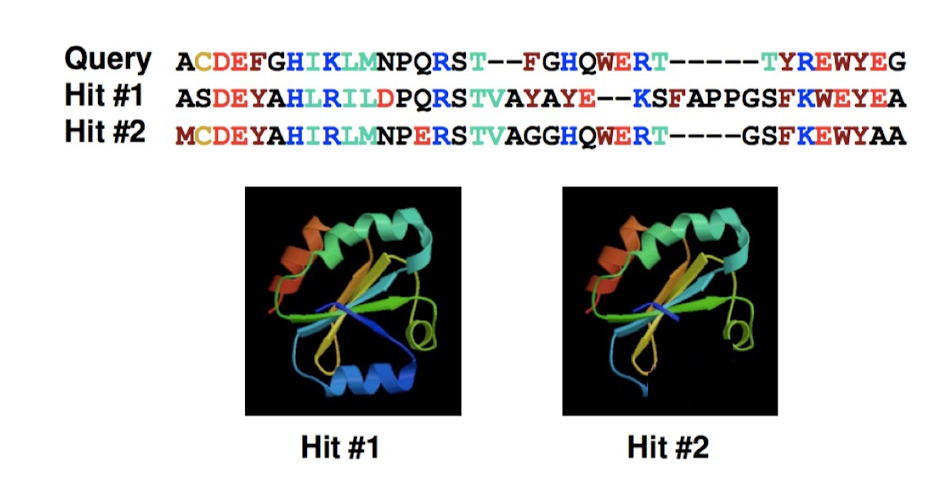
Homology Modeling Pipeline: Alignment
tools: Clustal Omega, MUSCLE, MAFFT
challenges:
correctly aligning gaps and insertions
handling low sequence similiarity
Importance: errors in alignment propagate to the final structure
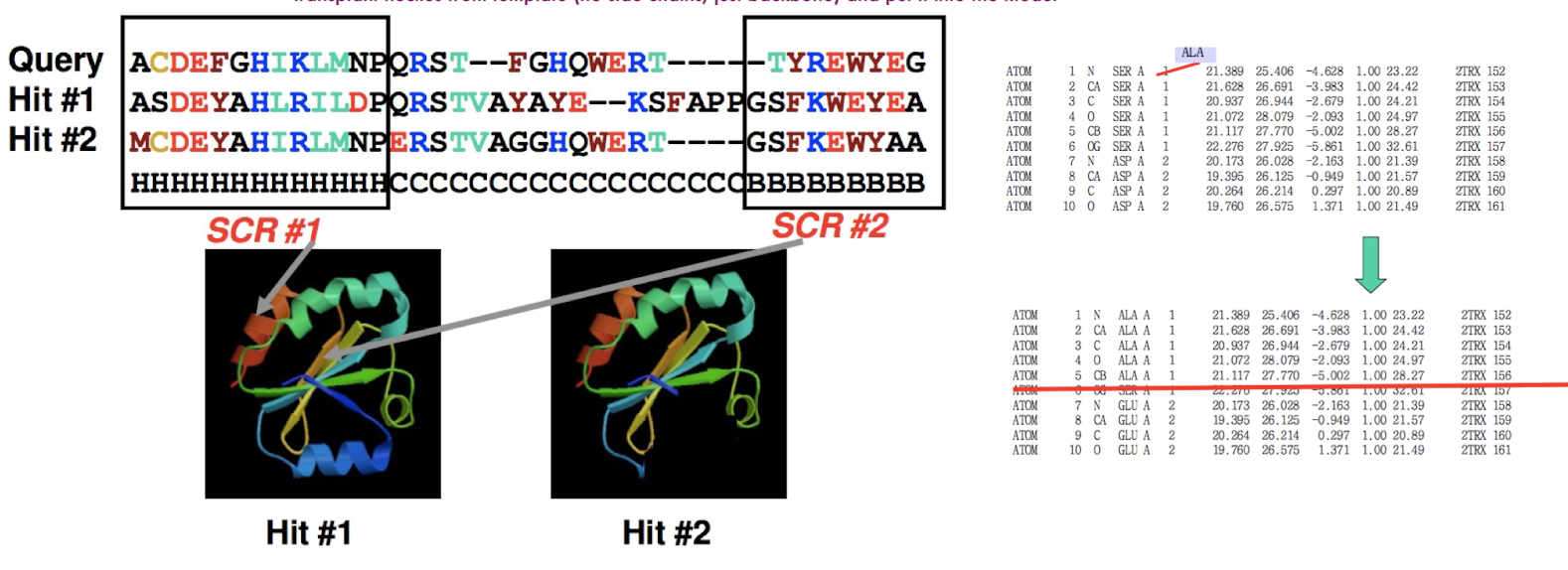
Homology Modeling Pipeline: Model Building
rigid body modeling: copying template backbone
transplant helices from template (no side chains, just backbone) and put it into the model
specifically, identifying sequence conserved regions and copying the coordinates of the backbone atoms
Homology Modeling Pipeline: Loop modeling
loop modeling: predicting missing regions
identifying sequence variable regions and testing different loop templates/models (database search)
loop regions may differ b/w homologs
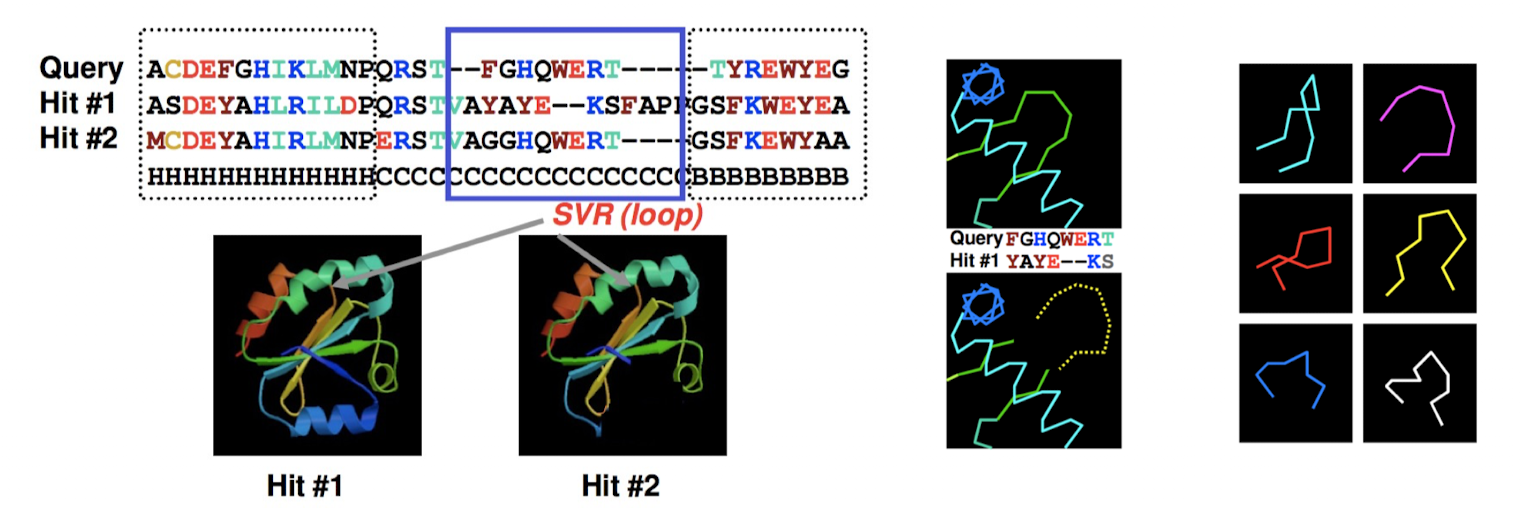
Homology Modeling Pipeline: Side-Chain Modeling + Model Optimization
side chain placement, trying rotamers (Rosetta)
modeling in the side chains; in conserved sequences just take from the template
side chains can only adopt 3 rotamers and how they fit into the structure to build the model
the first model might have lots of clashes, so then we put it into a force field to minimize clashes (force field automatically separates atoms)
goal is to optimize structure
energy minimization: corrects geometry (e.g bond angles, clashes0
MD simulations: improves local flexibility and structural accuracy
Iterative refinement: combines multiple round of modeling and energy minimization
Validation of Predicted Models
Ramachandran Plot: evaluates backbone dihedral angles
want to know if phi, psi bonds are ok
correct bond lengths and angles
model might accidentally push or pull bond angles and interactions, so make sure that the model is consistent w/ chemistry
energies of models (Force field, DOPE Score used by MODELLER)
burying of hydrophobic residues and exposure of polar residues
combining these features: QMEAN in the SWISS-MODEL server