H3 Toth- Cytoskeleton II - actin network
1/8
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
9 Terms
describe the structure and assembly of actin filaments
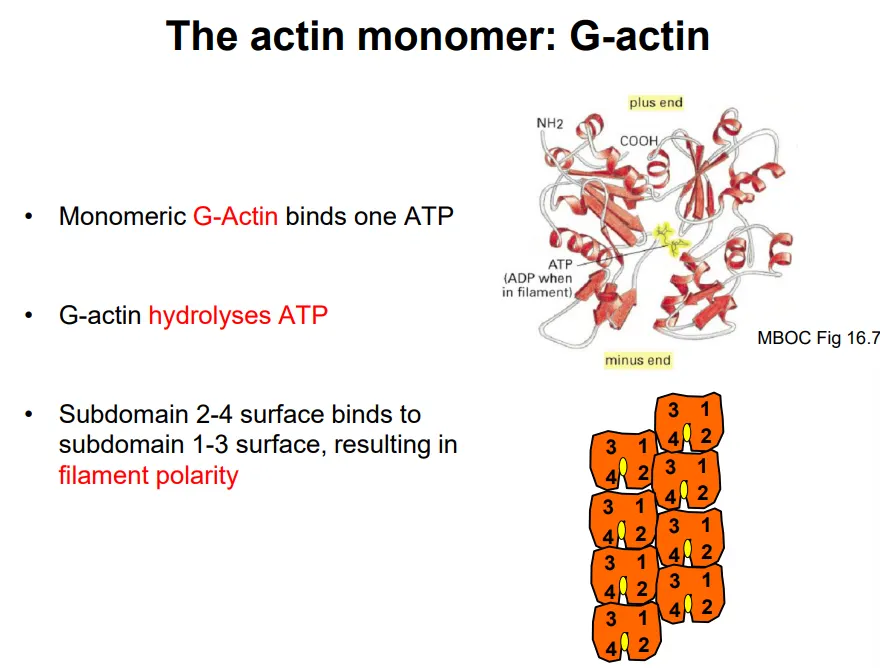
F-actin (filamentous actin) is a linear polymer of a single globular monomer, G-actin
it is a left-handed helix with 13 monomers per turn (37nm)
each G-actin binds and hydrolyses one ATP to spontaneously self-assemble (once three monomers have bound together)
one surface (2-4 subdomain) binds preferentially to the opposite surface (1-3 subdomain), so the filament is polar
polymerisation happens at the plus end, which has a greater binding affinity to ATP-bound G-actin
after addition, the G-actin monomers hydrolyse the ATP slowly, so that the minus end is made up of ADP-bound G-actin, which can dissociate (this is called treadmilling)
however the rate of filament assembly and disassembly is normally controlled by actin-binding proteins, by:
nucleation- holding the three monomers together to allow for self-assembly
capping- binding to the plus end to prvent further polymerisation
severing- cutting the filament to induce depolymerisation
sequestering- binding to monomers to prevent incorporation
bundling- bind to actin filaments to connect them
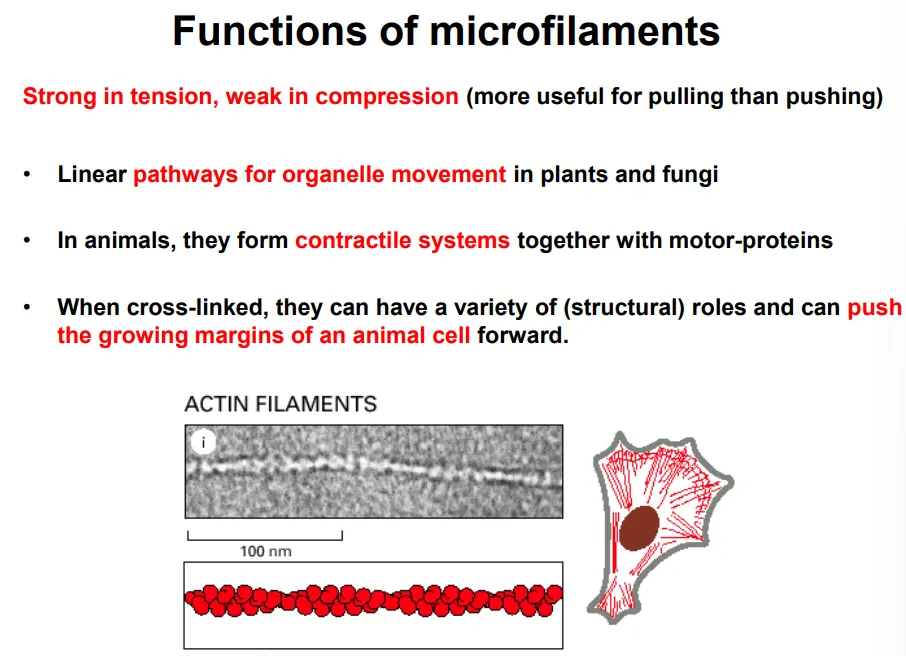
what are the functions of actin filaments?
stronger in tension than compression (used for pulling)
in conjunction with myosin motor proteins, they are used for:
organelle movement in plants and fungi
changing the cell shape in contractile systems in animals, eg:
sarcomeres in muscle cells
contractile rings in cytokinesis
stress fibres anchored to focal adhesions
cell movement
how are organelles moved around the cell by actin filaments?
in plants and fungi, the large vacuole means organelles and vesicles must be moved around by myosin motor proteins, causing the cytoplasm to also move by cytoplasmic streaming (overcomes diffusion barrier)
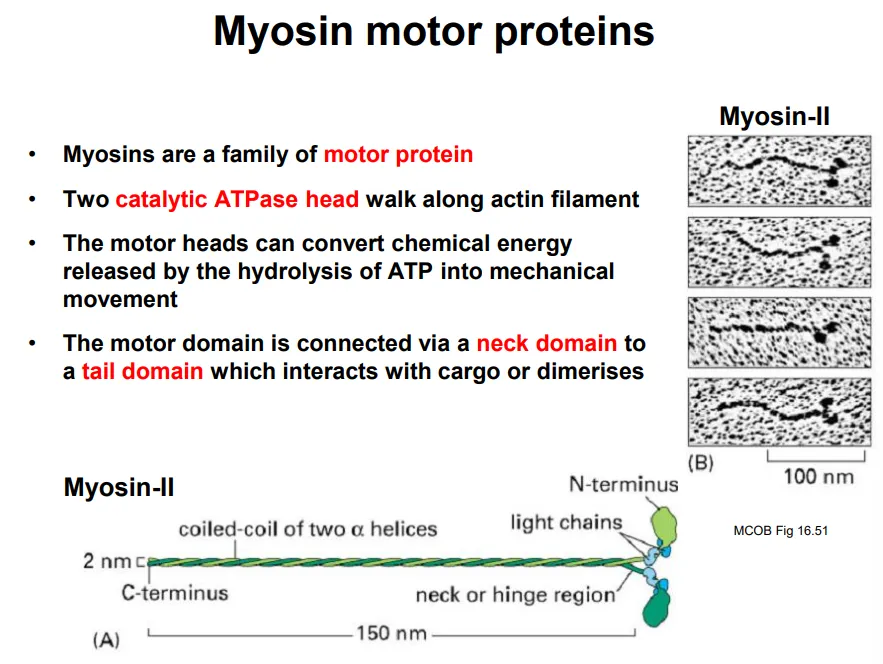
two catalytic heads walk along the actin filament in steps by conformational changes from hydrolysing ATP
this is connected to a neck domain, with a tail domain on the end to bind to cargo
how are actin filaments organised in skeletal muscle cells?
in animal skeletal muscle tissue, actin and myosin filaments (myofibrils) form contractile arrays in multinucleated syncitial cells (myoblasts fused together)
muscle myosin proteins assemble into bipolar myosin filaments (thick), where the heads project out along the length in specific radial positions (43 nm repeat)
each repeating unit is a sarcomere, which is striated:
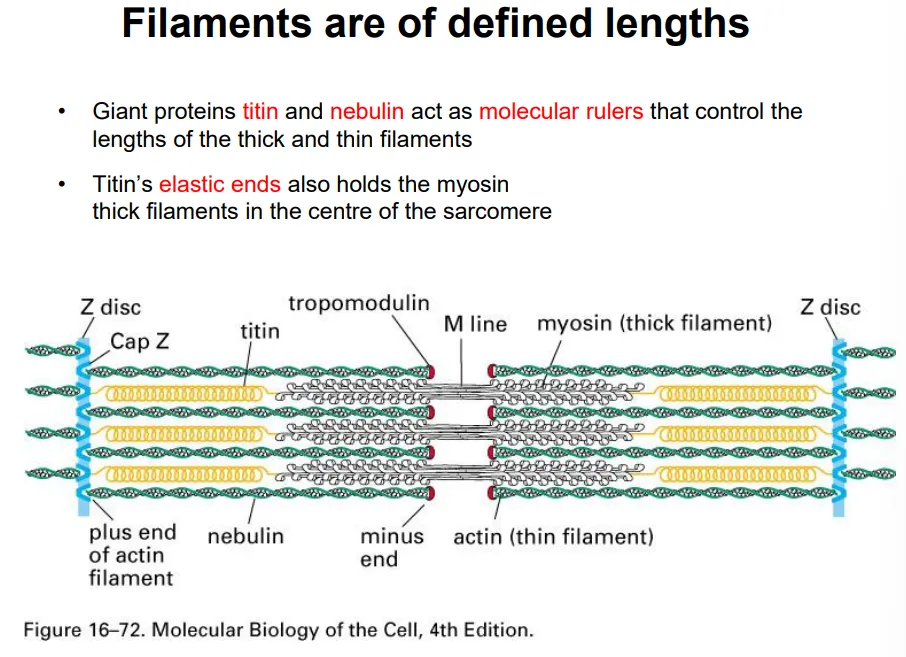
the plus ends of the actin filaments (thin) are anchored at the z disc, and the minus end is capped by tropomodulin (can’t polymerise/depolymerise)- these are all the same length due to attachment to a long nebulin protein
elastic titin proteins attach to the z disc and connect to myosin thick filaments to keep them centred along the m line
this shortens according to the sliding filament model
the thick filaments are hexagonally packed very regularly
describe how sarcomeres shorten
each sarcomere in skeletal muscle cells contracts according to the sliding filament model:
when relaxed, the titin proteins (connected to the z disc and myosin filaments) are stretched, so the actin and myosin only just overlap
when the muscle contracts, the myosin filaments walk along the actin filaments, pulling the z discs on either side towards the m line in the centre
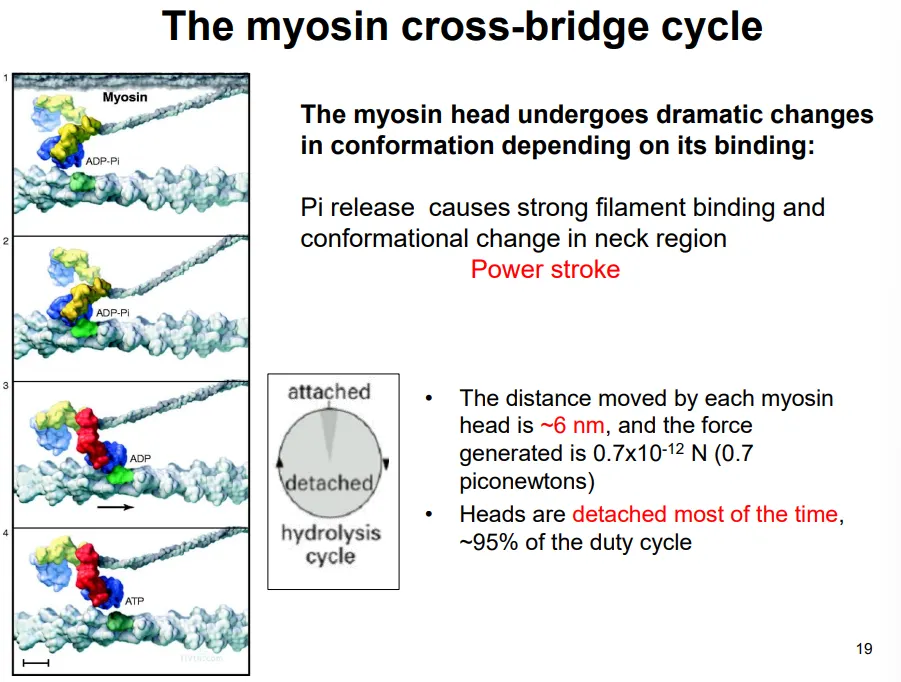
this occurs by the myosin cross-bridge cycle, driven by ATP hydrolysis:
the two myosin heads of each protein act independently
they bind to the actin and release a phosphate (from previous hydrolysis), causing the power stroke, which pulls the actin filament 6nm
the ADP dissociates, and the next ATP binds
this causes the head to detach from the actin (detached 95% of the time)
the hydrolysis of this ATP causes the head to move back to its original conformation
the spacing of the myosin heads doesn’t match up with that of the myosin binding sites of actin, and the nine radial positions of the myosin heads doesn’t match up with the hexagonal packing of actin around each thick filament
this means that not all the myosin heads can bind to actin at one time- this means there are always some heads attached to maintain tension
how is muscle contraction controlled?
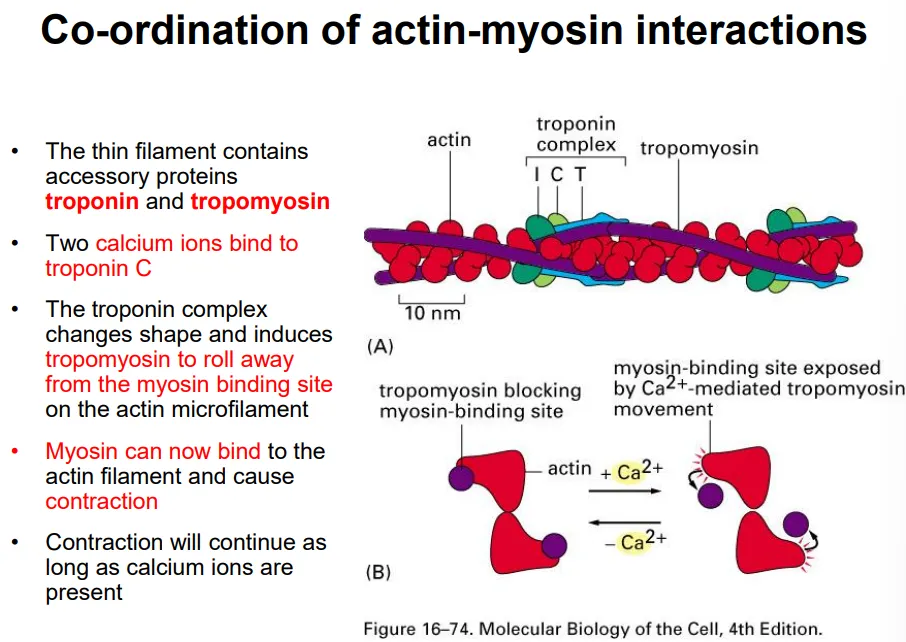
tropomyosin proteins are wrapped around the thin actin filament and cover the myosin binding sites
when an action potential arrives at the cell, it is transmitted through invaginations of the plasma membrane
this causes the voltage-gated release of calcium ions from the sarcoplasmic reticulum
two calcium ions bind to a troponin complex, which changes shape and causes the tropomyosin to detach
this allows myosin to bind to the actin filament, causing contraction by ATP hydrolysis as long as the calcium ions are present
what are the three actin filament organisations in crawling cells?
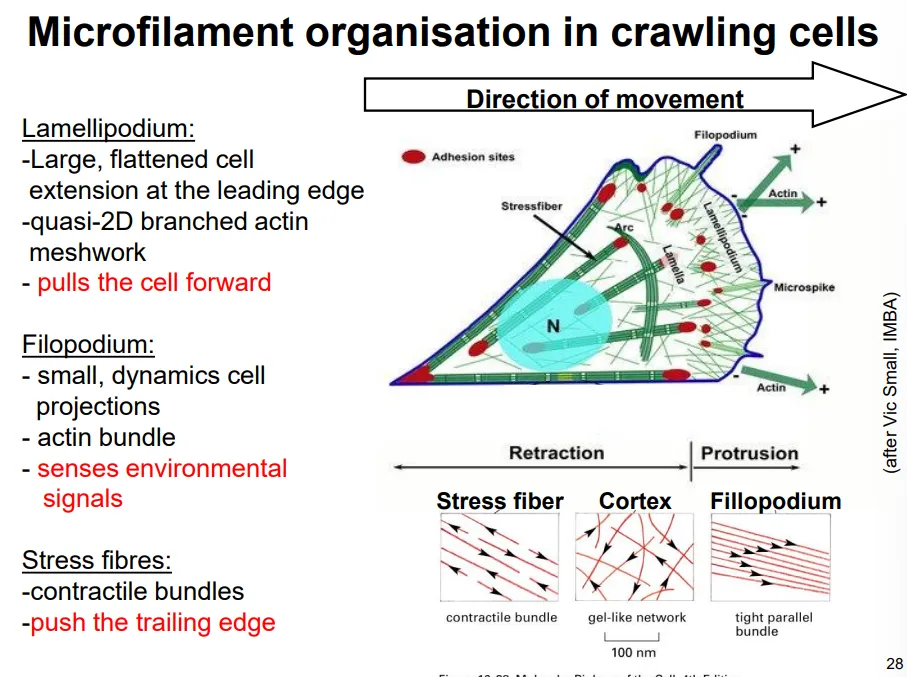
in the lamellipodia, the actin is branched at 70 degree angles due to the binding of actin-related proteins (ARPs) that facilitate new filaments forming, by holding together a trimer
this extension occurs via a treadmilling model, where the actin filaments grow a short distance before being capped, at which point they act as a site for new filaments to be assembled, while the older end of the network is disassembled by severing proteins
new focal adhesion points will be made and stress fibres attached, which will contract when this becomes the trailing end
this protrusion force is provided purely by actin polymerisation, not by myosin II
how are the different kinds of actin structures initiated?
GTP-binding proteins (G proteins) spatially organise actin polymerisation:
Rho stimulates stress fibre production
Rac stimulates lamellipodia extension
Cdc-42 stimulates filopodia formation
these are spatially separated around the cell in the relevant areas, to promote each actin structure
these act as molecular switches, active when bound to GTP, which they hydrolyse to become inactive
how are cytoskeletal elements interconnected?
many processes require the coordination of multiple cytoskeletal components
they can be connected by different crosslinkers to coordinate them:
static crosslinkers- neither can move
motor protein complexes- both proteins can move along their elements
motor protein-binding protein complexes- one is fixed, one can slide