GN Exam 4
1/73
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
74 Terms
How do you calculate allelic frequencies for a population?
f(x)=(#x alleles)/(total # alleles)=
f(y)=1-f(M)=
How do you calculate genotypic frequencies for a population?
f(xx)=#xx/total=
f(yy)=#yy/total=
f(xy)=#xy/total=
Hardy Weinberg Law
allele and genotypic frequencies will arrive at and remain at equilibrium frequencies after one generation of random mating if all assumptions are met
What are the 5 assumptions that need to be met?
1) infinitely large population
2) random mating
3) no selection (all genotypes are equally fit)
4) no migration
5) no mutation
Whats the formula for genotypic frequencies at equilibrium
p²AA+2pqAa+q²aa
What do p and q represent in this formula regarding A and a alleles?
the frequencies
If the gametic array was 0.6A and 0.4a, what is p and q?
p=0.6 and q=0.4
Set up a punnett square using these values and calculate/
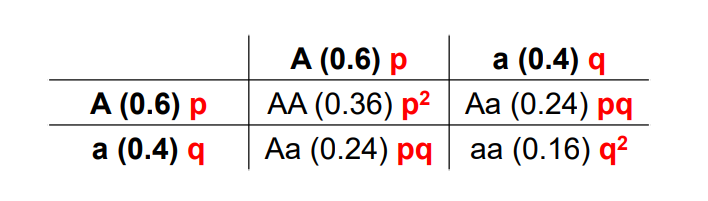

Plug values into formula (remember according to Hardy, the values you get will stay the same forever in that population)
0.36AA+0.48Aa+0.16aa
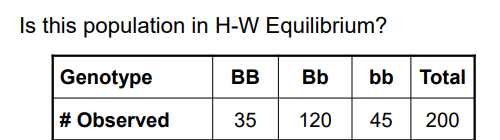
Use a chi-square test to solve this Hardy-Weinberg Equilibrium problem.
Step 1: calculate the allele frequencies so that expected genotypic frequencies can be obtained
f(B)=p=[(35×2)+120]/400= 0.475
f(b)=q=1-0.475=0.525
Step 2: calculate the expected Genotypic frequencies if population is in H-W equilibrium
BB= p²=(0.475)²=0.226
Bb= 2pq=2(0.475)(0.525)=0.499
bb= q²=(0.525)²=0.276
Step 3: use the expected genotypic frequencies to find expected counts of each allele in the population.
BB= 0.226×200=45.12
Bb= 0.499×200=99.76
bb=0.276×200=55.12
Step 4: calculate total (o-e²)/e
BB= (35-45.12)²/45.12= 2.27
Bb= (120-99.76)²/99.76= 4.106
bb= (45-55.12²)/55.12= 1.858
Total= 8.234
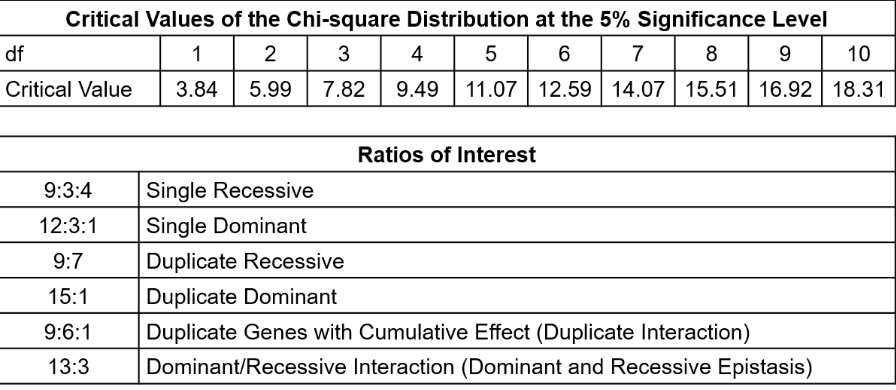
Use chi-square table to find critical value and compare with chi-square statistic.
DF= (# of genotypes-# of alleles)=3-2=1
1 DF= 3.84
8.24>3.84
Whats the conclusion for this test?
We reject Ho. At the 5% signifigance level, the B locus is not consistent with Hardy-Weinberg proportions.
How can you calculate allele frequencies from genotypic frequencies? Say you have genotypic array of 0.12MM+0.43MN+0.45NN.
f(M)=f(MM)+1/2 f(MN)=0.335
f(N)=f(NN)+1/2 f(MN)=0.665
Assume HW equilibrium to find frequencies of carries (Aa) in a population. The frequency of albinism (autosomal recessive aa) in a certain population is 1 in 10,000. What proportion of the population is expected to be heterozygous for albinism?
Albinism=aa
Frequency= 1/10,000= 0.0001
aa q²= 0.0001
q=sqrt of 0.0001= 0.01
p= 1-0.01= 0.99
Find heterozygotes 2pq= 2(0.99)(0.01)
Fitness
the ability to survive and reproduce
Directional selection
favors one extreme or the other so becomes homozygous after an infinite # of generations
Disruptive selection
advantage for both extremes so both alleles remain in population
Stabilizing selection
heterozygotes are favored so both alleles remain in population
If we’re given a starting population with starting genotypic frequencies and fitness values for each genotype, what can we determine?
the frequency of survivors after selection and the frequency of the offspring of these survivors
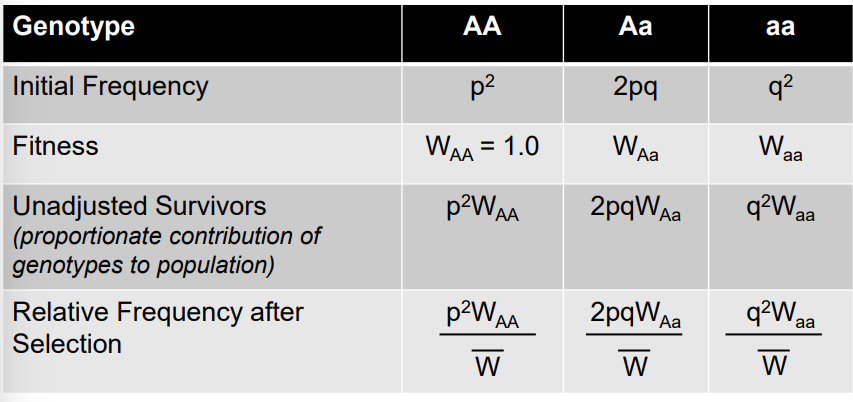
Formula for average fitness
W = p²WAA + 2pqWAa + q²Waa
Selection sample problem: What is the frequency of A after selection? AA=8, aa= 18, Aa= 24
Fitness W: AA=1.0. Aa=0.875, aa=0.5
1) total population is 50
2) f(AA)=8/50=0.16
f(Aa)= 24/50= 0.48
f(aa)=18/50=0.36
3) p=f(A)=f(AA)+1/2 f(Aa)= 0.16 + 1/2(0.48)= 0.4
q=f(a)=1-p=1-0.4=0.6
4) AA survivors: 0.16 × 1.0= 0.16
Aa survivors: 0.48 × 0.875= 0.42
aa survivors: 0.35 × 0.5= 0.18
5) W= 0.16+0.42+0.18=0.76
6) f(AA)=0.16/0.76= 0.2105
f(Aa)= 0.42/0.76= 0.5526
f(aa)= 0.18/0.76= 0.2368
7) f(A)= 0.2105 + 1/2(0.5526)= 0.487!!!
How does natural selection affect the elimination, maintenance, or increase in the frequency of various types of alleles?
If it reduces fitness, an allele can be eliminated. An allele can increase in frequency if it gives a survival advantage. Alleles are maintained when heterozygote has an advantage.
Genetic drift
random variation in gene frequency from generation to generation due to small population size leading to random fixation or loss of alleles over time
Founder population
small population that colonizes a new area
Genetic bottleneck
population’s size suddenly drops due to environmental events leaving a small and genetically less diverse population
Inbreeding
mating between relatives is more likley in a small population
Assortative mating (positive and negative)
mating based on phenotype where positive mating is when like individuals get together while negative mating is when opposite attracts keeping diversity in pop
Explain how the effects of assortative mating are different than the effects of inbreeding on genetic diversity.
Interbreeding increases the chance of being homozygous by receiving the same allele from a common ancestor. In other words, you’re matching alleles that are identical. Assortative mating, on the other hand, increases homozygosity for specific traits. If we mate tall with tall, they don’t have to be related, but alleles will look similar and most likely produce a tall offspring.
Why are rare recessive disorders more frequent in inbred populations?
If the common ancestor carries a rare allele and both relatives inherit that allele, if they mate, that child has a higher chance of getting one “a” from mom and one “a” from dad.
What does the variable F represent in inbreeding?
inbreeding coefficient which measures the probability that 2 alleles are identical by descent
What does it mean when F=0 and when 0<F<1?
F=0 when there is no inbreeding and 0<F<1 when inbreeding occurs
Whats the genotypic array formula for a population that undergoes systematic system of inbreeding?
(p2 + Fpq) AA + 2(1 – F)pq Aa + (q2 + Fpq) aa
How much inbreeding is occurring in a population? Assume that this population is undergoing a systematic inbreeding scheme and currently has a genotypic array of 0.60 AA + 0.20 Aa + 0.20 aa. Calculate F.
1) p=f(A)=f(AA)+1/2(Aa)→ 0.60+1/2(0.20)= 0.70
q=1-0.70=0.30
2) solve for F using part of the genotypic array equation
2(1-F)pq=Aa→ 2(1-F)(0.70 × 0.30)=0.20→ 2(1-F)(0.21)=0.20→ F=0.524
What does our calculated F=0.524 represent?
There’s about 52% chance a random individual’s 2 alleles at a locus are identical because of inbreeding
When individuals migrate from mainland to island, what 2 formulas are used?
mainland PA+Qa and island pA+qa
For a migration rate problem, identify these variables: P, m, p, 1-m, p’
P= frequency of A allele in mainland population
m=proportion of migrants after immigration occurred
p=frequency of A allele on island initially
1-m= probability that a parent will come from the island
p’= frequency of A allele after migration
If you initially have 175 individuals and 25 migrate in, whats your m?
immigrants/total→ 25/(175+25)=0.125→ 12.5%
Whats the formula for p’?
(1-m)p+mP
Migration sample problem: 200 immigrants migrate from mainland to join 1000 natives on the island.
starts here
Calculate m and explain what it means
200/1200=16.67% → island gets immigrants from mainland at a rate equal to 16.67% of total island population per generation
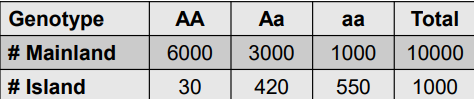
Use chart to calculate this:
• Mainland population: P = f(A) =
• Mainland genotypic array :
• Island initially: p = f(A) =
• Island genotypic array :
Mainland population: P = f(A) =
Evolution and Phylogeny
starts here
What 3 things can evolution lead to?
1) new species forming known as emergence
2) species becoming different known as divergence
3) species disappearing known as extinction
Reproductive isolation
species become distinct when they no longer exchange genes
What are 2 reasons why reproductive isolation occurs?
1) they don’t choose to mate with each other or can’t mate with each other (pre zygotic)
2) their progeny are sterile or inviable (postzygotic)
Describe the prezygotic mechanism.
If populations mate at different times, use different behaviors, or occupy different habitats, they never produce a zygote. Therefore, genetic difference keep accumulating and speciation continues unchecked.
Describe the postzygotic mechanism.
If zygote dies early, is sterile, or weak, gene flow is still blocked which reinforces separation and completes speciation.
Biological species concept
members of a species are capable of inter-mating and producing fertile progeny
Allopatric speciation
geographic barrier initiates speciation by blocking gene flow
How does allopatric speciation arise? Use Darwin’s finches example.
Ancestral species of finches migrated to Galapagos Islands and, as populations separated on different islands, their beak shape changed due to available food source.
Sympatric speciation
arises within a single interbreeding population without geographical barriers to gene flow
How does sympatric speciation arise? Use apple maggot fly example.
Original flies fed on hawthorn fruit, but mutation allowed some to feed on apples. The apple flies mostly mate on apple trees, and the hawthorn fly mates on hawthorn trees. They rarely interbreed
Phylogenetics
study of the relationship among and between species, individuals, or genes/alleles based on their characteristics
OTUs (operational taxonomic units)
what’s being compared in a phylogenetic tree such as a species, population, or single organism
Describe the steps for creating a distance matrix from OTU sequence data.
1) identify homologous genes and properly align them
2) calculate distances based on fraction of nucleotides that are different between OTUs using formula distance/total # of genes x 1000
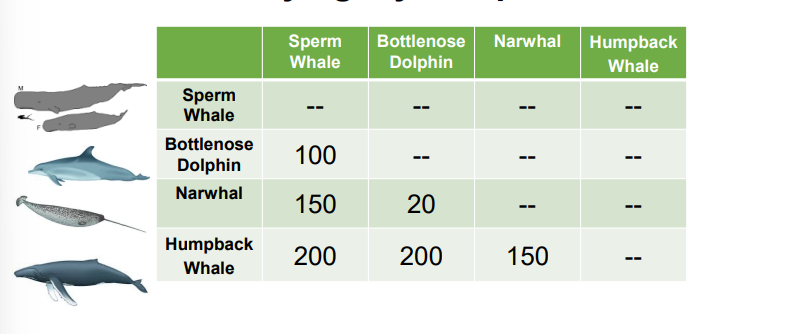
Phylogeny sample problem
Sperm whales and bottlenose dolphins have 100 differences. Bottlenose dolphin and narwhal have 20 differences. So, bottlenose dolphin and narwhal are most similar. Make a node with these two groups together first. Find next smallest distance, which is sperm whale and bottlenose. Make that the next node. ???
Anagenesis
evolution within a lineage over time
Cladogenesis
splitting of one lineage into 2, where, once cladogenesis occurs, the branches evolve separately from each other, leading to biological diversity
Paralogs
homologous sequences found in the same species and arrive through gene duplication
Orthologs
homologous sequences found in different species
True or false: Not all DNA evolves at the same speed, some parts change fast and other parts are strongly protected.
True
Where are highest mutation rates observed?
in regions of the genome that have the least effect on function
Explain why rates of nucleotide substitutions vary among different parts of a gene.
The 5’ flanking region (promotor) controls gene expression, so mutations can change gene expression levels. Therefore, there’s lower substitution rates. The UTRs, not translated into protein, regulate RNA stability and translation efficiency so mutations can alter protein levels. Therefore, there’s moderate substitution rates. Since introns get removed and pseudogenes are nonfunctional, they have the highest substitution rate.
Molecular clock
way to estimate when two species diverged by comparing their DNA or protein sequences
Explain how the molecular clock is used in studies of evolution.
Scientists look at the same genes in 2 species. They count the differences where more mutations means more time since divergence and less mutation means more recent split. If scientists know how fast mutations accumulate, they can convert differences into time.
Cancer Genetics
starts here
Cancer cells can bypass cell division checkpoints. lndicate the three key checkpoints in cell cycle control.
G1/S