2. RNA Synthesis (Transcription)
1/47
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
48 Terms
What is transcription?
Process of synthesizing RNA from a DNA template using complementary base pairing
What is the central dogma?
DNA → RNA → Protein.
What is the structure of prokaryotic RNA polymerase?
Core enzyme = α₂ββ’ω; + σ factor → holoenzyme (required for initiation).
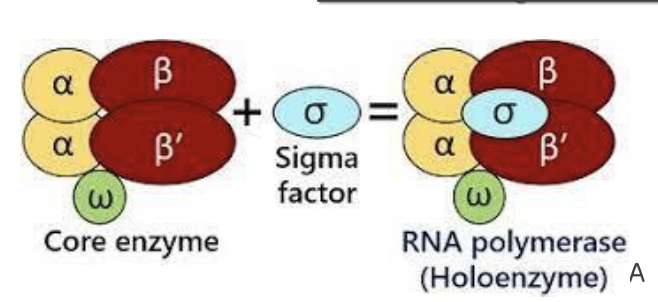
What are the functions of RNA polymerase subunits?
α (2): assembly + promoter/regulatory interaction
β: catalytic activity (RNA synthesis)
β’: DNA binding
ω: stability
What is the role of the sigma factor?
Recognizes -10 and -35 promoter regions, directs RNA polymerase to start site, then dissociates after initiation (~10 nt).
What does RNA Polymerase I do?
Synthesizes rRNA
What does RNA Polymerase II do?
mRNA + snRNA
What does RNA Polymerase III do?
tRNA + 5S rRNA
What is mRNA?
Carries genetic info from DNA
amino acid Template for protein synthesis
In eukaryotes → made by RNA polymerase II
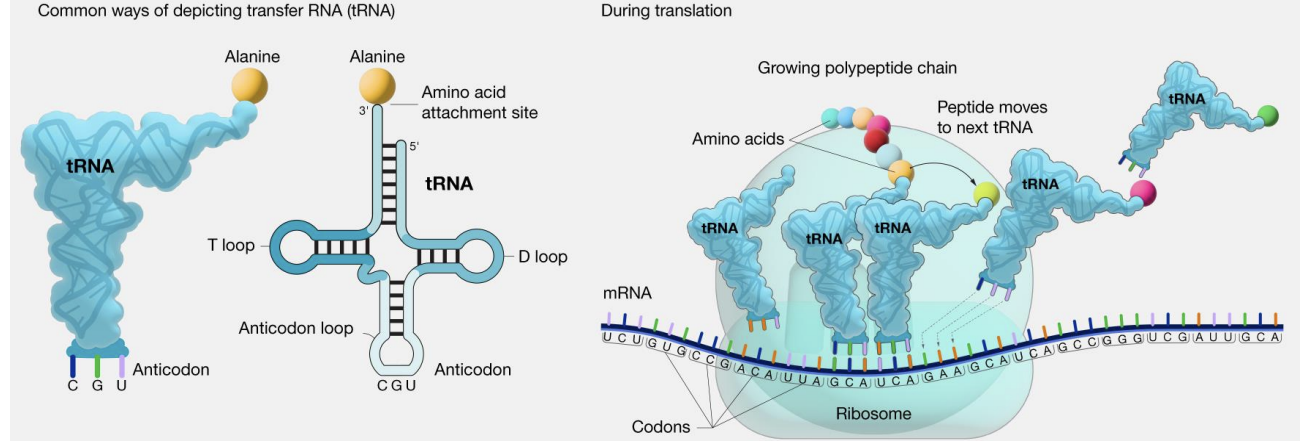
Function of tRNA?
curved croissant
Transfers amino acids to ribosome to make proteins
anticodon on tRNA matches/reads the codon on the mRNA
Synthesized by RNA polymerase III
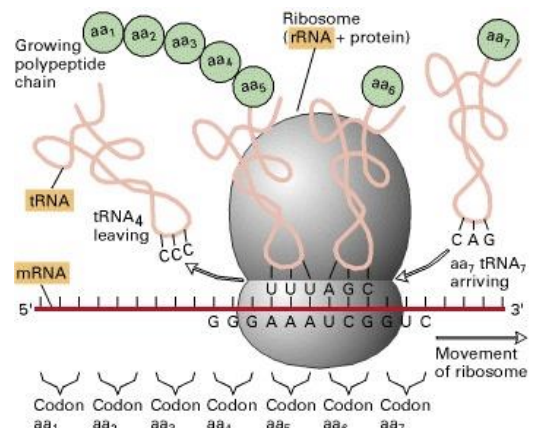
Function of rRNA?
Structural + catalytic component of ribosome
measured in Svedburg units (how fast a particle sediments during centrifugation)
~60% rRNA, 40% protein
Synthesized by RNA pol I (mostly) + III
Key prokaryotic promoter elements in prokaryotes?
-7 to -10 nt (Pribnow/TATAAT)
-35 box at -35 region (TTGACA)
upstream promoter (UP) element (A-T rich)….. found in highly expressed genes, bound by a subunit of RNA pol
What factor recognizes prokaryotic promoters?
σ factor
Other regulatory sequences in prokaryotes? to control transcription
Activator binding site
Operator (repressor binding site)
Key elements in eukaryotic promoters?
TATA box (core promoter)
CAAT box (high transcription genes)
GC box (↑ transcription, may lack TATA)
bind transcription factors and recruit RNA polymerase
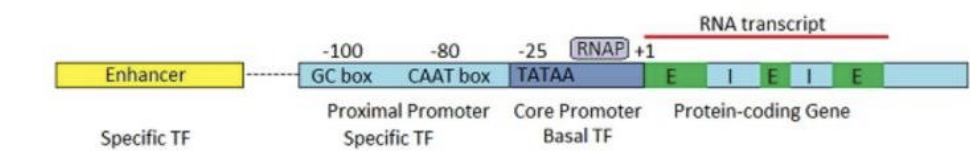
Where are eukaryotic promoters located?
~25–200 bp upstream of transcription start
Difference between enhancers and silencers?
Enhancers → ↑ transcription (activators bind)
Silencers → ↓ transcription (repressors bind)
Can be far away (upstream/downstream)
What are CpG islands?
CG-rich promoter regions
Often unmethylated in active genes
Effect of CpG methylation?
Represses transcription by proteins that bind to methylated CpG islands
Blocks transcription machinery
Seen in cancer (tumor suppressor silencing)
Effects of histone modifications?
Methylation → heterochromatin (condensed) → ↓ transcription
Acetylation → euchromatin (relaxed) → ↑ transcription
Coding vs template strand?
Coding (sense): same sequence as RNA. DNA = RNA.
non-coding Template (antisense): used for transcription
Steps of transcription?
Initiation
Elongation
Termination
Steps of initiation in prokaryotes?
RNA pol holoenzyme binds promoter → closed complex
DNA unwinds → open complex
Transcription bubble forms
σ factor released
How does prokaryotic elongation occur?
RNA pol moves 3’→5’ on DNA
RNA synthesized 5’→3’
Bubble ~8–9 nucleotides
Two types of prokaryotic termination?
Rho-independent
Rho-dependent
Rho-independent
Q: Mechanism?
Hairpin loop forms 15-20 nt before end
U-rich region weakens binding
RNA dissociates
Rho-dependent
Q: Mechanism?
Rho binds rut site (rho utilization site)
Moves along RNA (ATP-dependent)
Detaches mRNA from polymerase
What is the transcription bubble and why is it important?
A small unwound region of DNA where transcription occurs
Size: ~8–9 nt (prokaryotes), ~17 bp (eukaryotes)
RNA polymerase keeps it open while moving along DNA
Allows base pairing between template DNA and growing RNA
DNA re-anneals behind the polymerase
What is required for eukaryotic initiation?
Transcription factors
Pre-initiation complex
RNA pol II recruitment
Key features of elongation in eukaryotes?
RNA pol II phosphorylation required
initiation factors shed
~17 bp transcription bubble
Produces pre-mRNA
What does Actinomycin D do?
Intercalates DNA
Blocks elongation in proks and euks
Used in chemotherapy
Function of 5’ cap?
Stability
Nuclear export
Translation initiation
Function of poly(A) tail?
Stability
Translation efficiency
Degradation when shortened in bacteria
What is mRNA splicing?
Removal of introns from pre-mRNA
What is the spliceosome?
ribonucleoprotein complex: snRNA + proteins (snRNPs)
Removes introns
What is the difference between introns and exons?
Introns → non-coding sequences
Removed during splicing
Not translated into protein
Exons → coding sequences
Kept in mature mRNA
Translated into protein
👉 Memory tip:
Exons = Expressed
Introns = In between (cut out)
Major spliceosome components?
U1, U2, U4, U5, U6
>150 proteins
▪ Introns 5’ end splice site GU; 3’ end splice site AG
Key splice site sequences?
5’ → GU
3’ → AG
Steps of splicing?
U1 binds 5’ site
U2 binds branch point
U4/U5/U6 assemble
Introns removed
Exons joined
Why is alternative splicing important?
1 gene → multiple proteins
~90–95% human genes use it
‘3 UTR and ‘5UTR
Role of UTRs?
Control mRNA stability, localization, translation. recognized by miRNA, their associated machinery, and RNA binding proteins
Steps in tRNA processing?
Remove 5’ leader
Remove introns
Add CCA tail
Base modifications
What is copy number?
Number of transcripts per cell
Does copy number vary?
Yes — depends on gene function and cell needs
Which strand is used for transcription?
Template (antisense) strand
What is the first RNA product in eukaryotes?
Pre-mRNA (contains introns + exons)
Does mature mRNA contain introns?
❌ No
Which element can be far from gene and increases transcription?
Enhancer