DNA last 2 lectures
1/52
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
53 Terms
Multiple depositors, low-level contributor, secondary transfer
Name the different types of mixture DNA
_______ ___________: Two or more individuals physically deposit biological material at the same location
___-_____ __________: One contributor is present at a very low quantity, difficult to detect
_________ ________: DNA is indirectly deposited require an intermediate surface
Not all contributors are equally visible
Numbers of contributors, contributor ratio, allele sharing, and total DNA quantity
What are factors that affect mixture detection?
D18S51
What is the high diversity locus that makes the most informative markers for mixture interpretation (about 50 possible alleles)?
Major contributor
Individuals who contributed more DNA
Minor contributor
individuals who contributed less DNA
Mixture ratio
Estimate the mixture ratio - the relative amount of DNA each person contributed
Large differences in peak height = Larger major : Minor Ratio
5:1 Noticeably smaller peaks present- causes minor still detectable interpretation
10-20:1 some minor peaks missing - causes allele dropout
Detection depends on Peak height (RFU thresholds)
If you do not see the minor contributor, it does not mean they are not there.
70
Small peak / Large Peak > __% indicates a single source profile
if the ratio is below __%, indicates mixture; degradation; Inhibition
10-15
Normal stutter peak: small; <__-__% of the main peak; Always appears in a predicable position (n-1)
Mixture peak: Looks like stutter but larger
Not all small peaks are stutter artifacts.
Stutter can hide or mimic minor contributors, making mixture interpretation more difficult
Steps approach to mixture interpretation
Is it a mixture? more than 2 peaks at a locus; big peak height ratio; Unexpected allele patterns
Label the peaks: above analytical threshold
How Many contributors. Max alleles / 2 = minimum contributors
Who is Major and Minor, Tall peaks and shorter peaks
What genotype combination make sense
Compare to known profiles
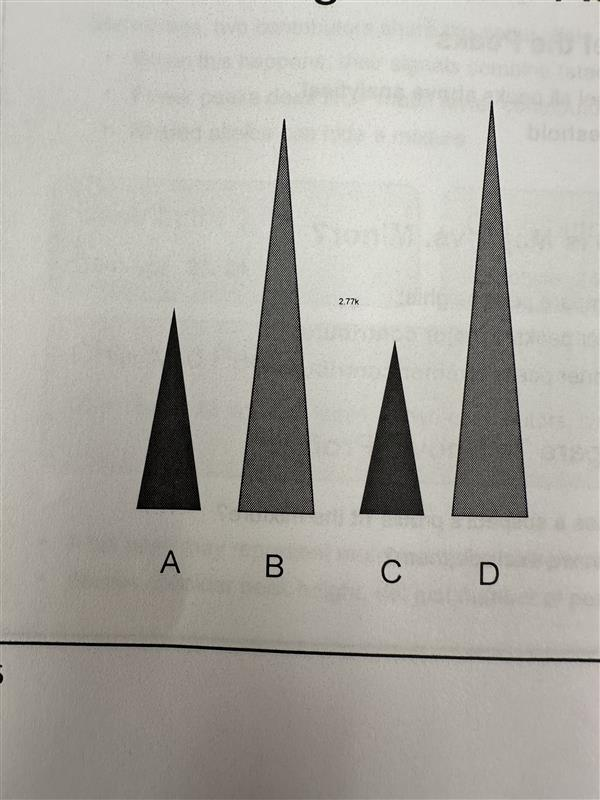
Calculating mixture ratio
Major contributor (B+D)
Minor contributor (A+C)
Ratio = (B+D) / (A+C) = 2.3 : 1
Round to approximately 2:1 - The major contributor contributes roughly twice the DNA of the minor contributor
Allele sharing, more than two contributors, low template DNA, stochastic effects, degradation, and inhibition
What are some challenges in Mixture interpretation?
Probabilistic genotyping
What type of interpretation method is used by software like STRmix to analyze complex DNA mixtures?
STRmix
This software is important because it uses real forensic labs, accepted in courts and helps analyze mixtures that are too complex to interpret manually
It tests millions of possible genotype combinations and uses the entire electropherogram. Accounts for peak height, dropout and stutter
Preforms interpretations faster, more consistently and with statistics
Single nucleotide polymorphisms (SNP)
A single base change in DNA occurs at a specific location
There are so many of them
Advantages:
Extreme Abundance: every 1000 bp
Genome-Wide Coverage - located across all chromosomes
Give a big picture of DNA
Especially useful when DNA is damaged or limited
Disadvantages:
Low power per marker
Need many SNPs, 50-100+ markers; more cost
Mixtures are Harder- Limited variation and difficult to separate contributors
Limited Databases
Technology Enables: Next-generation sequencing (NGS) panels
Reading results:
Each peak = a base (ATCG), each color = a different base
One peak = homozygous; Two Peaks = Heterozygous
STR vs SNP
Fill in the blank…
___: Repeat length, Often 10+ per locus, high distinguish power, length (100-400+ bp), stutter artifacts: present complicates interpretation. Databases: CODIS, NDNAD
___: Single nucleotide substitution. only 2 bi-allelic, Lower discrimination power on one locus, Very small (<100bp), no stutter artifacts, clear signal, No standard forensic databases.
Direct sequencing, primer extension (SNaPshot)
This SNP method reads the actual DNA sequence. Use next-generation sequencing (NGS).
The other common SNP method is ______ _______ (_______) that targets specific SNPs. Targets a single base using fluorescently labeled dNTPs.
Less commonly used: Microarrays and pyrosequencing
SNaPshot
A method to identify SNP alleles and looks at one base in DNA
DNA region is amplified (PCR), primer binds next to the SNP, One base is added (ddNTP) and that base is fluorescently labeled.
Each base=different color
Reads only one base. Color = SNP allele
Protocol:
PCR - Clean up extra primers and ddNTP - Extension add one base - electrophoresis (read color)
Massively parallel sequencing (MPS)
What type of sequencing technology analyzes thousands to millions of fragments at once?
Ancestry informative markers (AIMS)
What are specific single-nucleotide polymorphisms (SNPs) that show significantly different frequencies between human populations, making them powerful tools for estimating biogeographical ancestry.
Internal size standard
What is used during capillary electrophoresis to ensure accurate sizing of DNA fragments?
Amalyase
What enzyme is commonly used to identify saliva?
Acid phosphatase (AP)
What enzyme is targeted in presumptive tests for semen?
Test for contamination
What is the purpose of an extraction control (reagent blank)?
variable number tandem repeats (VNTR)
What type of repeated DNA sequence was used in early DNA fingerprinting before STR?
Microvariant
Which allele call contains an incomplete repeat (eg 9.3)
Normalization
What is it called when you adjust DNA input volume to ensure consistent amplification across samples?
Threshold cycle (Ct)
What value in qPCR represents the cycle at which fluorescence exceeds the detection threshold?
Low DNA level
What does a high Ct value indicate about a DNA sample?
(hint: it took longer for sample to hit the threshold)
Stutter
What is the most common biological artifact in STR analysis caused by strand slippage?
Proteinase K
What enzyme is used to breakdown proteins during DNA extraction?
Measure inhibition
What is the purpose of an internal positive control (IPC) in qPCR? (quantitation)
Linkage equillibrium
What conditions must exist between loci to justify multiplying probabilities across STR markers?
Porous
What type of surface may initially capture more DNA but may make recovery hard?
Solid phase extraction
DNA extraction method that binds DNA to solid surface like silica or magnetic beads
What happens if too much DNA is added to PCR reaction
Overamplification and split peaks
DTT
What is added to break disulfide bonds in sperm cells during differential extraction?
Product rule
Which rule allows forensic scientists to multiply genotype frequencies?
Background DNA
What is the term for DNA already present at the crime scene?
RMP
What statistical value represents how rare a DNA profile is within a population?
Pull-up
What artifact occurs when signal from one dye appears in another color channel?
Analytical
Which threshold separates true signal from background noise in STR analysis?
Heat stable
Why is Taq polymerase used in PCR instead of regular DNA polymerase?
Tetranucleotide
Which STR is preferred in forensic science due to low stutter?
GeneMapper
What software is commonly used for STR genotyping and data interpretation?
Dye-labeled primer
This type of primer is useful in STR analysis as it separates different fragment sizes for a more clear result.
Denaturing
What type of electrophoresis conditions keep DNA as single strands for better resolution?
Null allele
What is the name for an allele that fails to amplify due to a primer binding size mutation?
Fragmented (degraded)
What is it called when DNA has been broken into smaller pieces?
Stacking
What is the term for when two contributors share the same allele causing peaks to combine?
DNA sequence
What does next generation sequencing (NGS) measure that STR analysis does not?
Prosecution / defense
The likelihood ratio (LR) includes two hypothesis. What are they (in order)?
Theta (θ)
What is the statistical adjustment when there is not enough DNA data for population genotype frequencies? (hint: 0.3)
Both hypothesis are equally likely
What does a LR=1 indicate?