Unit 4.4 Investigating Biotechnology and Bioethics
1/83
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
84 Terms
What does “Biotechnology” translate into?
“Life Tools”
What is biotechnology?
Any technological application that uses biological systems/living organisms/derivatives to make or modify products or processes for specific use
What is the Barcode of Life?
Genetic sequencing used for biodiversity conservation. Species have short segments of genetic code that are the same across the whole species + different from other species. If we can map all these sequences then we can quickly identify each species simply by reading short sections of their DNA
What is a real world example of the barcode causing controversy?
NYC students proved that the fish being sold as a high price wasn’t what the label said but actually a much cheaper fish
What are some difficulties with biodiversity conservation?
Hard to identify different species, not enough trained specialists, most of the world’s biodiversity is in developing nations
How long did it take to catalogue 15% of the world’s biodiversity?
250 years!
What is a very real fear we have about the research of biodiversity?
That most species will go extinct before we can get to them
What are some pros of DNA barcoding? What are some real world examples of benefits?
Fast, cheap, accurate; pest control in agriculture, stop illegal trafficking of endangered species
Why is biotechnology controversial?
Has legal/ethical/economic/political/environmental implications. Risk vs. benefits. E.g. a new medical benefit could harm the environment to produce

Timeline: 8000 BC to 1589
Selected seeds for planting, livestock for breeding, mated different species, utilized yeast/bacteria to ferment food and drink
Timeline: 1590 to 1862
Janssen invents the microscope; van Leeuwenhoek discovered microorganisms; Mendel describes laws of inheritance; Pasteur determines fermentation is the result of bacterial metabolism
Timeline: 1919
Ereky coins “biotechnology”
Timeline: 1953
Franklin records double helix structure of DNA; Watson and Crick describe DNA’s full structure
Timeline: 1959
Use gel electrophoresis to separate large molecules (like DNA and proteins) based on their size
Timeline: 1973
Boyer and Cohen transfer genes between species (i.e. genetic engineering)
Timeline: 1978
Boyer makes synthetic insulin from genetically-engineered bacteria + recombinant DNA technology
Timeline: 1984
Mullis develops Polymerase Chain Reaction (PCR) technique that can quickly replicate small samples of DNA (→ lead to rapid DNA sequencing)
Timeline: 1987
First criminal caught and convicted using DNA fingerprinting
Timeline: 1994
Tomato that was genetically modified approved for sale
Timeline: 1996
Recombinant DNA technology used to make soybean strain resistant to herbicide
Timeline: 1997
Dolly the sheep is cloned!
Timeline: 2000
Human Genome Project. Complete DNA sequence for all 30000 human genes
Timeline: 2002
Genome of rice is sequenced
Timeline: 2004 to 2005
Stem cells used to repair human tissue
Timeline: 2006 to 2009
Epigenetics found to play important role in genetic inheritance (epigenetics control the way genes are expressed→ new insight into the causes of diseases)
Timeline: 2010
Second Genetic Code. Explains how RNA segments get spliced together to express a gene (→ good for new genetic treatments for diseases like cancer)
Timeline: 2011
Malaria vaccine→ first vaccine against parasitic infection
Timeline: 2012
Wheat genome sequenced
What are the 3 most important discoveries?
1953→ discovering double helix and full structure of DNA
1959→ Gel electrophoresis
1984→ Polymerase Chain Reaction (PCR)
What are the 5 biotechnology techniques (all related to working with DNA) ?
Extraction
Electrophoresis
Amplification
Sequencing
Recombinant DNA
#1 What is extraction (1)?
Extracting DNA from cells
#1 What is step 1 of extraction? Describe
Physically break open the cell (lysis). Cell wall/membrane/mitochondria/chloroplasts must be broken so DNA can be released. Can be ground up or blended. Purpose is to increase the surface area for the next step
#1 What is step 2 of extraction? Describe
Remove the lipid membranes that protect the DNA. Use a detergent. Lipid-rich membranes are torn apart
#1 What is step 3 of extraction? Describe
Protect DNA from degradation by enzymes. After membranes are torn apart, DNA mixes with the cytoplasm where there are evil enzymes. Add a protease to destroy the enzymes OR keep cytoplasm very cold
#1 What is step 4 of extraction? Describe
Separate DNA from the other cellular molecules using a solvent. Since DNA is soluble in water it can be separated using a water filtration process OR extracted out using alcohol since DNA is not soluble in alcohol but the other molecules are
#1 Extracting DNA contains entire genome. Is this practical?
No, usually only want to study one gene/one part of a gene
#1 How can we isolate individual genes from the chromosome? How?
Using the enzyme Restriction Endonuclease. Recognize and cut out a specific sequence of nucleotides at a precise location
#1 What is the location “called” where RE will cut out a specific gene?
“Restiction site”
#1 What is a helpful term to call RE?
Molecular scissors
#1 How could two fragments, cut by the same RE, join together? What would this new thing be called?
Each fragment has unpaired nucleotides at each sticky end and join here, even DNA from different species; “Recombinant DNA”
#1 This whole process of cutting and joining together is what profession?
Genetic engineering
#2 Why do we need Electrophoresis (2)?
Because extracted DNA is invisible to the eye, gel electrophoresis determines if the sample contains the right fragments of DNA
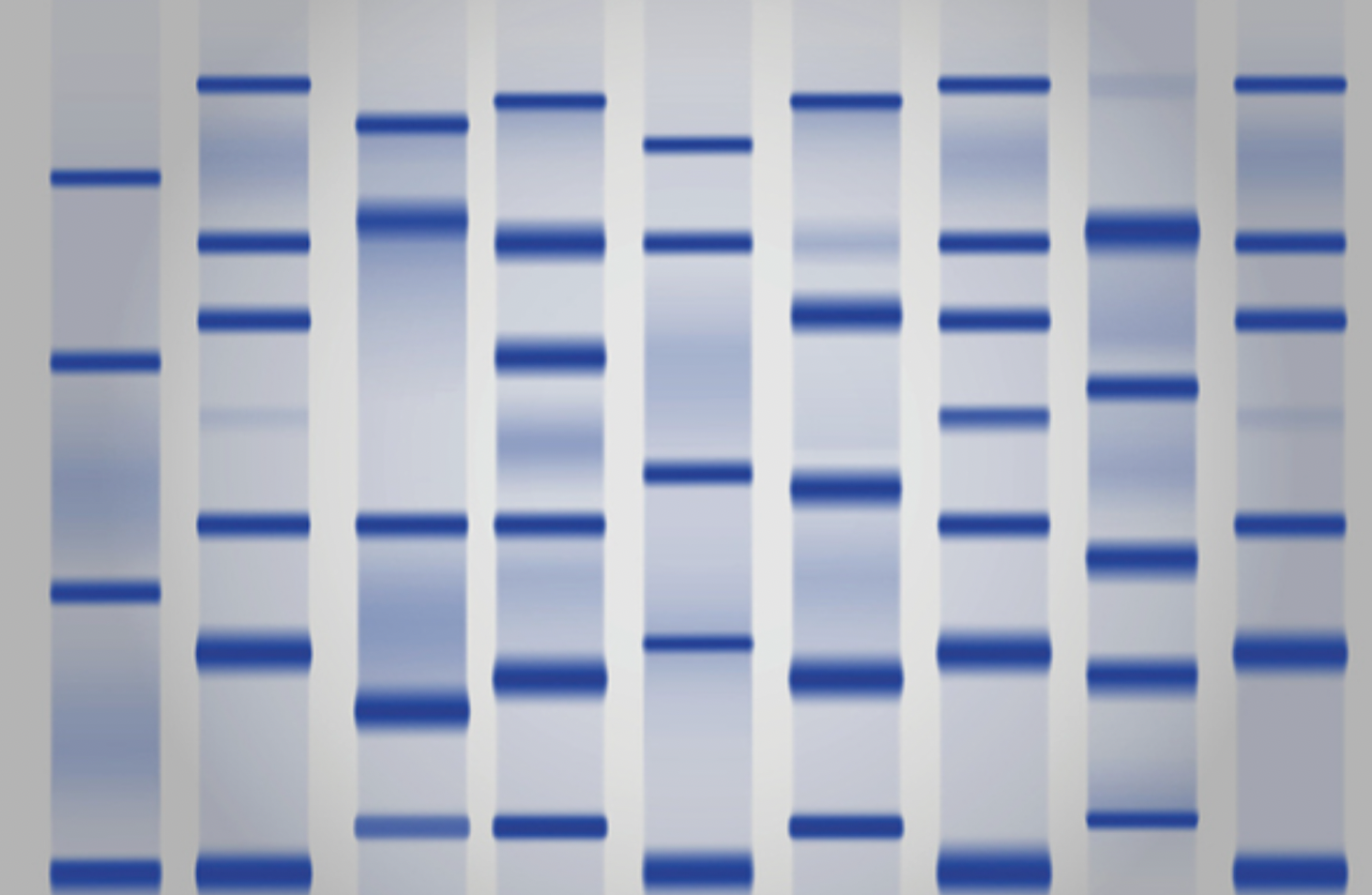

#2 How does gel electrophoresis work?
Negatively charged DNA molecules are forced through agarose gel which slows them down. They’re pulled towards the positively charged end. Large molecules are slow, small are fast. See how far the molecules have moved. Spread across the gel in bands
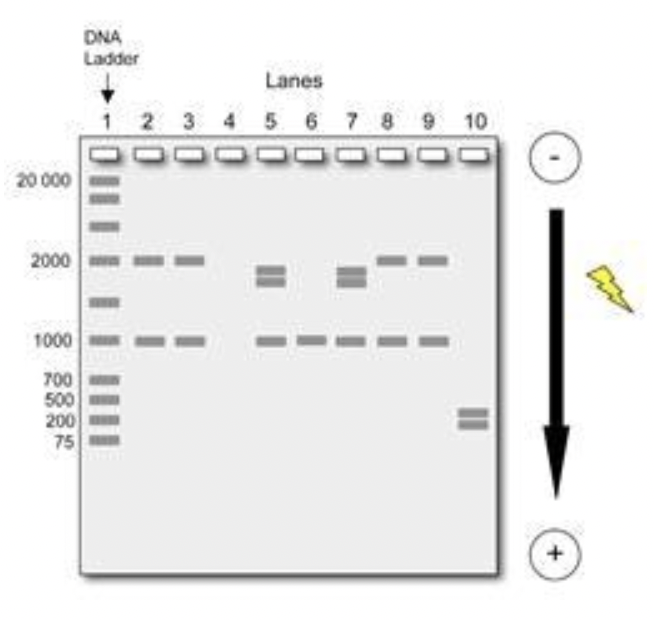
#2 How do we interpret gel electrophoresis?
There are 10 wells at the top end where the samples are deposited. The DNA moves down through the gel in lanes. The grey bands indicate the position of DNA fragments
#2 What is a DNA Ladder in electrophoresis?
The 1st lane is made up of a variety of DNA fragments with known lengths arranged from largest to smallest. Compare our sample to this one
#3 What is amplification (3)?
We need a lot of DNA material to perform genetic analyses so we need a fast way to make millions of copies of the DNA fragments in your sample → amplification!
#3 What is used for amplification? How does it do it?
Polymerase Chain Reaction (PCR); replicates specific sequences within a DNA sample
#3 What 4 things does amplification require?
DNA fragments of interest
A DNA primer (artificial, selects the correct DNA sequence)
Free DNA nucleotides (for replication)
Taq Polymerase
#3 What is Taq Polymerase?
A bacterium that lives in hot places.
#3 What is the result of amplification?
Exponential increase in the amount of DNA in the sample. Yields billions of copies in only a few hours
#3 What is a device that could cycle the temperatures for us with ease?
Thermocyclers
#4 What is gene sequencing?
Genetic blueprint→ exact order of A, T, C, G
#4 What is the purpose of gene sequencing?
To understand a gene; to identify a particular individual
#4 How many base pairs were identified in the human genome in the Human Genome Project?
All 3 billion base pairs
#4 What is gene sequencing more commonly called now?
“Chain Termination Sequencing”
#4 What are DNA oligonucleotides? Give an example
Short segments created from the original DNA fragment; if the original DNA fragment was ATGAC then the oligonucleotides are A, AT, ATG, ATGA, ATGAC
#4 What are Dideoxynucleotide Triphosphates (ddNTPs)? How are they different from regular dNTPs?
They are ‘terminators’. Like regular nucleotides but without the 3’ end bonding site for the next phosphate group. After ddNTPs are added, no other nucleotide can bind and elongation stops immediately
#4 What is the purpose of using ddNTPs?
When DNA is undergoing replication, regular nucleotides will bind, continuing elongation, and ddNTPs will bind, stopping elongation. The result is a mix of DNA pieces made up of all possible lengths. If these pieces are arranged in order by length and the last nucleotide is read, you would be reading out the original DNA fragment
#4 Once the chain termination sequencing reaction is complete, now we read the last nucleotide in each oligonucleotide. Happens in two steps. Describe the 1st step. What technique do we use?
Arrange the oligonucleotides in ascending order (smallest to largest) by length. Use capillary electrophoresis. Sample is passed through a thin tube (capillary) containing a gel which allows small molecules to move fast and large molecules to move slow. Result? Oligonucleotides gets sorted by size with the shortest coming out first
#4 Once the chain termination sequencing reaction is complete, now we read the last nucleotide in each oligonucleotide. Happens in two steps. Describe the 2nd step. What technique do we use?
Identify the nucleotides in the last position. ddNTPs need to be labelled in some way. Attach fluorescent molecules to them with a different colour for each base. When exposed to a laser they will glow giving off their specific colour. The result is 100’s or 1000’s of letters in one long string→ the DNA sequence of the original fragment
#5 What is recombinant DNA (5)? What is the process called?
Take individual genes from one genome and insert them into another genome; “recombination”
#5 What was Boyer’s amazing discovery in 1978?
Transferred human genes into a bacterium to create synthetic human insulin
#5 What is the name for a genetically modified organism that contains genes from another species?
“Transgenic”
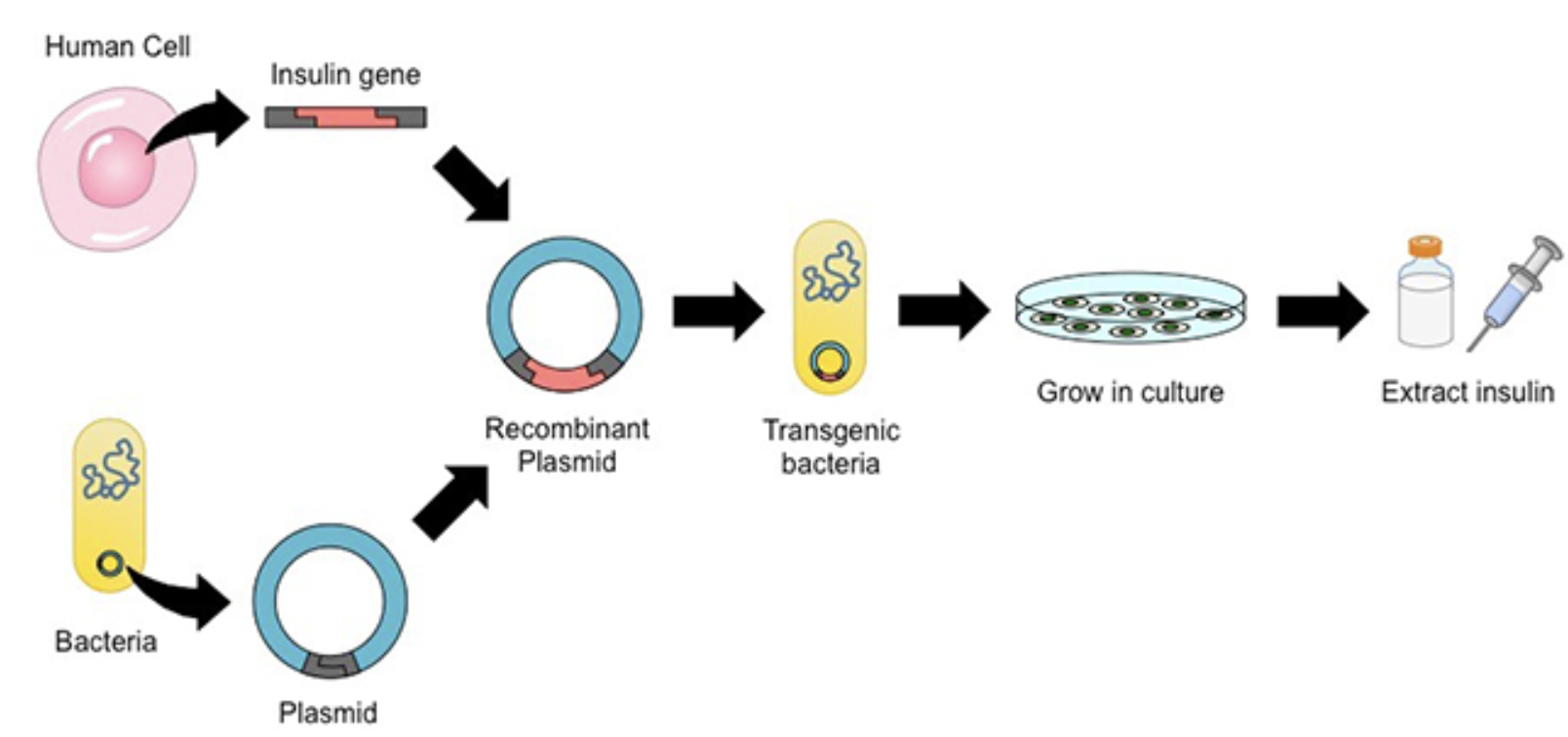
#5 In 1982, insulin hit the market- the first commercially available genetically engineered product created using recombinant DNA techniques. What are some of its pros?
Allergy-free, abundant, inexpensive
#5 Which usually gets inserted into who between prokaryotes and eukaryotes? Why?
Genomes of prokaryotes are easier to deal with so usually eukaryotic genes are inserted into their genome
#5 Genetic material in bacteria is stored in their chromosomes and where else?
In their plasmids
#5 What are plasmids?
Small circular ribbons of DNA that can move between bacteria
#5 When undergoing replication, are both the plasmid DNA and the chromosomal DNA copied?
Yes
#5 How does the enzyme Restriction Endonuclease (RE) fit in here? What enzyme is important here?
RE cut open the plasmid in the bacterium and also cuts up specific fragments of DNA in the eukaryotic cell giving them both ‘sticky ends’
Join together through when ligase guides the sticky ends together
#5 We now have a recombinant plasmid with foreign genes. What is its new name?
A “vector”
#5 What happens now to the vector?
It’s placed in a culture medium where the bacterium takes in the plasmid where it is treated like a normal part of the cell. Bacterium proceeds as usual→ makes proteins, replicates now with the new gene as a part of its permanent genetic makeup
#5 After all this, how would we get insulin (for example)?
Technicians can isolate and purify the desired protein for a specific use
What are the 2 sectors of biotechnology in Canada?
Reading DNA
Manipulating DNA
What are some examples of Canadians involved in reading DNA (1)?
DNA evidence in crimes, DNA fingerprinting, identifying genes related to learning disabilities, genetic counselling
What are some examples of Canadians involved in manipulating DNA (2)?
Vaccines, making trees with higher quality wood, bacteria that can absorb oil after an oil spill
Name the 4 steps of DNA extraction
Physically break open the cell (lysis)
Remove the lipid membrane protecting the DNA (using a detergent)
Protect the DNA from enzymes (keep cold)
Separate DNA with a solvent (alcohol)
In gel electrophoresis, the numbers indicate what?
Base pairs
In gel electrophoresis, DNA is negatively charged and the bottom is positive. What has to be applied to the agrose gel?
An electric current
Do high or low temperatures speed up the replication process?
HIGH
Recombinant DNA is the result of removing genetic material from one _____ and inserting it into another _____
Genome