Unit 12 RNA Splicing and Processing
1/170
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
171 Terms
How does mRNA processing differ between prokaryotes and eukaryotes?
Prokaryotes have very little RNA processing, so the primary transcript is mature mRNA; eukaryotic pre-mRNA is usually capped, poly-A tailed, and spliced before export as a mature mRNA
What is the primary transcript of a eukaryotic gene?
pre-mRNA and has the same organization as the gene.
What three major modifications occur to a eukaryotic pre-mRNA before it is considered mature?
It is capped, poly-A tailed, and splices
Where must mature mRNA be exported to for translation?
The cytoplasm
What is transcriptional coupling in the context of RNA processing?
It refers to enzymes for capping, tailing, and splicing being physically coupled to the transcriptional apparatus (RNAP II)
What part of RNA polymerase II (RNAP II) acts as a “loading pad” for processing factors?
The C-terminal domain (CTD)
How long is the fully extended CTD compared to the rest RNAP II
It is 10X longer
How do processing factors reach the nascent mRNA?
They “hop” onto it from the CTD
What helps the transcriptional apparatus clear the promoter?
The capping enzyme
Which amino acid residue on the CTD is phosphorylated by TFIIH during the transition to elongation?
Serine 5 (Ser5)
What specific processing enzymes bind to the Ser5-P (phosphorylated) CTD?
Capping enzymes
Which amino acid residue is phosphorylated on the elongating RNAP II later in the process?
Serine 2 (Ser2)
Which enzymes bind the Ser2/5-P CTD?
Splicing and tailing enzymes
What is usually the first nucleotide in an RNA transcript?
A pruine nucleotide triphosphate
Which mammalian enzyme is responsible for adding the 5’ cap?
Guanylyl-transferase
What substrate does guanylyl-transferase use to create the cap?
Uses GTP to create a 5’ ——5’ triphosphate linkage with the 5’ terminal purine nucleotide of the mRNA (created between the cap and the mRNA)
Which enzyme adds a methyl group to the terminal guanine of the cap?
Guanine-7-methyltransferase
Where specifically is the methyl group added on the terminal guanine?
The 7’ position
Where else can methyl groups be added to the transcript besides the 7’ position of the guanine?
To the 2’ position of downstream ribose sugars and the nitrogenous bases.
How far downstream of the initiation site does RNAP II pause to wait for capping enzymes to act”?
30 nt
Why is capping necessary?
Required for promoter clearance and transition to elongation
What is the risk of an uncapped nascent RNA
It is vulnerable to attack by exonucleases
What is Intron definition?
A recognition mechanism where the 5’ and 3’ splice sites are simultaneously recognized by components of the E complex. Introns are used for small, single-intron genes in unicellular eukaryotes.
What is Exon definition?
A recognition mechanism that takes advantage of small, consistently sized exons in multicellular eukaryotes, where introns are long and variable.
What do many sequences in introns resemble?
they resemble true splice sites
Is the paired recognition of splice site flanking efficicent?
No, it is inefficient
In exon definition, where does U1 bind relative to U2AF
U2AF binds to the 3’ splice site, and U1 binds to the 5’ splice site at the beginning of the next intron, which bridges the exon.
What associates with the complex after formation of the E complex?
other snRNPs and factors
How is the A complex formed?
U2 snRNP binds to the branch point and results in displacing BBP/SF1 and U2AF
What facilitates U2 binding to branch point?
Facilitated by base pairing between U2 snRNA and branch point consensus sequence
What is required for the formation of A complex
It requires energy, more specifically ATP hydrolysis
What forms the B1 complex?
A tri-snRNP complex composed of U5 and U4/U6 binds to the A complex
Why is the B1 complex considered the “true splicosome”?
It contains all the components needed for splicing
How is the B2 complex formed?
U1 and U4 are released from the B1 complex
How much energy is required to form the B2 complex?
The hydrolysis of 2 ATP molecules
What is the role of U4 snRNA before it is released?
U4 sequesters the U6 snRNA until it is needed.
What happens to U6 when U4 is released?
U6 pairs more extensively with U2 to create the active site
Which snRNPs bring the 5’ splice site into contact with the branch point?
U2 - already paired with the branch point
U6 - pairs with the intronic sequence downstream of the 5’ splice site
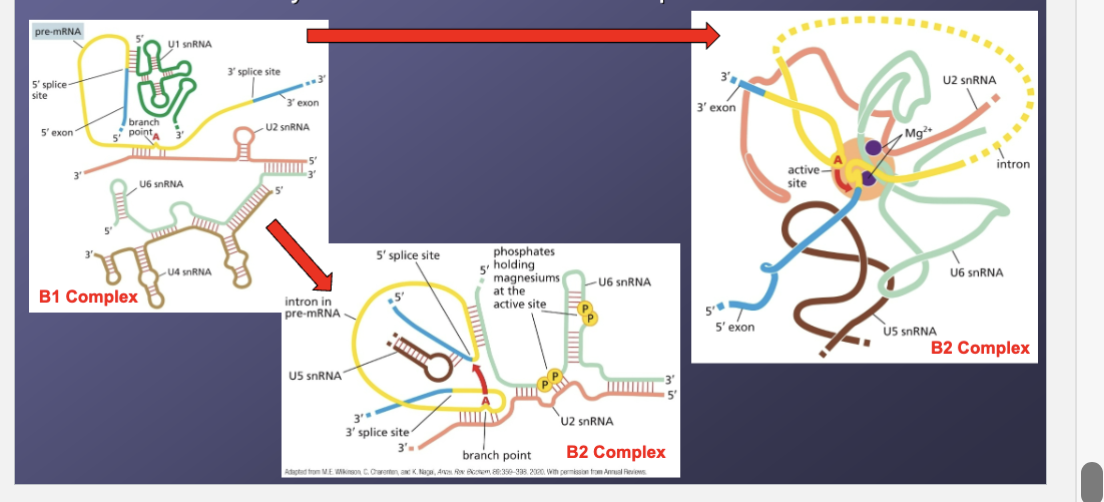
What does step 3 create?
Creates the entirety of the active site
Which snRNP assists by interacting with the upstream exon?
U5
How is C1 complex formed?
Several RNA rearrangements occur in the B2 complex
What occurs in C1 complex?
The first transesterification reaction between the 5’ splice site and branch point to form the lariat.
What is the role of the U5 snRNP in the C2 complex?
It positions U2 and U6 for the second transesterification reaction between the flanking errors
What happens to the snRNPs after the exons are joined?
They remain attached to the lariat, but are released as the lariat dissociates and is degraded
What is required per splicing event?
100+ proteins
5 snRNA molecules
Hydrolysis of 8 ATP molecules
Reassembly of the entire active site
Why might splicing be so “inefficient”?
It may be a consequence of the overexpansion of ancient self-splicing mechanisms. It is more complex than needed for regulation and accuracy
Why can’t the splicing process “backtrack”?
Because of the overall necessity of the splicing process for gene expansion
Which part of complexity is actually needed?
ATP hydrolysis
What is the primary use of ATP hydrolysis in the splicosome?
Used to break specific RNA-RNA base pairs so that others can be made that are specifically required for the sequential assembly of the spliceosome.
What happens if the initial correct base pairs dont form?
ATP hydrolysis will not occur and spliceosome assembly will not proceed.
Give two examples of base-pair exchange driven by ATP
Specific 5’ splice site pairing with U1 is broken by ATP hydrolysis and replaced by specific pairing with U6
Specific U6 pairing with U4 is broken by AP hydrolysis and replaced by specific pairing with U2
What is ATP hydrolysis also used for?
Kinetic proofreading
What is “kinetic proofreading” in splicing
ATP-mediated rearrangements are used to test the stability of base pairs; incorrect pairing will be less stable and are less likely to incorrectly proceed to the next stage of assembly than rearrangements that are correct. The correct base pairing is stronger than incorrect pairing, and the incorrect pairing will dissociate more quickly.
What is the exon-junction complex (EJC)?
A complex that is not correct. It deposited onto each exon-exon junction following splicing