BIOL 1101 FINAL
1/144
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
145 Terms
Opioid Receptors
Receptors on the surface of neurons that bind opioid drugs (ex. fentanyl, heroin, and naloxone) and is an integral membrane protein
Naloxone (Narcan)
An opioid receptor antagonist that binds the mu-opioid receptor without activating it, blocking fentanyl and heroin.
Antagonist
A molecule that binds to a receptor but does not activate it, blocking agonists from binding.
Heroin
An opioid agonist that activates the mu-opioid receptor but induces weaker and less dangerous signaling than fentanyl.
Fentanyl
A powerful opioid agonist that binds strongly to the mu-opioid receptor, causing intense signaling and high risk of respiratory depression.
Agonist
A molecule that binds to a receptor and activates it, turning on downstream signaling.
Mu-opioid receptor
A specific opioid receptor on neurons responsible for pain relief, respiratory depression, etc
Heroine, fentanyl, and naloxone bind to receptors on cells and cause different effects blc they…
Bind to the same receptors but cause different effects
How do fentanyl and heroin affect cells?
They act as ligands (molecules that bind to proteins) and bind to mu-opioid receptors on the outside of cells, this changes the receptor’s shape and triggers a signaling cascade inside the cell.
Fentanyl is so dangerous because it…
Particularly depresses respiration
A covalent bond
Shares electrons between two atoms
How many covalent bonds can a carbon atom make?
4
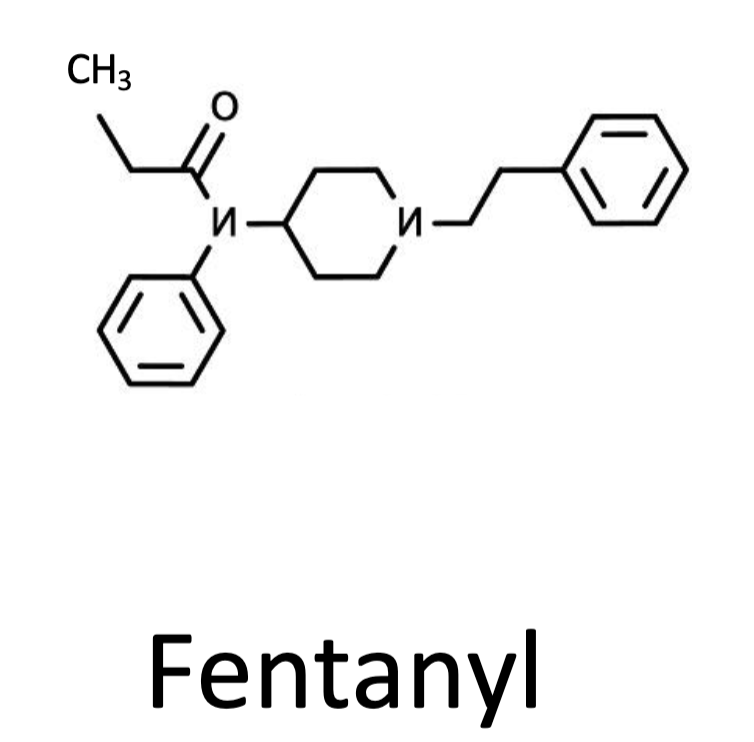
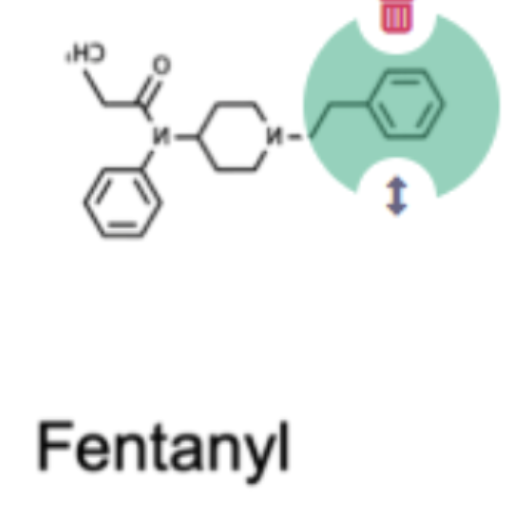
Pick the specific region on the fentanyl molecule that might explain why fentanyl has more potent negative effects than heroin
Non-polar ring that may be able to get inside of the pore of the protein more and interact with it there
Sickle Cell Anemia
A disorder caused by a single amino acid substitution (valine) in beta-hemoglobin that alters protein structure
Molecule
Multiple atoms held together by shared electrons
Polar
Uneven sharing of electrons (asymmetrical, ex. H2O)
Non-polar
Even sharing of electrons (symmetrical)
Property of forming hydrogen bonds in water
High cohesion, ability to exclude uncharged molecules, and ability to resist rapid temperature change
Covalent bond
Shared electrons between atoms ( electrons can be shared evenly/unevenly)
Structure of Sugars
Ring structures
Structure of Lipids
Long chains
Structures of peptides/proteins
Branching structures
Substances composed heavily of C and H (lacking other atoms)..
Crowd away from water (ex. oils, fats, lipids)
Hydrophobic molecules
“Water fearing”, want to interact w/themselves and shield themselves away from water
Why is water unique at ambient temperatures?
Resists rapid temperature change, dissolves many substances, excludes some molecules, and is cohesive (can be pulled/pushed in columns).
Hydrolysis
Uses water to break a bond (adds a water molecule)
Which parts of an amino acid participates in peptide bond formation?
Formed between the nitrogen of one amino group and the carboxyl group of another amino acid.
Macromolecules
Formed of repeating smaller (subunits), si
Macromolecules
Formed of repeating smaller units (subuntis) covalently bonded; four classes: proteins, nucleic acids, carbohydrates, and lipids
Polypeptides
Strings of amino acids residues connected by peptide bonds
Protein
Is a macromolecule formed on one or more polypeptides
Primary structure
The specific sequence of amino acids making up a polypeptide chain, from amino to carboxyl termini
Secondary structure
Either an alpha helix/beta pleated sheet (is generated by H-bonding in backbone of polypeptides/peptide)
SIDE CHAINS DO NOTHING
What drives protein shape in living organisms through polypeptide amino acid residue side chains?
Non-covalent bonds (ionic, H-bonding, van der Waals forces) and the hydrophobic effect
Denaturation
Break non-covalent bonds
Non-covalent bonds (electrostatic interactions)
Does not involve sharing of electrons, gives molecules structural flexibility: hydrogen bonding, ionic bonding, van der Waals interactions, hydrophobic interactions
T or F If water were less polar: increased cohesion would contribute to increased upward water transport potential in plants
FALSE: Cohesion: water molecules stick together because of hydrogen bonding and how transpiration happens where water goes up the plants (less polar = less sticky).
Why is “big” better for molecules?
Storage (energy, info)
Build/attach, construct bigger things
Bring things together, make stuff happen
Nucleotide
Monomer/building block of nucleic acids made up of a 5-carbon sugar, a phosphate group, & a nitrogenous base
Monosacchardie
A sugar (ex. glucose)
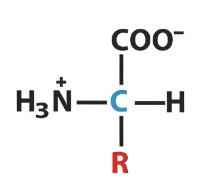
The -R group
An amino acid polymer
A peptide/polypeptide
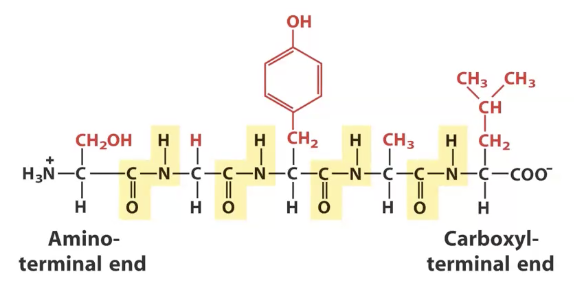
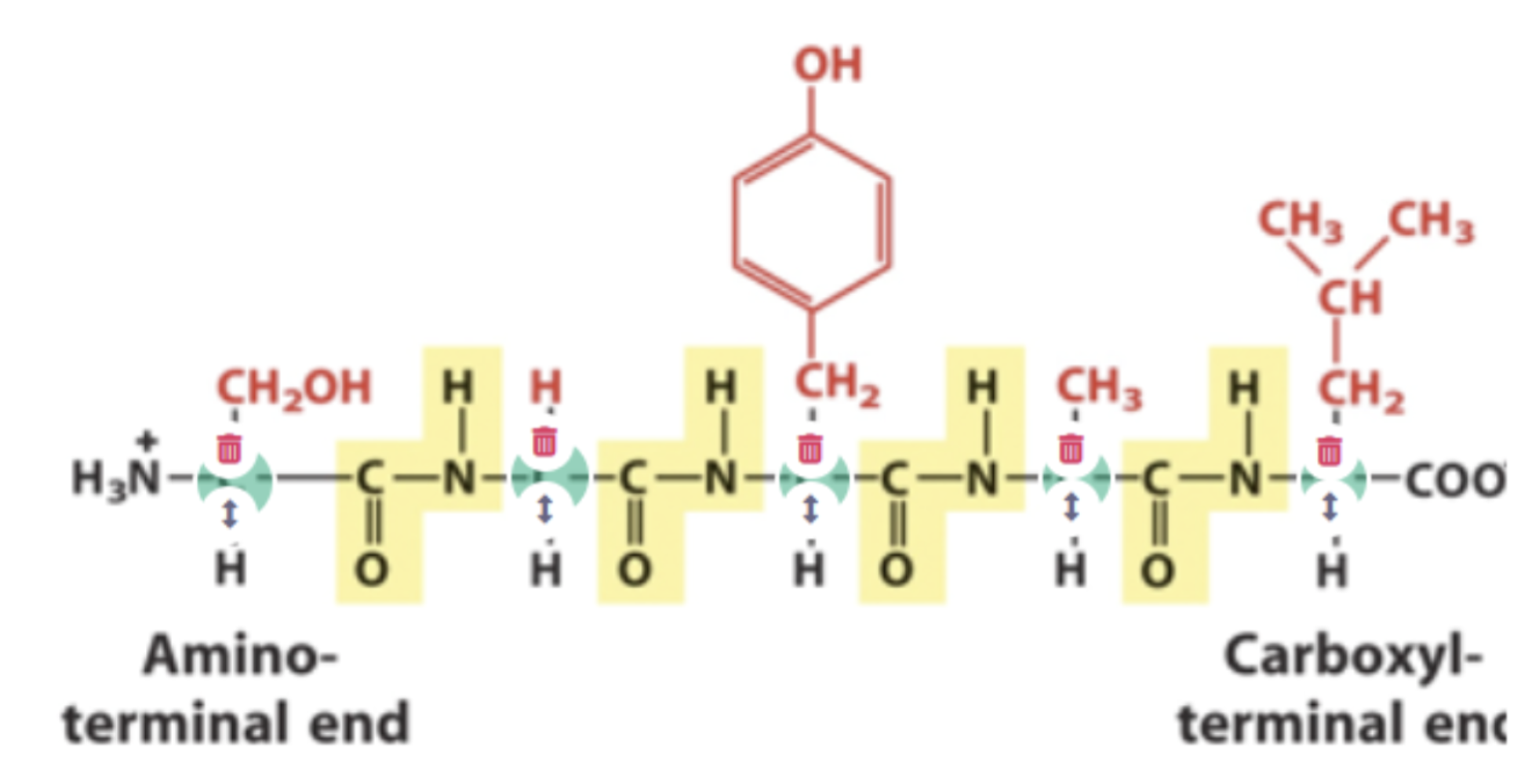
Polymer of amino acids, identify the alpha carbons and peptide bonds
Alpha carbons highlighted in blue, peptide bonds highlighted in yellow
Polymers
Made from covalently attaching monomers together in repeating reactions
Polymerization
Small molecules (monomers), chemically bond together to form large, chain-like molecules (polymers) - ex. dehydration (condensation)
Dehydration (condensation)
Removes a water molecule, forming a new bond
Depolymerization
The process of reacting monomers together to form 3-D networks of polymer chains (polymer gets shorter) - ex. hydrolysis
Tertiary structure
Whole polypetide will fold upon itself (one polypeptide)
Quaternary structure
Binding/folding of multiple polypeptides into a single protein
Outside cells are what type of bond?
Covalent bond (ex. Cysteine disulfide bonds)
Peptide bonds
Covalent bond between amino acids, the carboxyl group of one amino acids shares electrons with amino group of another amino acid. (generates potential for hydrogen bonding)
Which structures do the side-chain non-covalent bonds play an important role?
Teritarry & Quaternary structures
Hydrophilic
“Water-loving”, want to interact with water (polar/ionic molecules)
General protein structure follow 2 rules:
Hydrophobic residues are buried in protein interior, away from water.
The # of H-bonds within the protein is maximized.
What type of bond links amino acids together in primary structure?
Peptide bond
What type of bond stabilizes the local folding of a polypeptide into a beta sheet
Hydrogen bond
Where are non-polar R groups generally
In the interior of folded proteins
What type of bond stabilizes secondary structure?
Hydrogen bond
In sickle cell disease, beta-hemoglobin contains (blank) at amino acid position 6
Valine
CFTR protein
A receptor in the membrane, folded in ER and go to membrane of cell
If CTFR is misfolded what happens to the peptide?
Peptide never leaves ER, and is degraded by “quality control” enzymes
Phospholipids
Are amphiphilic, they have polar heads and non-polar tails. Composed of a glycerol backbone, a phosphate group, & 2 fatty acid chains
Glutamic Acid
Contains 2 oxygens, carries a full charge at physiological pH (anion), interacts favorably w/water
Valine
Composed of carbons and hydrogens, non-polar and hydrophobic, unfavorable interaction w/water
T or F: More double bonds in fatty acid tails increases fluidity
TRUE: . they introduce "kinks" in the hydrocarbon chains, which prevent them from packing tightly together.
T or F: Longer tail length of fatty acids increases fluidity
FALSE: longer fatty acid chain lengths decrease membrane fluidity and make the membrane more rigid since it increases surface area for intermolecular interactions.
How might a yeast cell adjust the contents of its cell membrane to protect itself from a sudden increase in temperature?
It might increase the # of long-chain fatty acids, number of sterols, and # of saturated fatty acids in its membrane
Saturated fatty acid
A fatty acid with no double bonds, making it straight and rigid.
Unsaturated fatty acid
A fatty acid with one or more cis double bonds, creating “kinks”.
Cholesterol (steroids)
A lipid, it can affect the fluidity of cell membrane
What is generally found in the bilayer of cell membrane
Phospholipids, sterols, integral proteins, glycolipids, and lipid rafts
Carbohydrates like those on glycolipids and glycoproteins are found
On the exterior of the cell membrane
Cell signaling pathway
A series of liked molecular interactions and reactions
Heterotrimeric G proteins
Can operate as molecular signaling switches and have alpha, beta, and gamma subunits
Hydrophobic signaling molecules
Can pass right through the membrane rather than use a cell-surface receptor
How to activate/promote cell cycle
Cyclin
Apoptosis
Programed cell death
Endocrine signaling
Long-distance cell-to-cell communication where specialized endocrine cells or glands secrete hormones into the bloodstream.
Paracrine signaling
Local cellular communication where a cell produces signaling molecules (ligands) that induce changes in nearby target cells within the same tissue
Autocrine singaling
Signal that is sent and received by the same or similar nearby cells.
“Juxtacrine” signaling
Occurs when signaling molecules on the surface of one cell interact with receptors on an adjacent cell.
Synaptic cleft
Space between neurons
Ligands
The signaling molecules that bind receptors
Examples of receptors that interact with cell surface
Adrenaline signaling, growth factor signaling
Examples of receptors that enter cell to interact with intracellular receptor
Steroid signaling (estrogen, cortisone)
Liver physiological response
Breakdown of glycogen and release of glucose into bloodstream
Adrenaline (epinephrine)
Signaling ligand that is a small polar molecule and signals through endocrine signaling responding cells in the liver
Adrenergic receptors
GPCrs found on cell surface that bind epinephrine/norepinephrine and trigger a signaling response.
GPCRs
G-protein coupled receptors. GPCRs work by producing lots of secondary messengers
Protein Kinase A (PKA)
Enzyme activates enzymes that release glucose from glycogen stores
Consequence of phosphorylation of proteins
Changes rate of heart muscle contraction/liver cell release glucose
During transmission of signals inside cells..
The signal is amplified and spread
Signal Amplification
A process where one ligand binding to a receptor triggers a cascade that activates many downstream enzymes/second messengers
How are signals turned off and “reset”?
Often through protein dephosphorylation (phosphatases) = removal of phosphate group
Signaling pathways must be turned (blank)
OFF to ensure proper cellular regulation, homeostasis, and recovery for future signals
G Proteins
Breaking a phosphodiester bond in GTP turns it into GDP (no protein kinase/phosphatase here)
What are the steps of the adrenaline GPCR signaling pathway?
Adrenaline binds GPCR → G protein activated (GTP binds) → adenylyl cyclase activated → cAMP produced → PKA activated → proteins phosphorylated → cellular response
Why is amplification important in adrenaline signaling?
Allows a small amount of concentration to produce a large response
What is the structure and function of GPCRs?
Seven transmembrane domains (crosses over membrane), the ligands bind on the outside, causing a conformational change inside the cell to transmit a signal.
Activating receptors leads to…
Conformational change