Genetics Module 4
1/97
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
98 Terms
directional selection
additive effects
all become 1of the homozygous extremes
disruptive selection
underdominance
advantage for both extremes
leads to bimodal population
Stabilizing selection
Overdominance
All become heterozygous
Both alleles remain in population
effects of small population size
genetic drift
founder population
inbreeding
genetic drift
founder population
founders will affect next generations
genetic drift
random fixation or loss of alleles over time from small population
positive assortative mating
mating similar individuals results in more homozygotes but only for loci under selection
negative assortative mating
mating opposite individuals results in more heterozygotes for the loci under selection
inbreeding
changes frequency of genotypes but not allele frequency
more homozygotes
affects all loci
cancer genetics
genetic but rarely heritable
cancer multi hit
Sporadic and influenced by environment
-siblings rarely have same cancer
-populations that migrate to new regions get cancer rates typical of that region
Cancer develops over time
-changes in cancer rates from new environment take decades
types of genes involved w/ cancer
tumor suppressor genes (prevent bad cells from dividing)
proto-onco genes (allow good cells to divide)
tumor suppressor gene mutation
recessive acting- must disrupt both copies to lose cell regulation
tumor suppressor genes
BRCA1
p53
RB
RB gene mutation
Tumor suppressor gene
40% cases inherited- inherit 1 bad type then another somatic mutation happens to cause cancer
RB gene function
G1 to S transition
normally prevents E2F from activating replication
BRCA1 and 2
tumor suppressor
used to repair double strand breaks
woman vulnerable to breast and ovarian
men vulnreable to prostate and breast
p53
Tumor suppressor
Colon, lung, breast, brain and is found in altered form in half of all tumors
if DNA damaged, p53 delays cell division until DNA repaired or dies
haploinsufficiency exception
loss of function of one copy and the other copy doesnt produce enough gene product to exhibit wild type phenotype
some cases of cancer w/ mutations in only 1 copy of tumor suppressor gene: bloom syndrome
bloom syndrome
defective DNA helicase enzyme that repairs double strand breaks
homozygous for mutated BLM gene have very high rates of cancer
hetero ppl have elevated colorectal cancer risk
oncogene mutation
only need 1 copy for cancer
burkitts lymphoma
reciprocal translocation between chromosomes 8 and 14 places c-myc ongogene next to enhancer
chronic myelogenous leukemia
reciprocal translocation involving chromosomes 9 and 22 places 2 oncogenes near each other
retroviruses causing cancer
mutate and rearrange proto-oncogenes
insert a strong promoter near proto-oncogenes
clonal evolution
over time tumor cells acquire more mutations that allow them to be progressively more aggressive in proliferation
mutations affect
cell cycle regulation
signal transduction
telomere length
chromosome segregation
vascularization
DNA repair
defective nucleotide excision repair= xeroderma pigmentosum
defective mismatch repair = colorectal, endometrial, stomach cancers
defective double strand break repair = BRCA1, 2
telomere length
telomerase works in germline cells but not in somatic cells, allows them to die and be replaced
tumor cells have telomerase, thought to contribute to the immortality
vascularization
angiogenesis- growth of new blood vessels important to tumor growth
angiogenesis inhibitors may be under expressed or inactivated
growth factors for angiogenesis overexpressed
classic cancer modle
cancer is a proliferative disease
cancer prevention
HPV vaccine
avoid environmental factors
get cancer screenings
height
quantitative trait with normal distribution
multi factor hypothesis
Expression depends on the additive effects of a number of genes.
The effect of each gene is small.
Environment plays an important role in expression of trait.
-smooths curve
phenotype formula
P = Genetics + Environment
Genotype formula
G = Additive + Dominance effects
A variable
A = Average effect of substituting A for a in genotype
D variable
D = dominance effects due to specific combinations of alleles at a locus
when does D = 0
if the value of Aa is exactly between the values of AA and aa
how to determine the values for G, E, A, D
We must look at a population and determine the values for G, E, A, D. and partition the variation we see in the phenotypes of the individuals in the population into G, E, A, D.
Heritability
proportion of the phenotypic variance due to genetic effects
broad sense and narrow sense
Broad sense heritability formula
H² = VG/VP proportion of phenotypic variance due to genetics
narrow sense heritability formula
h² = VA/VP proportion of phenotypic variance due to additive genetic effects
The higher the heritability…
the more progress we can expect from selection/breeding
heritability ranges
0 = all VP from environmental variation
1 = all VP due to genetic variation
who developed PCR
Kary Mullis
temp to heat up to PCR
95 c to allow strands to separate
then cool to 50-65 to identify target region for amplification
PCR steps
denature DNA by heating and allows strands to separate
cool, primers anneal to identify target region
Taq polymerase adds nucleotides to 3’ end of primer (72c)
PCR limitations
must know about sequence surrounding gene of interest
taq polymerase does not proofread and correct errors
fragments amplified are small
amplification of DNA
VNTR
allele based on length of DNA segment
must be polymorphic
recomb DNA restriction enzyme
endonuclease that recognizes a specific DNA sequence and cleaves dsNDA at that sequence
recomb DNA palindrome
reads the same 5’ to 3’ on either strand for a segment of DNA
cloning vectors have
origin of replication
selectable/insertional markers
multiple cloning site
cloning vectors selectable/insertional markers
allow cells containing vector and recomb molecule to be identified
cloning vectors multiple cloning site
has many restriction enzyme cut sites that can be used in producing a recomb DNA molecule
ligation experiment for recomb
join foreign DNA to vector
foreign DNA and vector cut w/ same restriction enzymes
DNA mixed and DNA ligase is added
insertional inactivation for recomb
inserted DNA inactivates a gene in the vector by inserting into that gene
allows cells which contain recomb DNA molecule to be identified
transformation experiment for recomb
allow cells to take up products from a ligation experiment
identification of different cell types for recomb
cells w/ no uptake, cells that took up the original vector, and cells that took up the recomb plasmid
Why Does Variation Exist in a Population?
Neutralist Theory and Selection Theory
Reproductive isolation can occur because
They don’t choose to mate with each other or cannot mate with each other (Prezygotic).
Or their progeny are sterile or inviable (Postzygotic)
prezygotic reproductive isolation
They don’t choose to mate with each other or cannot mate with each other
Postzygotic reproductive isolation
Their progeny are sterile or inviable
prezygotic types of reproductive isolating mechanisms
Ecological- habitat differences
Behavioral- different mating behavior
Temporal- reproduction at different times
Mechanical- anatomical differences
Gametic- not compatible
postzygotic hybrid breakdown
F1 hybrids are viable and sterile but F2 are not
Allopatric Speciation
Geographic barrier initiates speciation by blocking gene flow
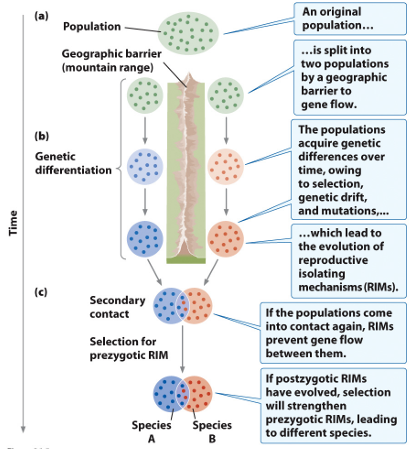
Sympatric Speciation
Arises within a single interbreeding population without geographical barriers to gene flow.
Hybridization that leads to allopolyploidy is another mechanism for sympatric speciation.
Races of the Apple Maggot Fly sympatric speciation example
Resource use is linked to mating preference.
Original fly fed on hawthorn tree fruit.
Mutation allowed feeding on apples.
Those with the mutation mated together more on apple trees (reproductive isolation)
Speciation not complete yet.
Gene flow only 2% between hawthorn and apple flies
Anagenesis
evolution within a lineage over time
Cladogenesis
splitting of one lineage into two
Paralogs
homologous sequences found in the same species and arrive through gene duplication.
Orthologs
homologous sequences found in different species
Molecular Clock
differences in sequence between present day organisms can be used to date past evolutionary events.
G1/S checkpoint
monitors for proper cell size and undamaged DNA.
G2/M checkpoint
holds up cycle until replication and DNA repair are complete.
M checkpoint
proper spindle formation and attachment
High MZ low DZ
indicates significant role of genetic effects
Low MZ but still much higher than DZ
indicates genetic predisposition, but environmental factors are important
how to make cDNA from mRNA
use reverse transcriptase (RNA dependent DNA polymerase)
-uses mRNA as a template to synthesize the first strand of cDNA
transformation of rice w/ daffodil PSY gene
need lots of DNA
need a promoter
need a way to get it into the rice genome
Rhizobium radiobacter
transfers DNA to plants
Modified Ti plasmid inserts cloned gene of interest into plant chromosome
Expression vector
allows inserted gene product to be produced
must contain sequences required for transcription and translation of the gene
what is needed for insertion into rice genome
PSY gene
crtl gene
promoters- endosperm specific
Poly A signals
Transgenic marker
LB and RB insertion into plant genome
blotting
process of transferring molecules that were previously separated to a membrane better able to support additional testing
Southern blot measures what
DNA fragments separated based on length
how does southern blot work
single stranded DNA fragments from a gel are transferred to a nylon membrane
Nylon membrane incubated w/ labeled single stranded probe DNA of interest
Probe binds to complimentary DNA fragments on the nylon
Northern blot
RNA fragments separated based on length
Western blot
Proteins separated based on molecular weight, isoelectric point, electric charge, etc
electrophoresis
DNA loaded at negative pole and migrates to positive pole
small migrates faster
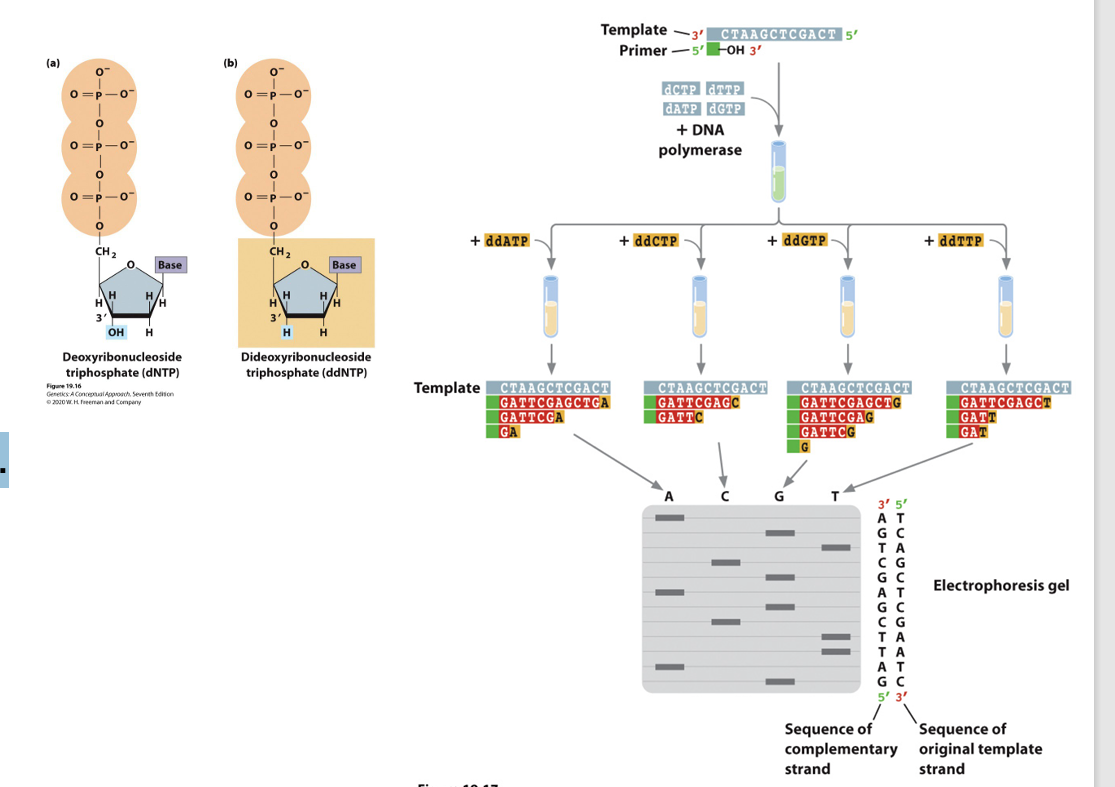
Dideoxy Sequencing Reaction
3’-OH required by DNA polymerase to form a phosphodiester bond.
• DNA replication reaction proceeds until a dideoxy nucleotide is incorporated.
• No further extension occurs.
• Detect which position the dideoxy nucleotide was incorporated bc that’s where the sequence terminates
golden rice results
rice with maize PSY genes makes higher b carotene than rice with daffodil gene
need 72g of that a day to get enough vitamin A
Forward Genetics
Start with a mutant phenotype and seek out the gene that causes that phenotype.
• Use chromosome mapping to identify the gene
Reverse Genetics
Start with a DNA sequence (a genotype), alter its function or prevent its expression and observe the effects on the phenotype
Reverse Genetics Transgenic Mice
Inject gene of interest into fertilized egg.
Implant in female.
Test progeny for presence of gene.
Mate to obtain mice homozygous for gene.
Study gene function
Reverse genetics knockout mice
Phenotype of knockout mice reveals function of gene
Reverse Genetics Knockdown Expression Using RNAi
Excess ApoB protein leads to high levels of cholesterol.
Inject lipid coated ApoB synthetic siRNA into Cynomolgus monkeys to ‘knockdown’ expression.
Monkeys that got more siRNA had lower cholesterol
Microarray analysis of RNA from cancer and noncancer cells
shows that some genes are more expressed in cancer cells (green) and some more expressed in noncancer cells (red)
RNA sequencing to determine expression of genes
RNA of interest isolated
enzyme reverse transcriptase used to make cDNA from mRNA
cDNA broken into overlapping fragments
adapters w/ sequences for amplification and sequencing are added to ends of fragments
fragments are amplified w/ PCR