Unit Questions
1/42
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
43 Terms
How do you define the genome of a prokaryote?
single circular dna molecule in nucleoid region
can include plasmids
In terms of the genome, how do archaea differ from bacteria, how are they the same?
both
prokaryotes and no nucleus
one circular chromosome
dna in nucleoid region
can have plasmids
differ
gene structure and machinery: archaea gene and expression similar to eukaryotes
dna packaging: archaea use histone like proteins to organize dna similar to eukaryotes
rna polymerase: archaea have complex rna polymerase similar to eukaryotes; bacteria are more similar
How are genes organized in bacteria? Operon
group of genes controlled together
transcribed into single mrna
structural genes - code for proteins and carry out cell functions
promoter (start of transcription) - binding site for rna polymerase and located upstream
operator (controls access or transcripton) - binding site for repressor protein
one switch controls multiple genes
How are genes organized in bacteria? Structural and control genes
structural
code for proteins (enzymes) that do work in cell
transcribed into mrna then translated into proteins
worker
Control (regulatory)
produce proteins (repressors/activators) that turn genes off/on
regulate gene activity
manager
How are genes organized in bacteria? Monocistronic and polycistronic mrna
monocistronic mrna
one gene → one protein
common in eukaryotes
Polycystronic mrna
multiple genes → multiple proteins
common in bacteria and operons
How are genes organized in bacteria? Regulon
group of genes or operons controlled by same regulatory protein
multiple operons controlled by one regulator
What are plasmids and what are some of their features?
circular double stranded dna molecules found in bacteria
own origin of replication (ori)
replicate independently
not essential for bacterial survival but beneficial
r plasmids - antibiotic resistance genes
virulence factors - toxin production genes
metabolic genes - break down substances
moves via conjugation (cell to cell) to spread traits
How are plasmids maintained in a cell?
ori
allows plasmids to copy themselves independently of bacterial chromosome
ensures plasmid dna is available for daughter cells
regulate copies
prevent over replication and maintains stability
partitioning
par genes to ensure plasmids distributed to daughter cells during cell division
post par killing/ toxin antitoxin systems
plasmids have toxin and antitoxin gene
if plasmid is lost, antitoxin disappears and toxin damages or kills cell
selective advantage
antibiotic resistance - plasmid cells survive antibiotics
How do low-copy-number plasmids and high-copy-number plasmids affect the cell and plasmid stability/expression?
low copy
few copies (1-10)
tight control systems and partitioning genes
lower metabolic burden
more stable
low gene expression
high copy
many copies (50-200+)
rely on replication control
higher protein expression
higher metabolic burden
slows growth of cell and adds stress
unstable
What is rolling circle replication?
dna replication used by some plasmids bacteriophages and viruses to rapidly produce copies of circular dna
single strand nick - enzyme cuts strand of circular dna
3’ is starting point - dna polymerase adds nucleotides to free 3’ OH end
strand displacement - new dna synthesized and old strand is unrolled
synthesis - circular template rolls, creating long single stranded dna tail
second strand synthesis - displaced single strand converted to double stranded dna and created multiple copies of original circular dna
What is transcription and what is the end result?
Copying genetic info from dna to rna
carried out by rna polymerase and uses dna strand as template
rna polymerase binds to promoter and reads dna, building rna
What is translation and what is the end result?
making protein from mrna template
occurs at ribosome
translates mrna genetic code into amino acid sequence
mrna binds to ribosome
ribosome reads mrna in codons (3 bases)
trna bring matching acids
amino acids link to make polypeptide chain
folds into protein
What are -35 & -10 sequences, i.e., what is their significance?
dna regions in promoter of genes that help initiate transcription
nucleotides upstream of transcription start site
rna polymerase
recognized by sigma factor
bind to promoter
-35 : recognition site
-10 : opening site (dna unwinding)
What component of RNA polymerase interacts with 010 and -35 sequences?
sigma factor
recognizes and binds to promoter regions
helps rna polymerase
locate promoter
bind to dna
initiate transcription
What is meant by promoter strength?
how effectively a promoter initiates transcription
strong - rna polymerase binds frequently → high transcription → more mrna
weak - rna polymerase binds less → lowtranscription → less mrna
How is basal level gene expression increased to high level expression?
basal - low transcription
positive regulation/activation increases expression
activator protein binds near promoter so rna polymerase binds more easily
What is the function of the Shine-Delgarno sequence? Where is it found?
ribosome binding site on mrna
aligns ribosome with start codon
What is meant by vertical gene transmission and horizontal gene transmission? What are examples of how each of these processes occurs in prokaryotic cells?
vertical
normal cell division - low genetic variation
transfer genetic info from parent cell to daughter cell
binary fission - dna replicated and split into identical cells
horizontal
transfer of genetic material between cells not parent and offspring - high genetic variation
can occur between diff bacteria or species
conjugation - direct transfer via sex pilus (plasmids)
transformation - uptake of free dna from environment (comes from dead/lysed cells)
transduction - bacteriophages (infectious viruses)
What is the difference between core and flexible gene pools?
core
genes in all strains of species located on main chromosome
encode housekeeping functions for survival and reproduction
stable pool and changes slowly (vertical)
flexible
genes present in some strains found in mobile elements
specialized functions for special niches/stress
dynamic and updated through horizontal
What are genomic islands?
large segments of dna in genome acquired from other organism via HGT
provide upgrades for bacteria to adapt to environments
Describe the horizontal gene transfer mechanisms and components involved: Transformation; artificial v. natural transformation; Gram-negative cell vs. Gram-positive cell
transformation
bacterium in competence state
tightly regulated and triggered by stress or high cell density (quorum sensing)
competence factors
signaling molecules that trigger expression of transformation genes
transformasome
protein complex in cell membrane capturing extracellular dna and facilitates entry
translocasomes
machinery that pulls dna strand into cytoplasm
gram positive
uses transformasome
non selective
gram negative
move dna across 2 membranes (type iv pilus)
highly selective
natural
cell’s own encoded machinery (transformasome)
spontaneously in wild
Genetic diversity, DNA repair, or DNA as a nutrient source
artificial
damage membrane to create temporary pores
chemical - treated with CaCl2 followed by heat shock, neutralizing negative charge and creating pressure gradient
electroporation - hit with high voltage electric pulse, destabilizing membrane and creating pores for dna entry
Describe the horizontal gene transfer mechanisms and components involved: Conjugation (include role/function of: sex pilus, fertility factor, Hfr formation, F prime factor)
f factor
specialized dna containing genes for conjugation
f+ are donors that have f factor
f- are recipients lacking f factor
codes protein needed to build conjugation bridge and enzymes to initiate dna transfer
sex pilus
protein produced by donor cell
pilus attaches to receptor on recipient cell and pulls cells together forming conjugation bridge
Describe the horizontal gene transfer mechanisms and components involved: Transduction. What other process of horizontal gene transfer does specialized transduction resemble and why?
transduction
virus that infects bacteria carries dna from one cell to another
generalized
lytic cycle - accidental packaging
when phage is chopping host dna for its own, it puts bacterial dna instead of viral dna
random gene
specialized
lysogenic cycle where virus hides inside dna and waits
viral dna integrates on bacterial chromosome and removes itself imprecisely, taking bacterial genes with it
specific genes near viral site
Describe the horizontal gene transfer mechanisms and components involved: Transposition: what is this and how does this operate? What is conjugative transposition? Compare a simple transposon to a complex one. What are insertion sequences?
moving dna within cell - chromosome → diff spot
jumping genes because dna sequences jump around
insertion sequences
carries bare minimum tools to move
transposase genes - scissor enzyme cuts dna and puts it in new spot
inverted repeats - identical sequences at both ends as markers for transposase
simple transposons
carry cargo genes between inverted repeats
complex transposons
2 separate insertion sequences to move everything in the middle
replicative
copy paste - transposon replicated, one stays, one moves
non replicative
cut past - transposon cut and moved
conjugative
can jump out of chromosome of one cell, use conjugation, and jump into new chromosome in new cell
Describe the horizontal gene transfer mechanisms and components involved: What is a partial diploid cell?
carries normal set of genes plus second copy of specific genes
f’ conjugation
f factor takes bacterial genes when exiting chromosome and new cell has original genes on chromosome plus second set of genes on new f’ plasmid (plasmid/sex pilus)
specialized transduction
virus carries host genes to new cell and adds on to existing copies (virus)
What is the role of regulatory proteins in a bacterial cell?
control when and how genes are expressed
bind to dna regions (promoters/operators)
activate expression → rna polym start transcription
repress expression → block rna polymerase
Describe the levels at which control of gene expression can occur in a bacterium
transcription control
controls if mrna is made from dna
regulatory proteins turn genes on/off
involves operons
post transcription control
mrna stability and availability
rna processing and modification
translational control
mrna translation to protein
post translational control
protein activation/inactivation
protein folding, modification, and degradation
What are constitutive genes?
genes continuously expressed
not regulated or switched off
code for essential proteins
metabolism
cell structure
dna replication
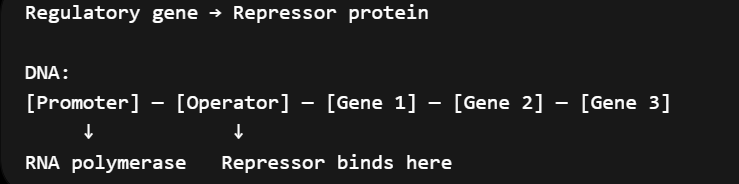
Describe the components of the operon model of gene organization and regulatory elements.
promoter
DNA sequence where RNA polymerase binds and starts transcription
operator
controls transcription
regulatory proteins/repressors bind here
structural genes
code for proteins
regulatory gene
codes for regulatory protein
repressor
binds to operator, blocks rna polymerase, and turns genes off
activator
helps polymerase bind to promoter, increasing transcription
inducer
inactivates repressor
corepressor
activates repressor
What are inducible and repressible operons?
inducible
turned on when inducer is present
repressor active by default
inducer then binds to repressor
repressible
turned off when corepressor present
inactive by default
corepressor binds to repressor and activates it, stopping transcription
What is a corepressor?
turns off gene expression by activating repressor protein
What is an inducer?
turns ON gene expression by inactivating a repressor protein
What is derepression?
process of turning gene expression back ON by removing repression
Describe what the lactose operon is, and how the lactose operon functions in both the presence and absence of lactose.
inducible operon controlling genes needed to break down lactose
lactose (allolactose) acts as inducer and binds to repressor, inactivating it
How does cAMP-CRP complex affect expression of the lac operon? How does the presence of glucose affect the expression of the lac genes?
cAMP - cyclic AMP
CRP - catabolite activator protein
cells make more cAMP when glucose is low to bind to crp
binds near lac promoter
helps rna polymerase bind
increases transcription of lac genes
What is diauxic growth?
pattern of bacterial growth where cells grow in two distinct phases when two different sugars are available
first growth
bacteria prefer glucose and growth is rapid
lag phase
glucose runs out and bacteria stop growing
adjust gene expression (activate lac operon)
second growth
bacteria use lactose and activate lac operon
growth resumes but slower than glucose
catabolite repression
glucose inhibits cAMP-CRP and turns off lactose genes
Describe what the tryptophan operon is, and how the tryptophan operon functions in both the presence and absence of tryptophan.
repressible operon in bacteria that controls the genes needed to synthesize the amino acid tryptophan
promoter
operator
structural genes
regulatory genes
absence - cell makes own tryptophan
repressor protein inactive and cannot bind to operator
polymerase transcribes genes
enzymes produced to synthesize tryptophan
presence - cell stops making tryptophan if available
tryptophan is corepressor
polymerase is blocked, blocking transcription, ceasing tryptophan
What is transcriptional attenuation? How does this work in the presence/absence of tryptophan?
gene regulation mechanism in bacteria that controls transcription
presence
ribosomes move quickly and forms terminator hairpin
polymerase stops transcription early
absence
ribosomes stalls at codons in leader sequence
anti terminator structure forms and polymerase continues transcription
What is phase variation? How does this work in the expression of Salmonella enterica flagellar protein?
expression of certain genes is reversibly switched ON and OFF in a random or controlled way
bacteria change surface structures to avoid immune detection and adapt to environments
What are small regulatory RNA (sRNA)? How do these function in regulation?
short RNA molecules that do not code for proteins but regulate gene expression at post transcriptional level
bind to target mrna molecules
can repress or activate translation
What are anti-sense RNA’s? How do these function in regulating gene expression?
single stranded rna compllementary to mrna in bacteria
can bind to mrna and affect expression
regulate genes by binding to mrna and blocking function
block translation
promote mrna degradation
What is the stringent response? How does this occur?
stress response when cell is nutrient limited
helps survive harsh conditions by slowing growth
triggered by lack of amino acids, stress, or uncharged trna accumulate
activation of relA
senses uncharged trnas at ribosome
synthesizes alarmone (ppGpp) and binds to polymerase
How do sigma factors figure into gene regulation?
proteins that help polymerase recognize and bind to promoters for transcription
control which genes are transcribed
bind to core polymerase
direct to promoter sequences