DNA Discovery/Replication + The Central Dogma + Tanscription/Translation/Mutation
1/56
Earn XP
Description and Tags
Probably don't need to know all the names of scientists but adding them just in case.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
57 Terms
What’s an Assay?
A way of measuring something (either a substance or an abstract phenomenon)
Friedrich Miescher - what did he discover/figure out?
Figured out a way to isolate the nuclei from cells found in pus and found that the main constituent of the nucleolus was a compound he called nuclein (which later turned out to be DNA)
Fredrick Griffith Experiment
2 bug strains: S (killed mice) & R (didn’t kill mice)
Inject heat killed S along with live R - killed mice because R had been transformed into S - substance responsible for transformation = transforming principle
What is the bacteria lifecycle?
Infection (of something)
Injection (of DNA)
Replication (of DNA)
Expulsion (to go infect more things)
Hershey and Chase experiment
Took a virus made of just DNA and Protein → labeled virus with EITHER the
DNA with 32P (radioactive phosphorus)
Protein with 35S (radioactive sulfer)
Allowed virus to infect bacteria and then threw the mix in a blender (taking off protein coat) → whatever stayed had to be the genetic material)
Infected bacteria produced more virus with 32P but basically no 35S
SO it had to be the DNA
What are Chargaff’s Rules?
# of A = # of T AND # of G = # of C
Even though ratio of A+T to G+C can vary widely from one organism to the next
Franklin and Wilkins used what technique to show what about DNA?
Used X-ray diffraction to show that DNA was double stranded and helical
Using Chargaff’s Rules and the findings of Franklin and Wilkins, Watson and Crick came up with what model?
Made DNA model where the n-bases are in middle and phosphate backbone runs along outside (backbone is negatively charged + exposed to water)
Pairs are always a purine and a pyrimidine → most stable bonds & distance btwn 2 strands stays constant
A from one strand always pairs with the T from the other strand
C from one strand always pairs with the G from the other strand
Strands are reverse complements of each other
Structure of DNA suggests what 2 things?
Anti-parallel (run in the opposite direction); reverse-complements (one strand complements the other - not mirrored)
sequence of nucleotides doesn’t affect overall structure → info can be encoded arbitrarily
the 2 strands bind by complementary base pairing so the they both contain identical information → separating the strands and binding them to 2 new strands = DNA replication
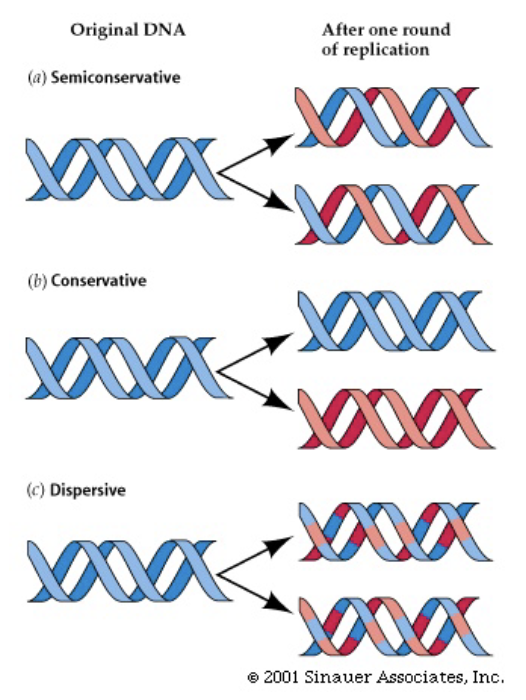
3 possible ways that DNA gets replicated + which is the correct way
Conservative replication = each strand makes a new strand and the 2 old strands re-bind to each other
Semi-conservative replication = each old strand made remains bound to the new strand (THIS IS WHAT ACTUALLY HAPPENS)
Dispersive replication = DNA breaks apart and rejoins to produce 4 strands, each with a mix of old and new DNA
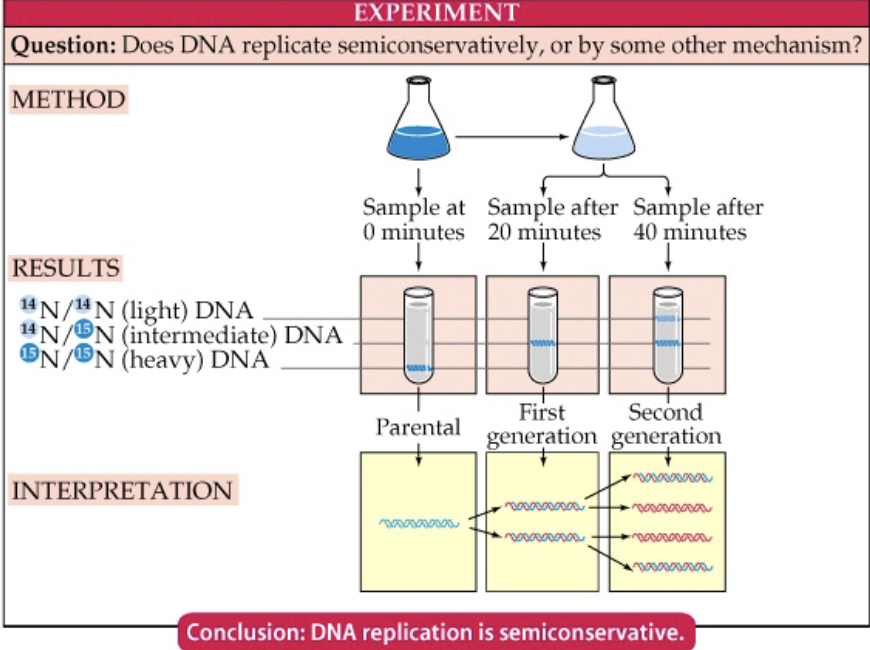
Meselson and Stahl experiment
Showed that DNA replication was semiconservative
labeled DNA by growing bacteria with a heavier isotope of nitrogen (15N) → measured weight with centrifuge
After ! duplication in the presence of normal nitrogen the duplicated DNA was ½ as heavy
After 2 duplications half the DNA was ½ as heavy and ½ the DNA was normal (light) weight.
Only semiconservative replication would produce this result
How does DNA polymerase work
Grabs nucleotide complementary to next nucleotide in line on template strand
Takes nucleotide and breaks bond btwn alpha phosphate (attached to the sugar) and the beta phosphate
Attaches alpha phosphate to the 3’ hydroxyl group of the last nucleotide on the strand thats being extended
(DNA synthesis always occurs 5’ to 3’)
What do the enzymes Helicase and Primase do?
Helicase expands energy to unwind the strands of DNA (making replication forks on either end where the old double stranded DNA is being split to act as a template for new DNA strands)
Primates adds short stretch of complementary RNA (a primer) to create a ragged end for the DNA polymerase (3) to work)
What are origins of replication?
The specific (non-random) spots where DNA polymerase 3 initiates replication
Bacteria have 1; Eukaryotes have many along the length of the chromosome
Leading vs Lagging strand
*Okazaki Fragments
Leading strand → continuous replication
Lagging strand → replicates in short stretches (bc it’s going backwards)
Continually puts down primers every few hundred bases
the short stretches of DNA = Okazaki Fragments
DNA Polymerase 3 vs DNA Polymerase 1
P3: DNA Synthesis (cloning)
P1: chews up RNA primers + fills in gaps within the lagging strand using newly synthesized DNA as a primer
What does DNA ligase do?
Joins the ends of newly synthesized strands → DNA replication then finished
What is the central dogma?
Crick, Brenner, & Jacob Proposed that the flow of information within genes is from DNA to RNA to proteins and is unidirectional
Garrod, who studied patients with alkaptonuria, proposed that Alkaptouria…
is/is not hereditary
is/is not autosomal recessive
“inborn error of __” caused by what?
Alkaptouria is hereditary and is autosomal recessive
“inborn error of metabolism” caused by the absence of an enzyme that metabolizes tyrosine
How does the idea of “inborn errors of metabolism” help you find mutations
Gene codes for enzyme SO mutant/absent gene = no/dysfunctional enzyme (which you can see as a phenotype probably)
Beadle and Tatum experiment with Neurospora (mold):
Hypothesis = there should be a mutant allele corresponding to every enzyme in a pathway.
Gene B codes for Enzyme B which turns ornithine into citrulline
Gene C codes for Enzyme C which turns citrulline into arginine
If strain has no Gene B, pathway can’t continue BUT if you add citrulline downstream of missing Enzyme B, then Enzyme C can still turn it into arginine
Auxotrophs vs Prototrophs
Both are microorganisms
Prototrophs make their own nutrients
Auxotrophs are have a mutation causing them to need external/added nutrience to survive
Information flow from gene to protein
DNA (in nucleus) → RNA (nucleus) = Transcription
RNA(nucleus → ribosome)
RNA(ribosome)→ Protein = Translation
Transcription: What step in information flow?
What enzyme? how does it work?
Step 1: DNA → complementary RNA
RNA polymerase binds to DNA promoter sequences and unwinds helics
Then reads DNA template in 3’ to 5’ direction → produces RNA transcript by adding nucleotides to the 3’ end (replaces T with U)
When RNA polymerase reaches termination site, RNA transcript (mRNA) is set free from the template
Newly synthesized mRNA can take parent DNA info from nucleus and bring to ribosomes (in cytoplasm)
Transcription: sense strand vs template/antisense strand
Sense Strand = not transcribed, has the same sequence as the mRNA
Template/antisense strand = transcribed, has sequence complementary to mRNA
A sequence of [#] DNA letters, called a __ corresponds to 1 amino acid
3 DNA letters = a codon
Start vs stop codons
Start: tells ribosome where the proteins starts (AUG)
Stop: tells the ribosome that the protein is done (UAA, UGA, UAG)
What does it mean when code is degenerate?
When several different codons code for the same amino acid.
What’s the issue with adding mRNA to a ribosome? what is the fix?
Ribosomes can’t rely on the natural affinity of a codon to an amino acid (unlike how polymerase relies on natural affinity of base pairs to each other) → need an adapter → tRNA acts as adapter
Explain the shape/structure of tRNA
Clover-like shape - loop has 3 nucleotides that stick out, called the anticodon → anticodon binds to a codon in mRNA according to base-pairing rules
The other end of tRNA molecule = the amino acid that’s encoded by that codon → at least 1 tRNA per amino acid
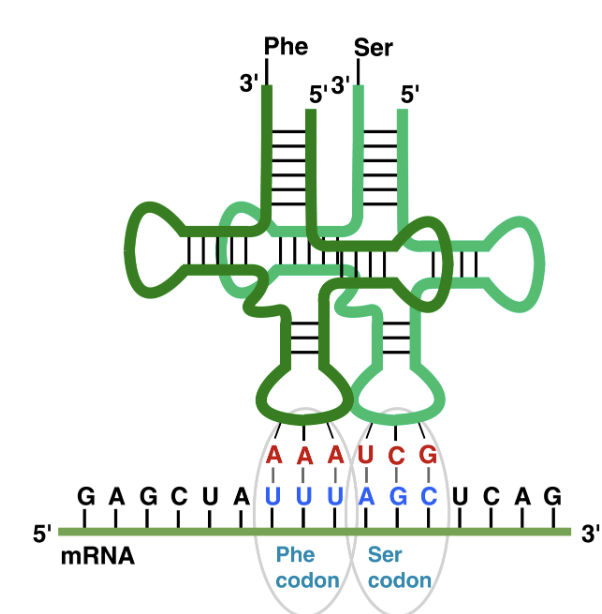
Codon vs Anti-Codon
Codons are in mRNA; anti-codons are in tRNA → they bind together according to base-pairing rules
How does the ribosome create a string of amino acids whose sequence is determined by the mRNA?
Ribosome causes tRNA anticodons to sequentially bind to mRNA codons + catalyzes the polymerization of amino acids on the other end of tRNAs
What is the process of converting info in the mRNA sequence to the protein sequence?
Translation
What determines which amino acid gets attached to each tRNA?
A class of enzymes called "Aminoacyl tRNA Synthases” catalyze the attatchment of specific amino acids to a tRNA
Notes about the Genetic code:
is it arbitrary (assigned by chance)?
is it fixed?
is it universal?
Genetic code is arbitrary, is fixed, and is almost universal → probably comes from a common ancestor of all organisms
Transcription vs Translation
Transcription = when DNA is transcribed into mRNA → happens first → done by RNA polymerase
Translation = when mRNA info is converted into proteins → after transcription → done by ribosomes
How does RNA polymerase know where a gene begins/ends + which strand to transcribe?
DNA has sequences called promotors → RNA polymerase recognizes them as the beginning of a gene → promotors are NOT transcribed
There are also termination sequences telling RNA polymerase when to stop
From the 2 strands of DNA, are genes transcribed always from one of the strands (ex: only the top strand) or can it be from both strands (ex: 1 from top, next 2 from botttom, next from top, etc)?
Genes can be transcribed from both strands, but always in the 5’ to 3’ direction → so in a diagram the direction indicates which strand
What is rRNA
A non-coding type of RNA that serves as the primary structural/functional component of ribosomes
Ribosomes consist of [#] subunits that are large complexes of __ and __
2 subunits of rRNA and proteins
Steps for Translation (until stop codon is reached)
Within a ribosome…
The small subunit finds the first start codon (AUG) within the mRNA and binds to it and also to the methionine tRNA (mRNA sandwich w/ small subunit + tRNA as bread)
This “sandwich”, or complex, recruits the large ribosomal subunit which has 2 binding sites: A & P
Large subunit binds so that the tRNA is in the P site → the tRNA that corresponds to the next codon in line binds to the A site
The carboxyl group of the methionine is transferred from the 3’ hydroxyl of the 1st tRNA (in the P site) to the free amino group of the amino acid bound to the 2nd tRNA (in the A site), forming the first peptide bond
Ribosome releases the 1st tRNA and shifts one codon in the 3’ direction → the tRNA that was in the A site (which now has an NH-met-ser-3’hydroxyl) moves to the P site
tRNA corresponding to the next codon comes to the A site → process repeats → continues until ribosome reaches a stop codon
In translation, what happens when a stop codon is reached?
A protein called a release factor binds to the A site → last tRNA to release the peptide chain + ribosomal complex disassembles
Peptide chain folds into a functional protein
What does Wobble pairing refer to?
Within translation: some tRNAs can use more than one codon → the 5’ residue of the anticodon has some wobble = doesn’t always have to match up perfectly with the 3’ residue of the codon
Wobble paring rules:
[G, C, A, U, I] pairs with __
G → C or U
C → G
A → U
U → A or G
I (inosine) → A, U, or C
Are the steps in the process of converting DNA info to protein sequential?
Not always:
In prokaryotes there’s no nucleus so translation can begin before transcription is complete
many ribosomes can bind to the same transcript, starting at the start codon, one after the other (called a polysome)
What are the 2 general catagories that mutations fall into?
Point mutations = small changes in a single gene
Chromosomal mutations = changes that affect a large protion of a chromosome, often affecting many genes
What does a Silent mutation do?
(Point mutation) They do not affect the protein sequence because of the degeneracy of the code (meaning the code is redundant or not functional)
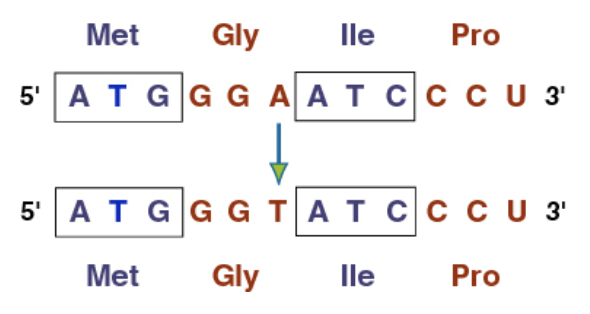
What does a Missense mutation do?
(Point mutation) They change one amino acid to another
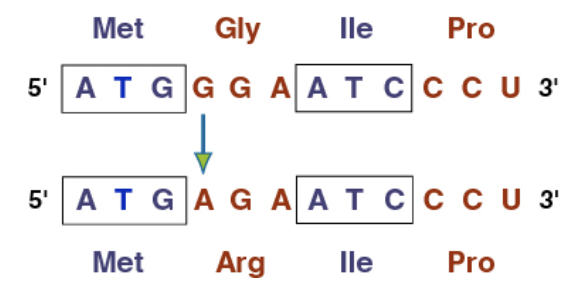
What does a Nonsense mutation do?
(Point mutation) They change an amino acid to a stop codon, cutting off the protein
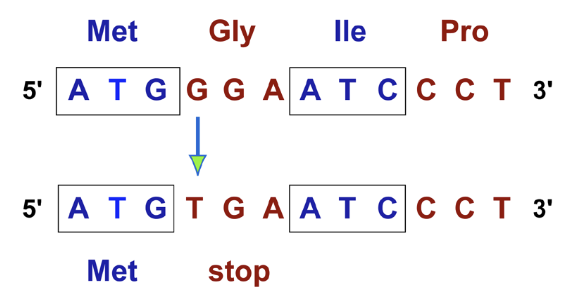
(Point mutation) What does a Frame-shift mutation do? What do they result from
(Point mutation) They change the reading frame from that point onward → result from either a deletion or insertion
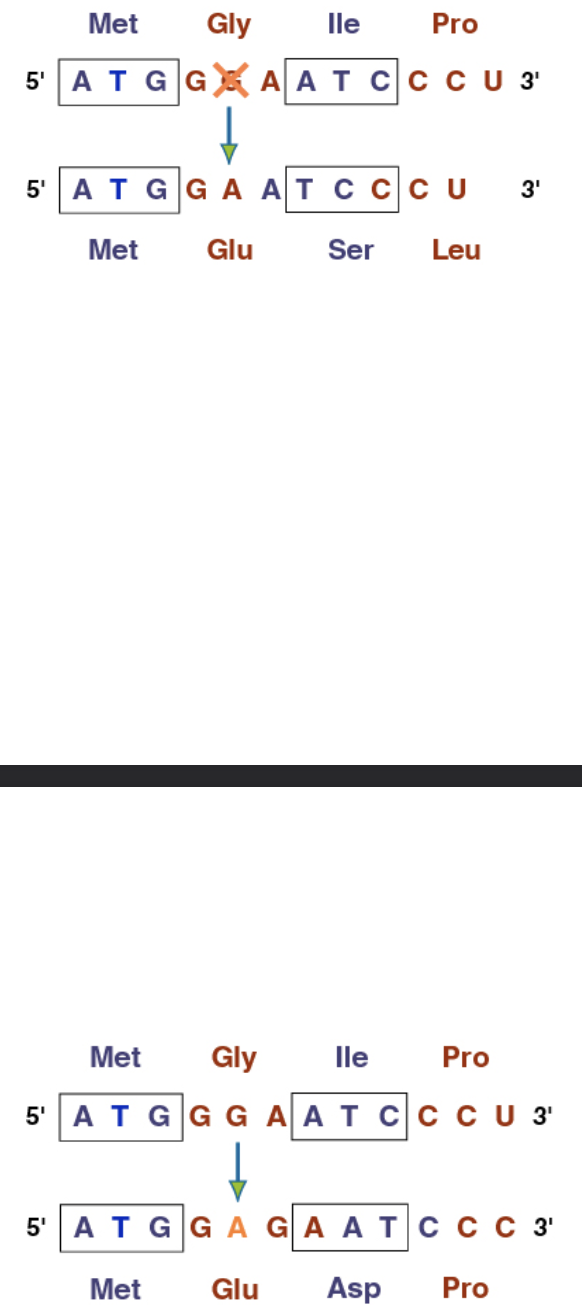

Chromosomal mutations: Deletion
Remove a large piece of the chromosome including many genes
Chromosomal mutations: Duplication
Duplicate large chunks of the chromosome
Chromosomal mutations: Inversion
When a piece of the DNA flips around, re-entering the chromosome in the reverse orientation
Chromosomal mutations: Translocation
When a piece of DNA jumps from one chromosome to another chromosome
Chromosomal mutations: Reciprocal translocation
Where two chormosomes trade place
The term “Mutagen” refers to…
Anything extraneous to an organism that causes changes
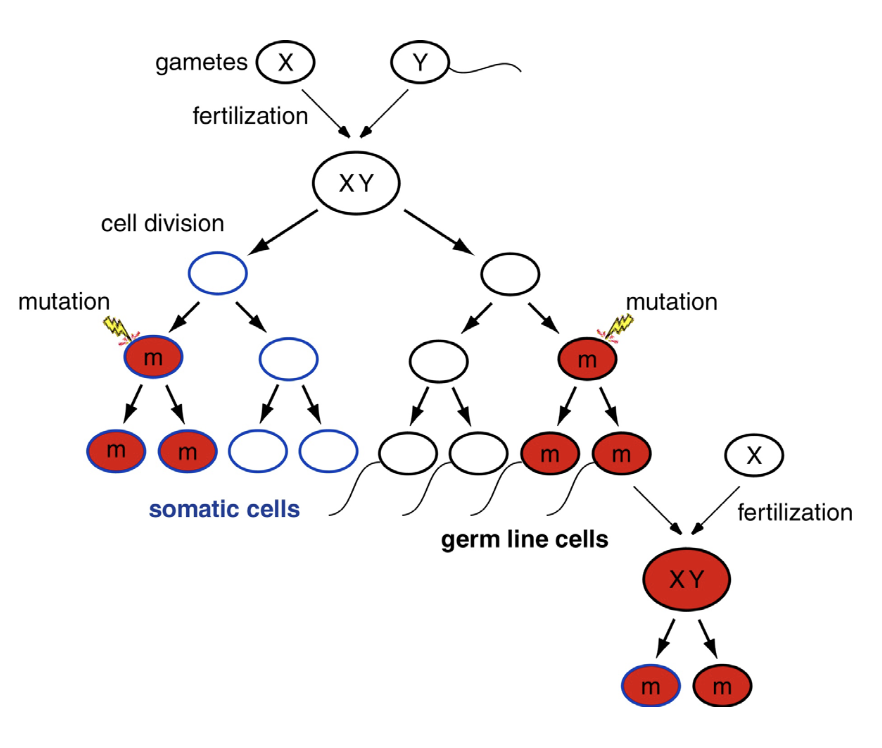
Mutations in somatic cells vs in the germ line
Transmitted to progeny?
In somatic cells: mutations could kill the cell or make it sick/cancerous → mutation prob inherited by mitotic progeny of the cell, BUT not transmitted to progeny bc somatic cells don’t make gametes
In the germ line: there’s possibility that mutation ends up in egg/sperm cell & transmitted to progeny → mutation, if not too deleterious, will become part of the allele repertoire in that population of organisms