BME 304 - Midterm 2
1/83
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
84 Terms
- EDTA is used, instead of guanidinium thiocyanate, to inhibit DNAse
- RNAse is added
Selection all of the following that are true about the process of DNA extraction.
- Chloroform is added to inhibit RNAse
- EDTA is used, instead of guanidinium thiocyanate, to inhibit DNAse
- A neutral to slightly basic phenol solution is used to allow DNA to migrate to the aqueous phase
- RNAse is added
- DNA ligase joins adjacent DNA fragments
- DNA polymerase is able to remove mismatched pairs of nucleic acid bases, improving the accuracy of DNA replication
Select all of the following that are true about DNA replication
- Okazaki fragments are formed when DNA polymerase synthesizes DNA in the 3’ to 5’ direction on the lagging strand
- DNA replication is very accurate because the strong covalent bonds formed between A and T or G and C are more stable than mismatches
- DNA ligase joins adjacent DNA fragments
- DNA polymerase is able to remove mismatched pairs of nucleic acid bases, improving the accuracy of DNA replication
All of the above
What is/are true about Polymerase Chain Reaction (PCR)?
- The starting and ending material of PCR are both DNA molecules
- PCR goes through cycles of denaturation, annealing, and elongation
- Each cycle of PCR doubles the amount of nucleic acid in the previous step
- All of the above
All of the above
What is true about real time rt-PCR?
-Fluorescence is proportional to the number of DNA molecules produced
- Reporter molecules are used to facilitate the visualization of DNA molecules amplification in real time
- Relative difference between the starting quantities of copies of the template DNA used can be estimated between different samples (e.g. control vs. experimental)
- All of the above
- A gene that is expressed constitutively and abundantly for basic cellular function
- It is used to remove variability resulting from e.g. minor pipetting error when comparing experimental and control samples in PCR studies
Select the following points that are true about a housekeeping gene
- A gene that is expressed constitutively and abundantly for basic cellular function
- It has variable expression level regardless of normal or pathological conditions
- It is used to remove variability resulting from e.g. minor pipetting error when comparing experimental and control samples in PCR studies
- The use of DNA ladder (or DNA markers) allows us to identify the molecular weight of different DNA fragments in a sample
Which of the following is/are true about gel electrophoresis?
- The positive charge of DNA allows it to travel in a properly set up gel electrophoresis
- The use of DNA ladder (or DNA markers) allows us to identify the molecular weight of different DNA fragments in a sample
- The heavier a DNA fragment, the further it travels away from the sample well
Northern blot - RNA (cDNA probes)
Southern blot - DNA (DNA fragment with a specific sequence)
Western blot- Protein (Antibody labeling)
Match up these choices with Northern, Southern, and Western blot
- Protein (Antibody labeling)
- RNA (cDNA probes)
- DNA (DNA fragment with a specific sequence)
- ddNTPs are used to terminate DNA synthesis because they have an H instead of an OH at the 3’ end
What is the purpose of using ddNTP is Sanger Sequencing?
- ddNTPs are used to terminate DNA synthesis because they have an OH instead of an H at the 3’ end
- ddNTPs are not used at all during Sanger Sequencing. Only dNTPs are used
- ddNTPs are used to extend the growing strands during DNA replication
- ddNTPs are used to terminate DNA synthesis because they have an H instead of an OH at the 3’ end
All of the above
Why are DNA libraries useful?
- The collection of DNA fragments cloned in vectors allows researchers to further identify and isolate their DNA fragments of interest for more investigation
- In cDNA libraries, the removal of introns (the non-expression region) from eukaryotic genes allows researchers to further study the function of that gene using simpler prokaryotic cells instead
- Genomic DNA library allows us to further study the entire genome of an organism
- Human genome project is an example of the application of DNA library
- All of the above
Garbage in garbage out (GIGO) effect should be avoided
For bioinformatics, which of the following statements is/are correct?
- Garbage in garbage out (GIGO) effect should be avoided
- An example of primary data is the collection of sequence data arranged by organism
- A coordinated database management can be made possible by using different annotation by different sequencing provider
- Endonucleases; phosphodiester bond
What kind of enzyme is used to cleave a polynucleotide chain in specific locations (not at the end of the chain)? What type of bond does it hydrolyze?
- Endonucleases; phosphodiester bond
- Phosphodiesterase; phosphodiester bond
- Exonucelases; hydrogen bond
- Exonucleases; phosphodiester bond
- 3’ hydroxyl extension
- 3’ sticky end
Based on the figure, what type of cut is made by Pvull in the recognition site below? Choose all that apply.
- 3’ blunt end
- 3’ hydroxyl extension
- 3’ sticky end
- 5’ hydroxyl extension
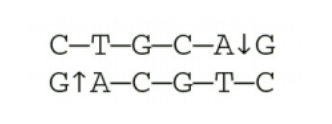
- MOI is the ratio of “agent (e.g. virus particle)” to “infection target (e.g. cell)”
- If the MOI is 10³, 10^4 virus particles are needed to infect 10 cells
Which of the following is/are correct about the Multiplicity of Infection? (MOI)? Choose all that apply.
- MOI is the ratio of “agent (e.g. virus particle)” to “infection target (e.g. cell)”
- MOI is the average number of cells that can be infected by one virus particle
- The most effective number of MOI is 2.5
- If the MOI is 10³, 10^4 virus particles are needed to infect 10 cells
- Own origin of replication that allows it to be replicated independently of the bacterial chromosome
Which of the follow is correct about plasmids?
- Small, circular pieces of DNA that exist and depend on the bacterial chromosome
- Usually necessary for the survival of bacteria
- Not natural, and can only be artificially made and grown in bacteria
- Own origin of replication that allows it to be replicated independently of the bacterial chromosome
- It is a M13 viral vector containing only essential genes
- It is a M13 viral vector containing some non-essential genes
- It is a Lambda viral vector containing only essential genes
- It is a M13 viral vector containing only essential genes
- It is a M13 viral vector containing some non-essential genes
- It is a Lambda viral vector containing only essential genes
- It is a Lambda viral vector containing some non-essential genes

- Surviving colonies after antibiotics incubation will be extracted
- Antibiotic resistance gene is normally added together with the gene of interest only to protect the bacteria that has successfully obtained the gene of interest when incubated with antibiotics
Which of the following is/are true about the isolation process involved in genetic engineering of bacteria? Choose all that apply.
- Surviving colonies after antibiotics incubation will be extracted
- Antibiotic resistance gene is added together with the gene of interest to destroy the bacteria that has successfully obtained the gene of interest when incubated with antibiotics
- The purpose of the isolation process is to eliminate bacteria of different sizes
- Antibiotic resistance gene is normally added together with the gene of interest only to protect the bacteria that has successfully obtained the gene of interest when incubated with antibiotics
PCR uses the DNA template to build, alongside primers, dnTPs, DNA polymerase, and more, to build the complementary strand with
How does PCR in the lab mimic this natural process of replication?
They can be tail-less or have a head and tail (lambda and M13 are what we are focusing on)
What phage groups exist?
M13 and Lambda phages, which both infect E Coli.
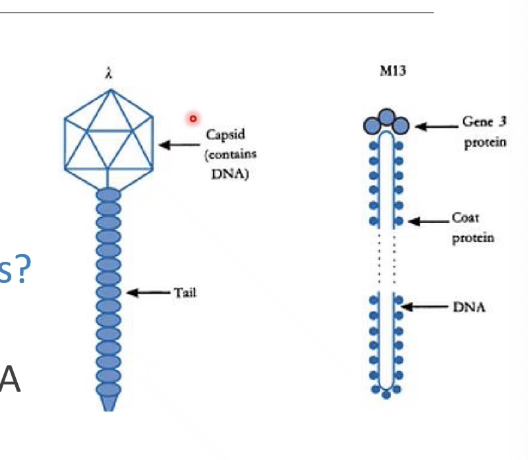
Lambda - large genome and has non-essential genes, and has dsDNA
M13 - smaller genome and only has essential genes, and only has ssDNA
What are the most common viral vectors, and what are the key similarities are differences?

Number of virus particles needed in infect 1 cell, and the formula is (number of viral particles or viruses/number of successfully infected cells)
What is multipicity of infection (MOI)? How do you calculate it?
Number of viruses / Number of cells successfully infected
(1. 50/20 = 2.5
2. 200/80 = 2.5
3. 2000/800 = 25
4. 5000/800 = 62.5)
How do you calculate MOI of these rows?
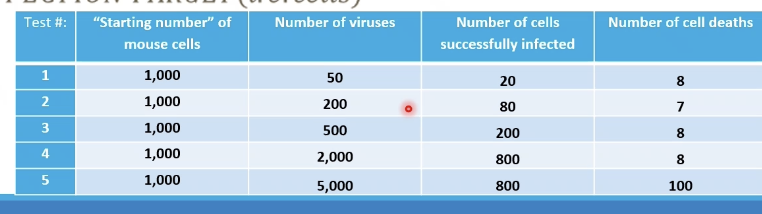
There is toxicity at the high virus concentration, and the cell becomes cytotoxic. It is more effective to have a low MOI instead of a high one!
High value of MOI means what?
Plasmids are small circular dsDNA that exist independently of chromosomal DNA.
What are plasmids?
They have their own origin of replication, and they can replicate without the bacterial chromosome
How are plasmids replicated?
Conjugation
_______ is when two bacterias are in direct contact with each other and they pass DNA, requiring direct contact
Transformation
______ is when a bacteria finds loose DNA in its environment and absorbs it as its own
Transduction
________ is when a virus infects bacteria and takes its DNA and injects it into the cell that it infects. The virus essentially becomes the unintentional middle man.
Cut open plasmid and paste in the gene, which relies on restriction enzymes
Insert the plasmid into the bacteria
Plasmids are transformed into cells and used as factories to create proteins
How do we clone DNA into plasmids, grow plasmid DNA, and transform plasmids into cells?
Conjugative plasmids carry transfer genes, are usually large and have a low copy number
Non-conjugative plasmids carry non-transfer genes, are usually small, and have a big copy number
What are conjugative and non-conjugative plasmids?
Origin of replication
Markers
Cloning sites
Promoting sites/regions
Name the typical elements of a plasmid.
YACs are yeast artificial chromosomes, carrying large gene inserts and less stable
BACs are bacterial artificial chromosomes, carrying smaller gene inserts and are more stable
What are YACs and BACs?
Type I cut DNA at random sites and are far from recognition sequence.
Type II cut DNA at specific sites and are close to the recognition sequence
What are Type I and Type II restriction enzymes?
NO - do not pay attention to Roman numerals
Can you tell the type of restriction enzyme based on the name?
5’ phosphate extension, 3’ hydroxyl extension (both sticky ends)
or a blunt end
What types of ends can be produced?
LacZ gene
What is used as a reporter to tell whether a plasmid has successfully taken up a foreign DNA insert?
plasmids
In many ______, multiple cloning sites are located in the Lacz gene
LacZ stays functional and produces blue colonies
What happens to LacZ if no foreign DNA is inserted?
It disrupts LacZ, and the colonies stay white
What happens to Lac Z when foreign DNA is inserted?
It is a quick color test to tell which bacteria got the plasmid with the foreign DNA insert. The goal is to pick white colonies because those are the ones most likely carrying the foreign DNA.
What is blue-white screening?
A blue colony is where lacZ still works, and foreign DNA is NOT inserted
A white colony is where lacZ is broken, and foreign DNA IS inserted
Explain the difference between a blue colony and a white colony
Bacteria phage lambda
What does the ci gene come from?
It works as a transcriptional repressor. If a DNA fragment is inserted into the ci gene, then ci is disrupted. This disruption tells whether colonies survive and how they appear.
How does the ci gene work?
They undergo lytic cycle
What happens is the ci gene for selection is disrupted?
If a bacteria takes up the plasmid, it can form a colony and survive
If a bacteria doesn’t take up the plasmid, it will die.
*Remember transformation is not perfect and not every bacteria will take up a plasmid!
How is an antibiotic resistance gene used for selection?
We use them for studying gene function, such as human disease, gene disruption, or replaced.
What do we use transgenic animal cells for?
Recombination is when DNA is integrated into the genome. The two major types and homologous and non-homologous
What is recombination, and what are the possible methods?
Homologous - uses matching DNA sequence to insert at the specific target location (desirable outcome)
Non-homologous - more random, and DNA is inserted at an unintended location (NOT specific)
What is the difference between homologous and non-homologous recombination, and what can we accomplish with each?
Positive - Neomycin-resistant gene (inserted IN the target gene)
Negative - Thymidine-kinase gene (inserted OUTSIDE (next to) the target gene)
What are examples of a positive selectable marker and a negative selectable marker? Where are they inserted?
Selectable markers
What is able to alter a cloned gene?
It is a positive selection marker. Cells are grown in neomycin/G418. Only cells that successfully took up the DNA construct with Neoᴿ survive.
Memory trick: Neo = “DNA got in.”
What does the neomycin-resistance gene (Neoᴿ) select for?
It is a negative selection marker. If a cell has TK, TK activates ganciclovir into a toxic form, so the cell dies. Cells without TK survive. Correct homologous recombination usually gives Neoᴿ TK⁻, so the cell survives both neomycin/G418 and ganciclovir.
Memory trick: TK + ganciclovir = death.
What does the thymidine kinase gene (TK) + ganciclovir select against?
It allows us to simulate DNA manipulation before doing the experiment in the lab. We can check restriction enzymes have cut, or if gene inserts are in the right direction
What does snapgene do?
Multiple cloning sites (MCS)
What is this part of a snapgene map called?
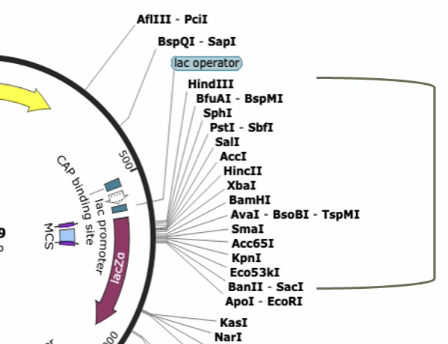
If a gene is inserted in the wrong direction relative to the promoter, it may not be expressed correctly. Snapgene arrows are very important!
Why does gene directionality matter in snapgene?
The origin of replication needs to be compatible with the intended host cell (it can determine what species the plasmid can replicate!)
It needs to be in the right orientation
Cutting site needs to be unique (can’t cut too many times because that makes the result hard to interpret)
The final plasmid has to has a selection marker
What are some important guidelines to follow when choosing restriction enzymes for cloning?
Gel electrophoresis can separate DNA fragments based on size.
Small fragments move down, and big fragments are towards the top.
How do you verify if cloning works?
Two bands, where one matches the size of inserted gene, and the other matches the size of the remaining vector backbone
For a successful double digest, we expect how many bands if we cut both sides of the insert?
2, where one band is the same size as your gene insert and the other is the same size as the original vector
How many bands should you see as a result of cutting the plasmid with 2 restriction enzymes?
Disruption (breaking cell wall), Separation (centrifugation), Recovery, (elution using water) (DSR)
What are the 3 major steps of isolating nucleic acids?
RNAse - isolating DNA (getting rid of RNA)
DNAse - isolating RNA (geting rid of DNA)
What are RNAse and DNAse used for?
It is similar because template DNA is natural
It is different because the primers and nucleotides (dnTPS) are artificial
How is PCR similar and different from natural replication?
Template DNA, primers, DNA polymerase, dnTPS, and buffers.
What are the common elements of PCR?
The number of copies increase exponentially (x → 2 to the x)
How does the number of copies increase over PCR cycles?
Denaturing, Annealing, and Extending
Name the 3 steps of PCR
The temperatures are important because PCR is basically forcing DNA to separate, primers to attach, and polymerase to copy, over and over again. PCR needs specific temperatures because each temperature controls a different “behavior” of DNA and the enzymes.
What temperatures are used for PCR and why?
aqueous buffer with Mg2+, primers 1 and 2, and a thermostable DNA polymerase
What chemicals are needed for PCR?
Reverse transcriptase, and its purpose is to detect and measure RNA
What does the RT stand for in RT PCR, and what is its purpose?
The two steps are RT step, and PCR step
RT: RNA goes to cDNA
PCR: cDNA is amplified
What are the two steps of RT PCR?
Real time RT PCR is able to monitor the amplification in real time, using signals to quantify RNA levels and gene expression.
What is real time RT PCR, and what is its purpose?
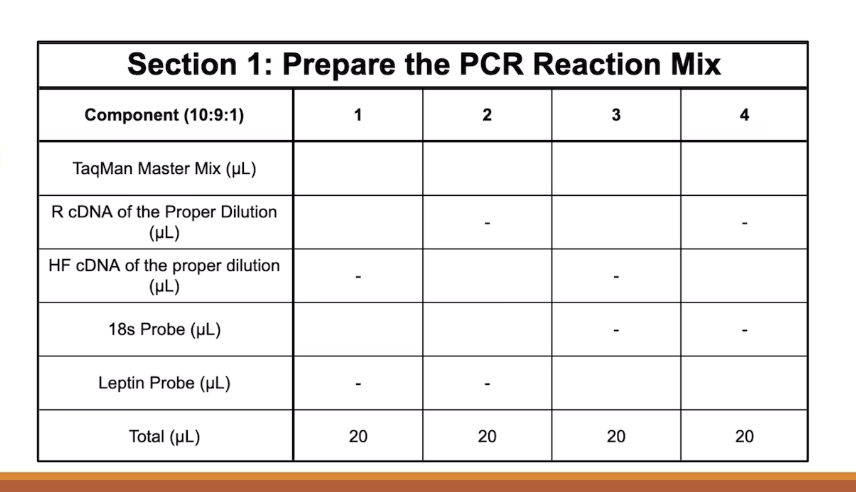
Basically each column’s blanks need to add to 20. So TaqMan Master Mix is always 10, and then going down it is 9 and 1
How do you fill the blanks of this table?
RNA (where DNAse can isolate it)
What is REQUIRED to start an rt-PCR evaluation?
An uptake of leptin because leptin is what controls hunger
What would you see in an obese mice’ leptin levels?
A housekeeping gene, which is used to normalize data from slight variations
What is used to normalized the Ct value?

What is the Fold Change equation?
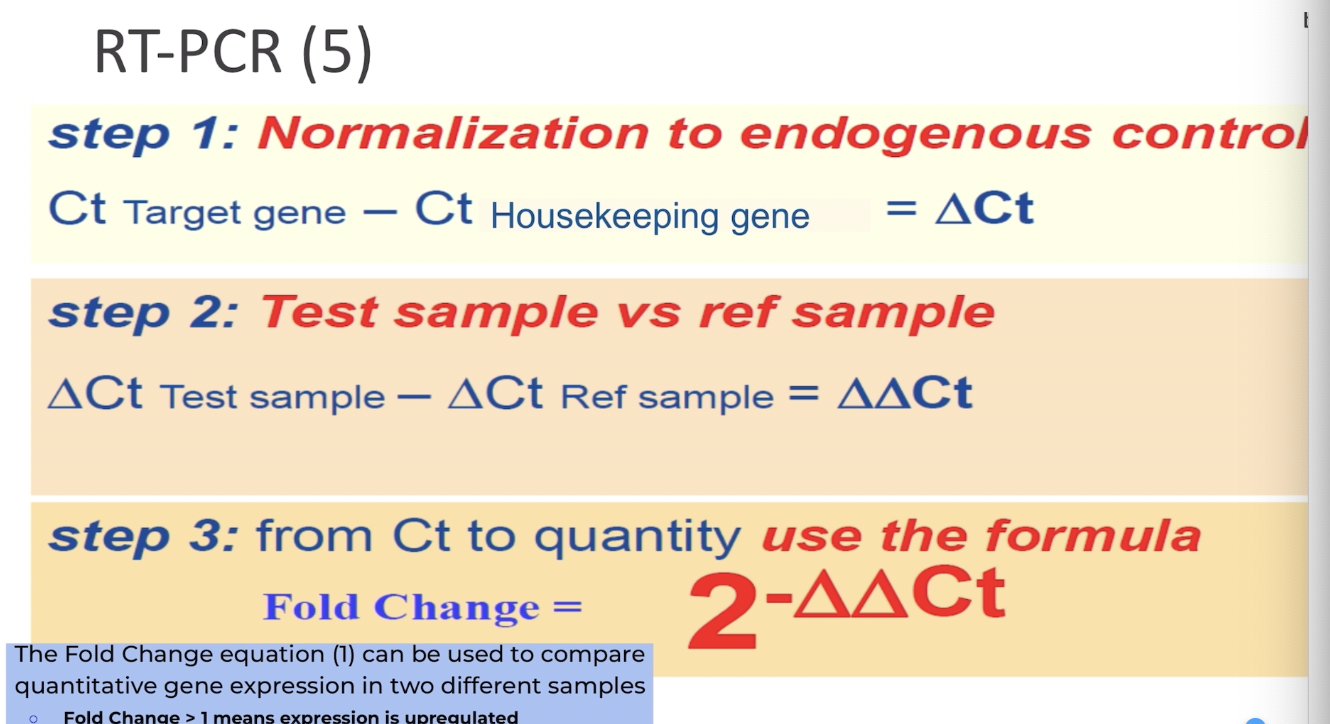

What are the rt-PCR equations using Ct? You need to memorize these!!
Fold change > 1 means expression is upregulated
Fold change < 1 means expression is downregulated
Fold change = 1 means there is no change
Explain what each of the following mean:
Fold change > 1
Fold change < 1
Fold change = 1
A DNA library is a collection of DNA fragments that have been cloned into vector. They are useful for researchers in order to identify certain DNA fragments.
What are DNA libraries, and what are they used for?
Main two are genomic and cDNA.
Genomic - contain the entire genomic DNA of a gene, containing introns and exons
cDNA - are made from reverse transcriptase, where there are only exons
What main types of DNA libraries exist, and what are the key differences?
Steps: isolation, digestion, insertion and amplification
How are DNA libraries produced?
PCR
How can we retrieve a clone that contains a specific sequence?
dNTPS and ddNTPS
What are the two molecules utilized by Sanger sequencing?
Molecule Y is the dNTP because of the OH group
Molecule Z is the ddNTP because of the H group
Why