FW 453 Exam II Flashcards
1/207
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
208 Terms
Why estimate population?
Management and conservation: understand population status, predict and evaluate management consequence/outcomes
Science: evaluate pop status and change via environmental change
What are the main challenges of estimation?
sources of error:
spatial variation
detection probability
is your pop “closed” or “open”?
Also, the methods we use must be relevant to the scale we want to estimate at!
pop, landscape, community
formula for detection probability
N^ = C/p^
(inferences (^) about N require inferences about p)
census
detection probability equals 1 and we count ALL individuals
index
the count represents some number equal to, or less than, the true number of individuals present
detection probability constant but <1
estimate
a measure of a state variable or vital rate based on a sample of observations
typically simultaneously estimates detection probability
Estimation examples in practice
distance sampling
capture-mark-recapture
occupancy modeling
What is distance sampling?
widely used method for estimating abundance and density (population size/area)
robust and very popular methodology
addresses imperfect detection
samples (typically lines or point) are conducted such that objects (e.g. animals, nests, dens, etc) are detected from the sample
auxiliary info is taken (distances to objects)
allows estimation of detection probability as a function of distance to the observer
Distance Sampling Types
line transects
point transects
Line Transects
sample unit is a line of length L placed at a location in the study area
observer moves along transect recording all individuals detected and their perpendicular distance to transect
count only objects observed from the line
Assumptions of distance sampling with line transects
animals directly on the line are always detected
animals are detected at their initial location (no movement response)
distances are measured accurately
transect lines are placed randomly
observation of animals are independent from each other
there are sufficient samples to estimate the detection function
Distance sampling with line transects
when detection is not perfect, then we must determine the area sampled in order to estimate density
D ̂= n/(2Lμ ̂ )
•μ ̂ is the “effective half width” of the survey
– Note: when detection is perfect μ ̂=w
Note that w in strip sampling is set by the user, here, the mew must be estimated!
Violation of independence: clusters
animals may be in groups
objects in groups are not independent
treat cluster as the object of detection
estimating density of clusters
need mean cluster size (and variance)
need to account for variation in detection (larger clusters more likely to be detected)
How to estimate μ (effective half width of the survey, which is equal to w when detection is perfect)
based on observed frequency distribution, f(x)
the relative frequency at which distances x from the centerline occur in the sample data
convert to a probability density function
What is f(x) in distance sampling line transects?
f(x) is the observed distribution
probability of observing an animal at distance x, given that the animal is present and detected somewhere on the transect
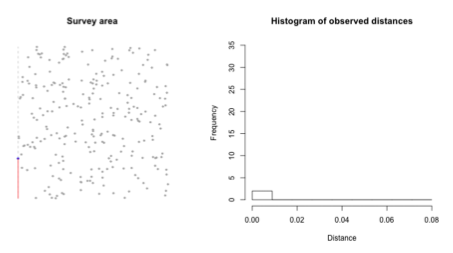
What is g(x) in distance sampling line transects?
g(x) is the detection function
probability of detecting an animal, given that it is at distance x from the centerline
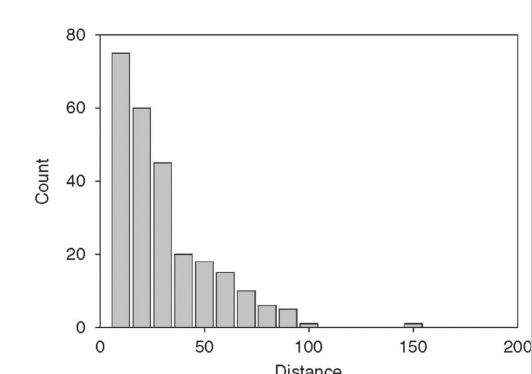
Where does μ come in with graphs?
f(x) = g(x) / μ
this makes f(x) a probability density function
these are easy to work with for estimation problems
There is a relationship between the detection function and the histogram of count data and that relationship is a function of the effective half width of the survey
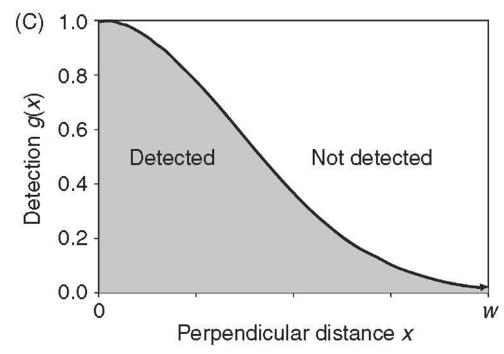
Estimation of distance function
f(x) = g(x) / μ
first assumption: we detect objects on the line 100% of the time
g(0) = 1
f(0) = 1/μ
μ ̂=1/(f ̂(0))
If we can estimate f(X), and evaluate at x=0, then we can estimate μ
How do we select a model for f(x)?
How do we use sample data to estimate f(0)?
Model for f(x)
Key function is a basic shape for the detection data that describes the way the detection changes as a function of increasing distance from the center line
We will let Program Distance fit the functions and we will use AIC to determine which is the best model for our specific data (we’ll do this in lab)
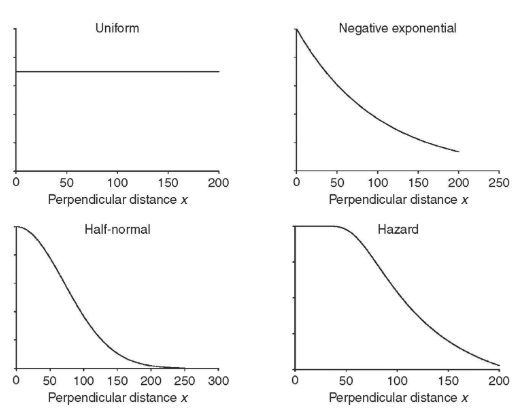
How do we use sample data to estimate f(0)?
after we use program distance to fit a key function to our data, it will use statistical methods to:
estimate f(0)
which will allow estimation of μ
which will allow estimation of density
Why do we add covariates to the model?
The model fit is often poor without the addition of covariates that are likely to influence detection probability
homogenous bias (observer bias)
temporal bias (weather/other conditions of survey)
series adjustments (i.e. covariates) are added to improve the model fit like…
f(x) = key(x)[1+series(x)]
(VIS = visibility; poor/fair (e.g., glare, light fog or rain) or excellent; LIGHT = light conditions; overcast, mostly cloudy, or partly cloudy/clear). ESW = Effective strip width
(meters). p = Detection probability. ^NN = Abundance estimate. Goodness of Fit metrics: C-S = Chi-squared; K-S = Kolmogorov-Smirnov; C-vM = Crame´r-von Mises
N<
estimated abundance
p<
estimated detection probability
Distance sampling of polar bears
line transect aerial survey to estimate abundance
compared against satellite-derived surveys which estimated abundance using capture-recapture
clouds are factors that may hamper detection
water conditions are factors that may hamper detection of bears. foam accumulating along edges of water could be confusing
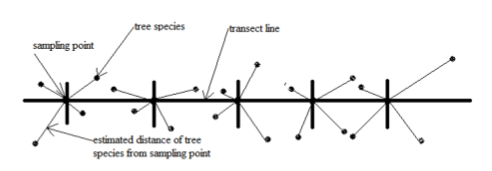
Point Transect Sampling
establish k points to sample, n observations are recorded at each point
a point is placed in the study area
observer spends a predetermined amount of time at the point attempting to detect individuals
the count of individuals is recorded as well the distance of each from the observer
Density estimation
D ̂=n/(kπw^2 )
k is the number of points surveyed
n is the number of animals detected
w is the radius of the circle and we assume detection probability is perfect out to this distance
kπw2
the cumulative area of the k points surveyed
the frequency of observations at each radial distance (r)
f(r)
analogous to f(x)
h(r)= g(x) detection function for ___
radial distances r
we use the same method: fit a key function to f(r)
assumptions are the same as for line transects
D ̂=(nf′(0))/2kπ
instead of using f(0) for density estimation, we use f’(0)
f’(r) is the derivative of f(r)
remember
μ ̂=1/(f′(0))
Sampling principles
What is the objective?
What is the target population?
What are the appropriate sampling units?
quadrats?
point samples?
line transects?
3 types of sampling units
Quadrats
point samples
line transects
Comparison of line transects vs. point counts
line transects
typically preferred for mammals
cover larger areas more quickly
distance estimation when moving may be more difficult
point counts
typically preferred for birds
more practical in rough terrain
fixed time at each station
more time to see or hear
especially important for birds in high canopy
less area covered to see or hear
especially problematic for mammals
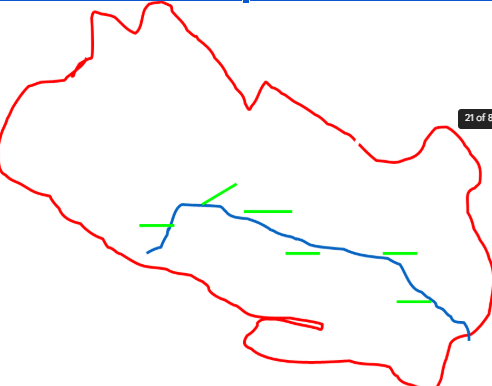
Advantages and disadvantages of samplings units—nonrandom placement
advantages
easy to lay out
more convenient to sample
disadvantage
do not represent other (off road) habitats
road may attract (or repel) animals
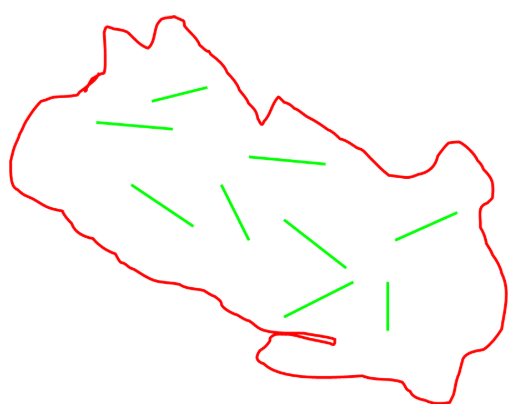
Advantages and disadvantages of samplings units—random placement
advantages
valid statistical design
represents study area
replication allows variance estimation
disadvantage
may be logistically difficult
may not work well in heterogeneous study areas
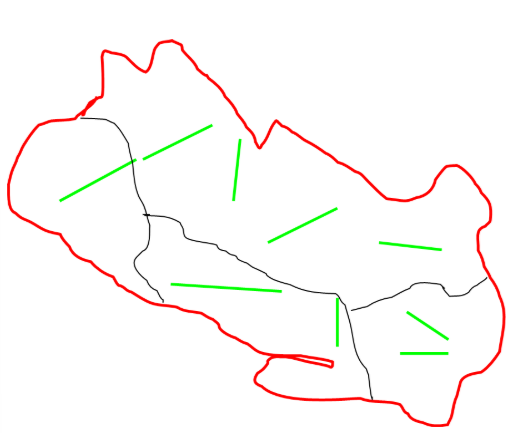
Advantages and disadvantages of stratified sampling
advantages
controls for heterogeneous study area
allows estimation of density by strata
more precision
disadvantages
more complex design
Takehomes for Strip transects
Detection is assumed constant across the transect
Tends to underestimate density
Take homes for line/point transects
Distances are recorded for each observation
Detection is modeled as a function of distance
Effective transect width/point area is modeled based on the detection function
Density estimates take into account the effective transect width and missed individuals
Variables N, C, and p in detectability meaning
N= abundance
C= count statistic
p = detection probability; P(member of N appears in C)
C = pN
Inferences about N require inferences about p
N> = C/P>
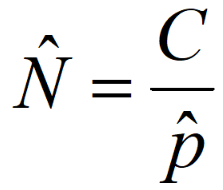
Lincoln-Petersen Method
Frederick Lincoln and Gustav Petersen
Capture, mark and release a sample of individuals from a population
At a later date, take a second sample
Proportion marked individuals in the second sample indicates proportion of total pop we sample each time
detection probability!
Marking animals for L-P: Batch marking
Groups of animals are given the same marks
i.e., marks are not uniquely identifying, animals are simply either marked or not
Examples?
Limitations?
the basic estimate for abundance
N = n1n2/m2
N = abundance
n1 = the number of animals caught and marked at capture occasion 1
n2 = the number of animals caught at capture occasion 2
m2 = the number of marked animals caught at capture occasion 2
What is detection probability (p) under the Lincoln-Petersen model?
p = m2 / n2
We commonly use an adjusted estimator that is unbiased for small sample sizes:
N = ((n+1)(n2+1))/(m2+1) — 1
If sample size is small, the LP estimator is biased. For example, what happens if the number of recaptures is zero?
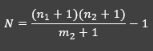
The estimated variance is calculated as:
var(N) = ((n1+1)(n2+1)(n1-m2)(n2-m2)) / ((m2+1)²(m2+2)
The approximate 95% Cls are
N ± 1.96 sqrt(var(N))
Rabbit abundance in Oregon example
Rabbits were captured in central Oregon and marked by dying their tails and hind legs with color
87 were initially captured and marked
Live trapping
14 were captured a second time, of which 7 were marked
Drive counts
Based on the results, we can set up the problem with:
n1 = 87
n2 = 14
m2 = 7
Then we use the adjusted L-P to find our estimate of N
𝑁=((𝑛_1+1)(𝑛_2+1))/(𝑚_2+1)−1= (87+1)(14+1)/(7+1) −1=164
We can also calculate the variance of N and the approximate 95% Cl
var(N) = 1283.33
The 05% Cl is (93.8 to 234.2)
Overall detection probability = 0.5
p = m2/n2 = 7/15 = 0.5
But they used 2 diff methods so we can see which had the higher detection probability
live trapping: p = 87/164 = 0.53
drive counts: p = 14/164 = 0.09
because of two different methods, our estimate of p is not very good, we need to estimate p separately for each method. Live trapping has much higher detection probability so is a better method for estimating N in this population.
The variation in capture probabilities between occasions is allowed in L-P!
Assumptions:
the pop is closed
all animals have the same probability of being caught within an occasion
first occasion capture does not affect second
there is no tag loss or other loss of marks
Catching an animal does not affect its probability of capture on subsequent occasions. Marks are accurately recorded.
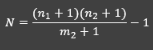
Closed population
Abundance is constant during the study
No gains or losses (i.e., births, deaths, immigration, emigration)
Most of the models we do, will make this assumption
Open population
Abundance may be constant or change
Subject to gains and losses
We can estimate survival, recruitment, movement, etc. if the sampling is well designed
Assumption of equal detection
All animals have the same probability of being caught within an occasion
What might impact this assumption?
i.e. when might it be violated?
Sampling technique
Gear
Time of year
Home range size differences between sexes
Violations of Equal Detection
Trap happy
Animals are more likely to show up as recaptures than would be predicted
p will increase, N will decrease
Trap shy
Animals are less likely to show up as recaptures than would be predicted
p will decrease, N will increase
Individual Marking
Marking animals individually gives us extra information
Can relax some assumptions of Lincoln - Petersen
Allows use of more complex models with multiple samples (occasions)
Allows us to account for variation in capture probabilities!
Multiple sample CMR
2 or more occasions
Advantages
more data to estimate N=more precision
avoid some assumptions of LP
all animals NO LONGER have the same probability of capture
probability of capture can be affected by previous experience of capture
M0 model
constant detection probability (p)
Over time, individuals, behavior
Mt model
only time effects
– L-P is a special case with k=2
Mb
only behavioral effects
Trap happy or shy
Mh
individual effects
Heterogeneity in capture probabilities
Combination of model effects
Mth, Mtb, Mbh, etc
What does 00000 mean in multiple sample CMR?
not captured at all
𝜔
individual encounter histories
encounter history
An encounter history tells us whether an individual has been captured/detected during each sampling occasion
Written as a 1 if an individual is encountered and a 0 otherwise
For 2 time occasions:
01: not captured first time, captured second sample
10: captured, not captured
11: captured, captured
00: not captured at all
Using the encounter history we can summarize the data into totals for each category:
𝑋10 number caught on the first occasion but not the second
𝑋01 number caught on the second occasion but not the first
𝑋11 number caught on both occasions
𝑋00 number never caught
𝑟 = 𝑋10 + 𝑋01 + 𝑋11 total number of different animals caught over the study
Total of all is our population size, so why not just add up all 4 categories?
How do encounter histories help us estimate?
We will calculate the probability of each unique combination of 1s and 0s (Pr(𝜔))
Using Pr(𝜔) and the observed frequency of each unique combination of 1s and 0s (𝑋𝜔), we can obtain estimates of capture and recapture probabilities
Via maximum likelihood, which we will leave to Program MARK
With this information, we can then estimate 𝑁 by what is basically an extension of the L-P estimator
𝑝𝑖𝑗
capture probability for each individual i at each time j
𝑐𝑖𝑗
recapture probability for each individual i at each time j
If we assume capture probability is constant over time then…
𝑝𝑖𝑗 = 𝑝𝑖 and 𝑐𝑖𝑗 = 𝑝𝑖
𝑘
the number of capture/recapture occasions during the study
Review of basic probability rules
Rule 1: Probabilities must be between 0 and 1
Rule 2: The sum of the probabilities for all possible outcomes of an event must equal 1
Rule 3: The probability that an event does not occur is 1 minus the probability that it does occur.
Rule 4: The probability that two events both occur together is found by multiplying the probabilities that they occur alone
Rule 5: If two events CANNOT occur together, the probability of them both occurring is found by adding their probabilities
Model M0 assumes…
constant and homogenous p
i.e., capture probability is constant across time and individual
Therefore, 𝑝𝑖𝑗= 𝑝 and 𝑐𝑖𝑗=𝑝
It has 2 parameters: N and p
Say we have:
𝜔 = 101
what is Pr(𝜔) under m0 model?
Pr(𝜔) = 𝑝(1 − 𝑝)𝑝
How about:
𝜔 = 000001
(what formula under M0?)
Pr(𝜔) = (1 − 𝑝)(1 − 𝑝)(1 − 𝑝)(1 − 𝑝)(1 − 𝑝) 𝑝
which can be written as:
Pr(𝜔) = (1 − 𝑝)5𝑝
IF 𝜔 = 000000…
under model M0, Pr(𝜔) =
Pr(𝜔) = (1 − 𝑝)6
Calculating Pr(𝜔) under Model Mt
𝑝𝑖𝑗 = 𝑝𝑗 and 𝑐𝑖𝑗 = 𝑝𝑗
So model parameters are N and p1…pk
Now, say we have
𝜔 = 101
"Pr(𝜔) = 𝑝1(1 − 𝑝2)𝑝3"
If we have k=6 occasions, then we might have
𝜔 = 101011
Pr(𝜔) = 𝑝1(1 − 𝑝2)𝑝3(1 − 𝑝4)𝑝5𝑝6
How about:
𝜔 = 000001
Pr(𝜔) = (1 − 𝑝1)(1 − 𝑝2)(1 − 𝑝3)(1 − 𝑝4)(1 − 𝑝5) 𝑝6
Calculating Pr(𝜔) under Model Mb
If we assume capture probability varies after the first capture, but is constant otherwise, then
𝑝𝑖𝑗 = 𝑝 and 𝑐𝑖𝑗 = 𝑐
For our previous encounter history
𝜔 = 101011
Pr(𝜔) = 𝑝(1 − 𝑐)𝑐(1 − 𝑐)𝑐𝑐
We can reduce this to:
Pr(𝜔) = 𝑝(1 − 𝑐)2c3
How about:
𝜔 = 100001
Pr(𝜔) = 𝑝(1 − c)(1 − c)(1 − c)(1 − c) c
Model M_h
This model allows capture probabilities to vary by animal, due to heterogeneity, but there is no trap response or time variation.
The parameters in the model are:
𝑁
p1…𝑝𝑖 - the capture probability of animal 𝑖
When might we use this model?
Calculating Pr(𝜔) under Model Mh
If we have individual effects, the model can get complicated:
𝜔 = 101011
Pr(𝜔) = 𝑝𝜔(1 − 𝑝𝜔) 𝑝𝜔(1 − 𝑝𝜔) 𝑝𝜔 𝑝𝜔
Basically a unique detection probability for each individual
How about:
𝜔 = 100001
Pr(𝜔) = 𝑝𝜔(1 − 𝑝𝜔)(1 − 𝑝𝜔)(1 − 𝑝𝜔)(1 − 𝑝𝜔)𝑝𝜔
What happens when you combine behavioral, time, and individual effects?
If we have behavioral, time, and individual effects, the model can get very complicated:
𝜔 = 101011
Pr(𝜔) = 𝑝𝜔1(1 − 𝑐𝜔2) c𝜔3(1-c𝜔4)𝑐𝜔5𝑐𝜔6
Basically a unique detection probability and recapture probability for each individual at each time step – difficult to fit without plenty of data!
Estimation for p and c
We will use a likelihood based approach
Use observed frequencies (𝑋𝜔) and (Pr(𝜔))
Multinomial likelihood
Estimates p and c probabilities
Use this information to estimate N!
Variables in model variation in capture probability
Model variation in capture probability as a function of
Time (t)
Behavior (b)
Individual heterogeneity (h)
Combinations (b*t, b*h, etc.)
Compare models
AIC
What is the best way to estimate abundance with 2 samples?
The Lincoln-Petersen method
Basic multi-sample CMR model types
M_0 — constant capture probability (p)
over time, individuals, behavior
M_t — only time effects
L-P is a special case with k=2
M_b — only behavioral effects
trap happy or shy
M_h — individual effects
heterogeneity in capture probabilities
combinations of effects (M_th, M_tb, M_bh, etc)
Cormack-Jolly-Seber Model
apparent survival (cannot differentiate death from emigration)
capture probability—but cannot estimate N!
Jolly-Seber model
estimation of survival and recruitment (theoretically)
can estimate N with strong assumptions but difficult to fit
Pollock’s robust design model
combines closed and open modes, N, survival and recruitment among other things
incredible amount of flexibility
pj
capture probability for at each time j
if we assume capture probability is constant over time then pj = p
φ𝑗
apparent survival for at each time j
if we assume apparent survival is constant over time, then φ𝑗 = φ
Why is there no N in the Cormack-Jolly-Seber model?
we are not interested in abundance, only apparent survival
Closed population vs open population diagrams
In the closed population, animals don’t leave, so there is just the “marked and release” tab with “captured” (p) and “not captured” (1-p) counts. But with open populatoin, we start with the “marked and released” tab which splits into “alive” (𝜙) and “dead/emigrated" (1-𝜙),” and “alive” is split into “captured” (p) and “not captured” (1-p)
Types of CJS models
M00
Mt0
M0t
Mtt
CJS model M00
constant detection and survival
2 parameters: 𝑝 and 𝜙
CJS model Mt0
time specific detection, constant survival
Separate p for each time period and a single 𝜙
CJS model M0t
constant detection, time specific survival
Separate 𝜙 for each time period and a single p
CJS model Mtt
detection and survival are time specific
Separate 𝜙 AND p for each time period
CJS M_00 — calculating Pr(𝜔)
Now we have p and 𝜙, the detection probability and survival rate respectively
𝜔=101
Pr(𝜔)=𝜙(1−𝑝)𝜙𝑝
We condition on the animal being captured in time 1. So we don’t estimate p or 𝜙 for time 1. We ONLY consider animals with a 1 for the first period
This 𝜙 is the probability the animal survives from the PREVIOUS time (Time 1)
We know it (the 101) survived since we captured it in Time 3!
We know this animal survived to time 3, therefore it was alive during Time 2 but not recaptured (1−𝑝)
This animal was recaptured in Time 3, therefore it was both alive and captured (𝜙𝑝)
These quickly get more complicated because we not only have to keep track of whether an animal is detected in a given period, but whether they were detected in subsequent periods
This gives us the information we need about survival to calculate P(𝜔)
Given the follow capture histories, what are the probabilities under M00?
111
110
101
100
𝜙p𝜙p
ignore the first 1 and only consider the next two 1s. Each one has 𝜙p
𝜙p(1-𝜙p)
Ignore first one and consider the 0. The second 1 gets 𝜙p but this time we also multiply by (1-𝜙p) because it was not recaptured (0)
𝜙(1-p)𝜙p
Ignore first 1. Now we have 0 and 1. The 0 indicates it wasn’t recaptured, so we do 𝜙(1-p). But then it was recaptured so it gets a 𝜙p
(1-𝜙) + 𝜙(1-p)(1-𝜙p)
Ignore the first 1. We only have 2 0s. So, we have (1-𝜙) + 𝜙(1-p)(1-𝜙p). The animal was never recaptured, we now have 2 possibilities (last row).
note: We condition on the animal being captured in time 1. So we don’t estimate p or 𝜙 for time 1. We ONLY consider animals with a 1 for the first period
Assumptions of the CJS
Homogeneity of capture and survival probabilities for the marked animals within each sample occasion and group
Individual heterogeneity can also be modeled similar to model Mh but is pretty complex
Instantaneous recapture and release of animals
Keep trapping short relative to the interval between trap occasions
All emigration from the study area is permanent (no temporary emigration)
Issues of Jolly-Seber model
Estimating the proportion of animals marked on each occasion is tricky and estimates of N may be biased as a result
As a result, estimating recruitment is difficult and models rarely converge
Estimates of abundance and recruitment are not robust to heterogeneity in capture probabilities
Difference between CJS and JS models
CJS models
Estimate survival and capture probability
Condition on marked individuals
JS models
Estimate survival, capture probability, abundance and recruitment
Must estimate the proportion of individuals in the population at each time step
Tricky
No heterogeneity in capture probabilities allowed!