How mirRNA silences translation
1/17
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
18 Terms
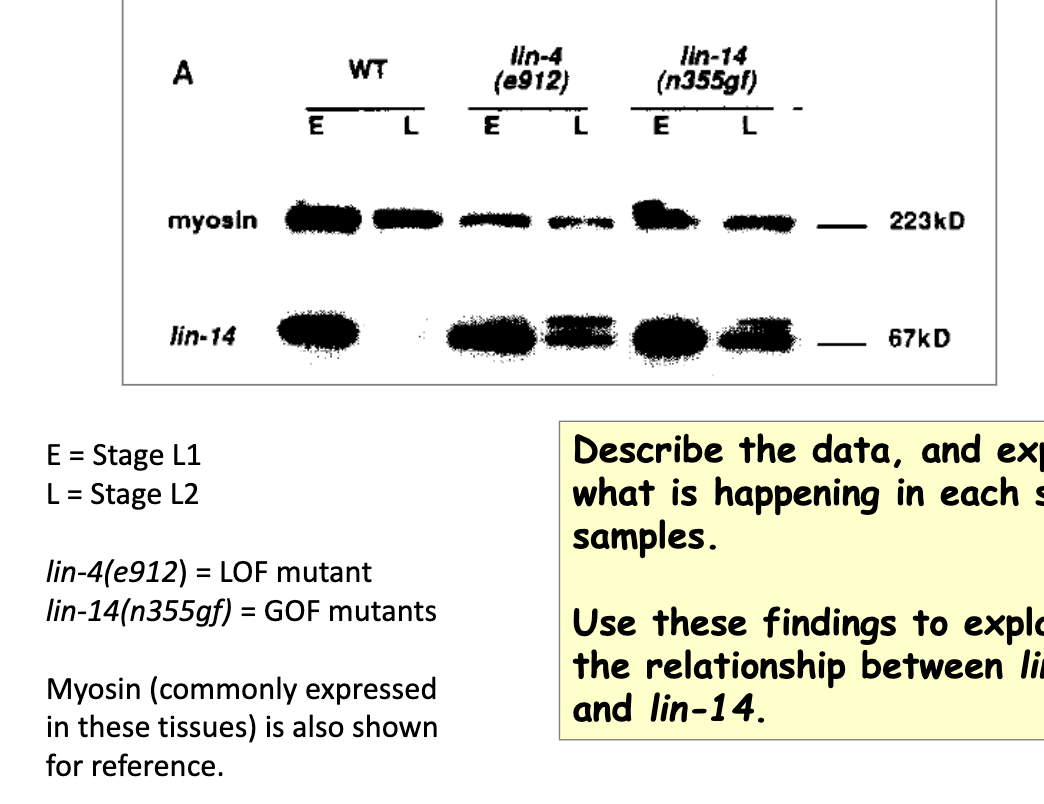
Western blot showing relative expression levels of Lin-14 protein
in WT: Lin-14 protein expressed in L1 but not in L2
in lin-4 lof mutant: Lin-14 still expressed in L2 → suggests that WT lin-4 responsible for preventing expression of Lin-14
in lin-14 gof mutant: Lin-14 still expressed in L2 → no longer responds to lin-4
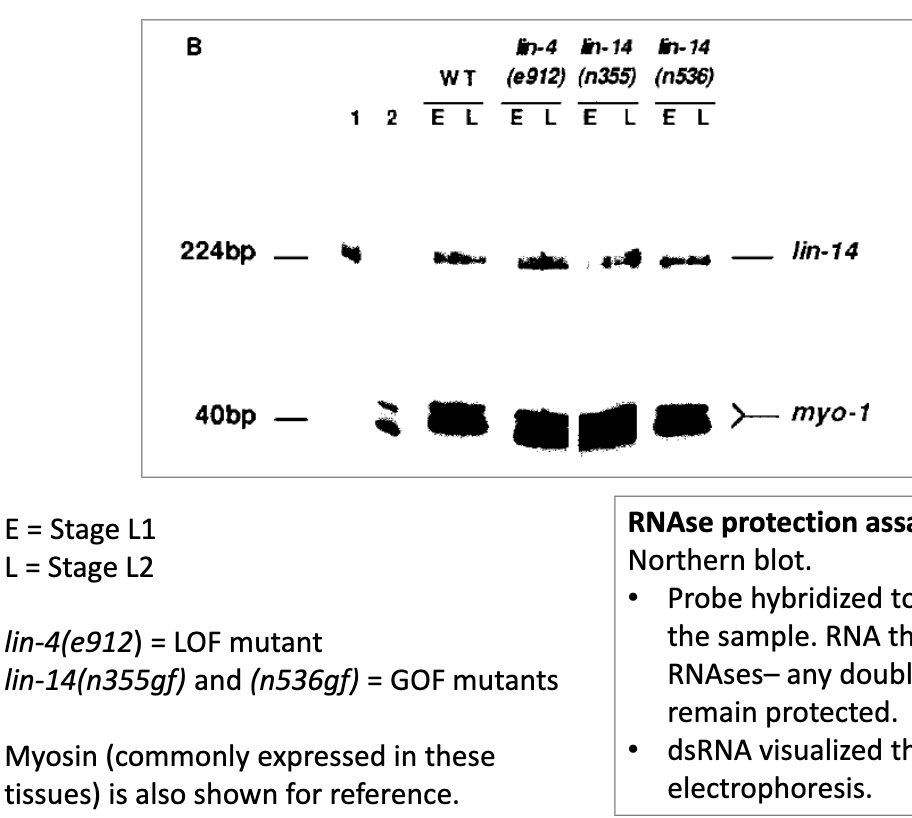
RNAse protection assay
similar to a Northern blot
probe hybridized to RNA targets in the sample.
RNA then exposed to RNAses → any double-stranded RNAs remain protected
dsRNA visualized thru gel electrophoresis
RNAse Protection Assay showing relative expression levels of lin-4 transcript
lin-14 transcript is expressed in both stages L1 and L2, in the WT and the mutant
lin-14 transcript is expressed in L1 and L2. It is only translated in L2 in the lin-4 lof mutant (in absense of lin-4), or in the lin-14 gof mutants (no longer responding to lin-4)
therefore inhibition occurs at translational level, not transcriptional level
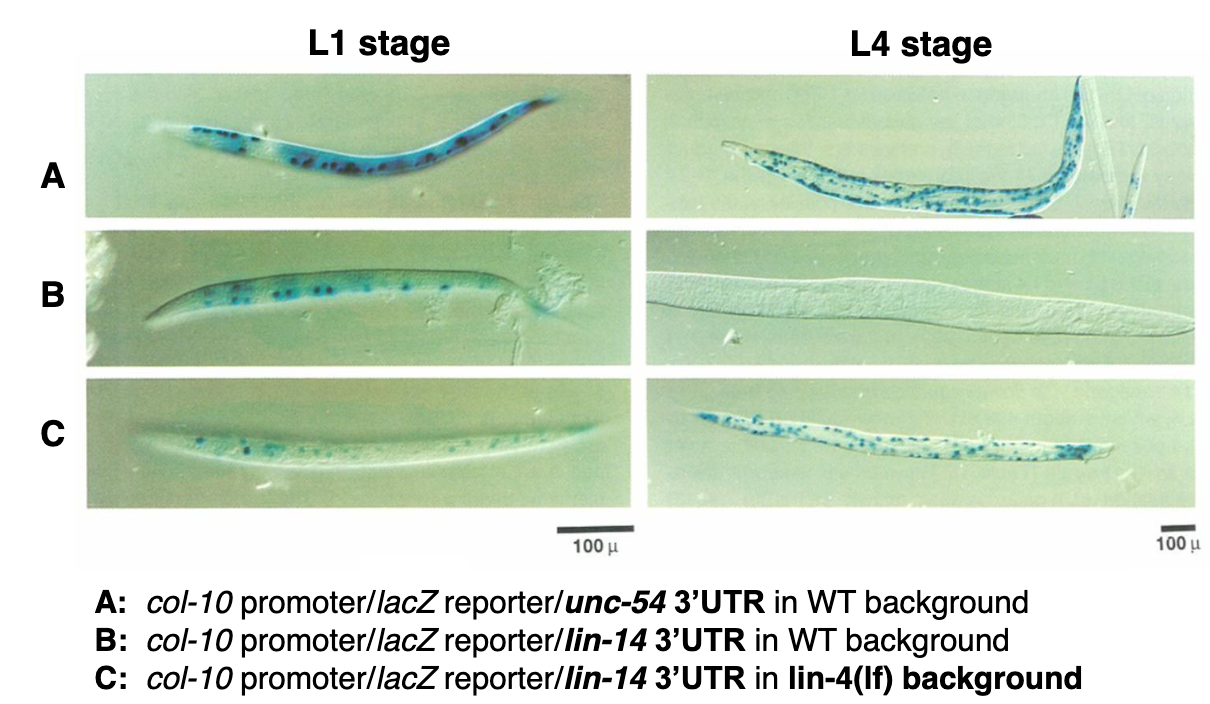
Pinpointing the regulatory elements within the lin-14 mRNA
What elements within lin-14 mRNA are required for regulation by lin-4?
Creation of transgenic C. elegans expressing lacZ reporter genes
col-10 collagen promoter: active in the same cells that accumulate lin-14. lacZ gene should be transcribed in those cells
test: lacZ gene fused to the lin-14 3’UTR: the 3’UTR is suspected to contain regulatory elements for lin-14 translation
miRNAs typically bind to 3’UTRs to repress translation
if lin-4 regulates thru the 3’UTR, then lin-4 should bind this region → reduced lacZ expression
control: lacZ gene fused to the unc-54 3’UTR (from a myosin gene): a control UTR that is known to be downregulated (but not completely suppressed) between stage L1 and L4
distinguishes between specific regulation by lin-4 and general developmental changes
Data from above slide
A is the control
B shows typical regulation of lin 14
What is the purpose of transforming C. elegans with the lacZ/unc-54 3’ UTR?
To show that any repression you see is specific to the lin-14 3′ UTR, not just a general effect
unc-54 3’UTR comes from a diff gene and is not a target of lin-4
it does show some normal developmental downregulation, but not due to lin-4
in the exp, if you see:
lacZ + lin-14 3′ UTR → NOT blue (repressed)
lacZ + unc-54 3′ UTR → still blue
then you can conclude the repression is due to specific sequence in the lin-14 3’UTR
without this control, you wouldn’t know if the drop in blue is due to lin-4 or just normal developmental changes (L1→L4)
Which panel(s) demonstrate the typical regulation of lin-14?
normally, in WT worms, lin-4 is present and represses lin-14 via its 3’UTR over development
so you expect Early (L1) = expression, later = repression
therefore, panel B demonstrates typical lin-14 regulation
Which panel(s) confirms how lin-14 is regulated?
B + C
B: shows repression happens with lin-14 3′ UTR
C: shows repression is lost without lin-4
lin-4 regulates lin-14 through the lin-14 3′ UTR
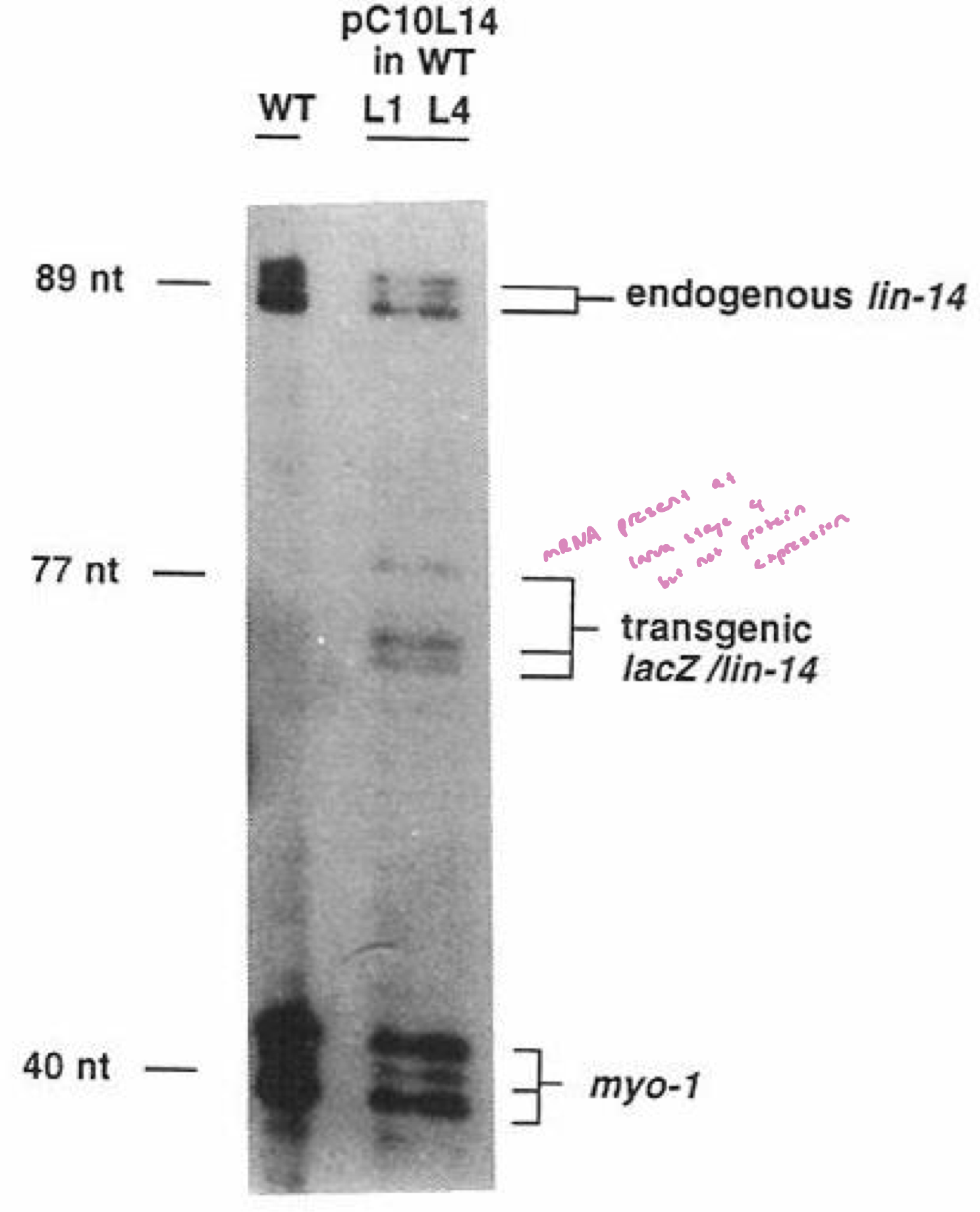
Is lin-14 regulation happening at the RNA level or the translation level?
RNA transcript for endogenous lin-14 and the lacZ transgenes are present in both L1 and L4
thus, inhibition of the lin-14 3’UTR does not depend on the loss of the lin-14 transcript
ie. lin-14 transcript is produced thru all larval stages, but Lin-14 protein is only translated in L1. Therefore, translation of the lin-14 mRNA must be suppresed after L1 (by Lin 4)
the lin-4 locus is very small and does not contain a recognizable ORF. However, its sequence is highly conserved b/w closely related species
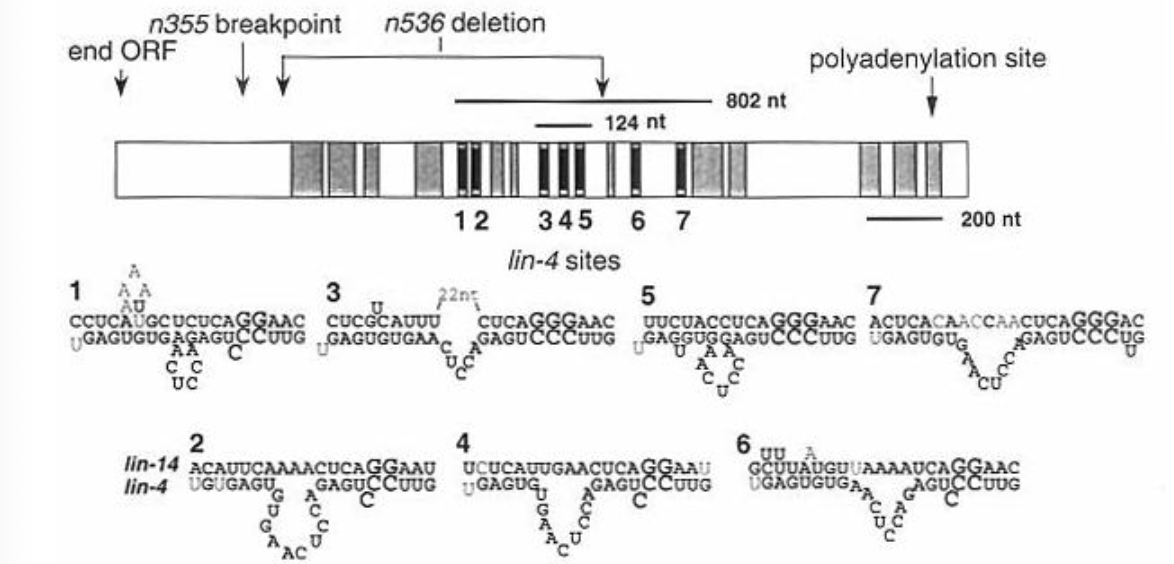
What is the mechanism of lin-4 reglulating Lin-14 protein expression?
lin-4 transcript hybridizes with the lin-14 3’UTR to suppress lin-14 translation
the lin-4 sequence is complementary to sequences within the lin-14 3’UTR
the lin-14 GOF mutations delete the 3’UTR, explaining why lin-4 is no longer able to suppress Lin-14 protein expression in those strains (lin-4 binding sites are gone)
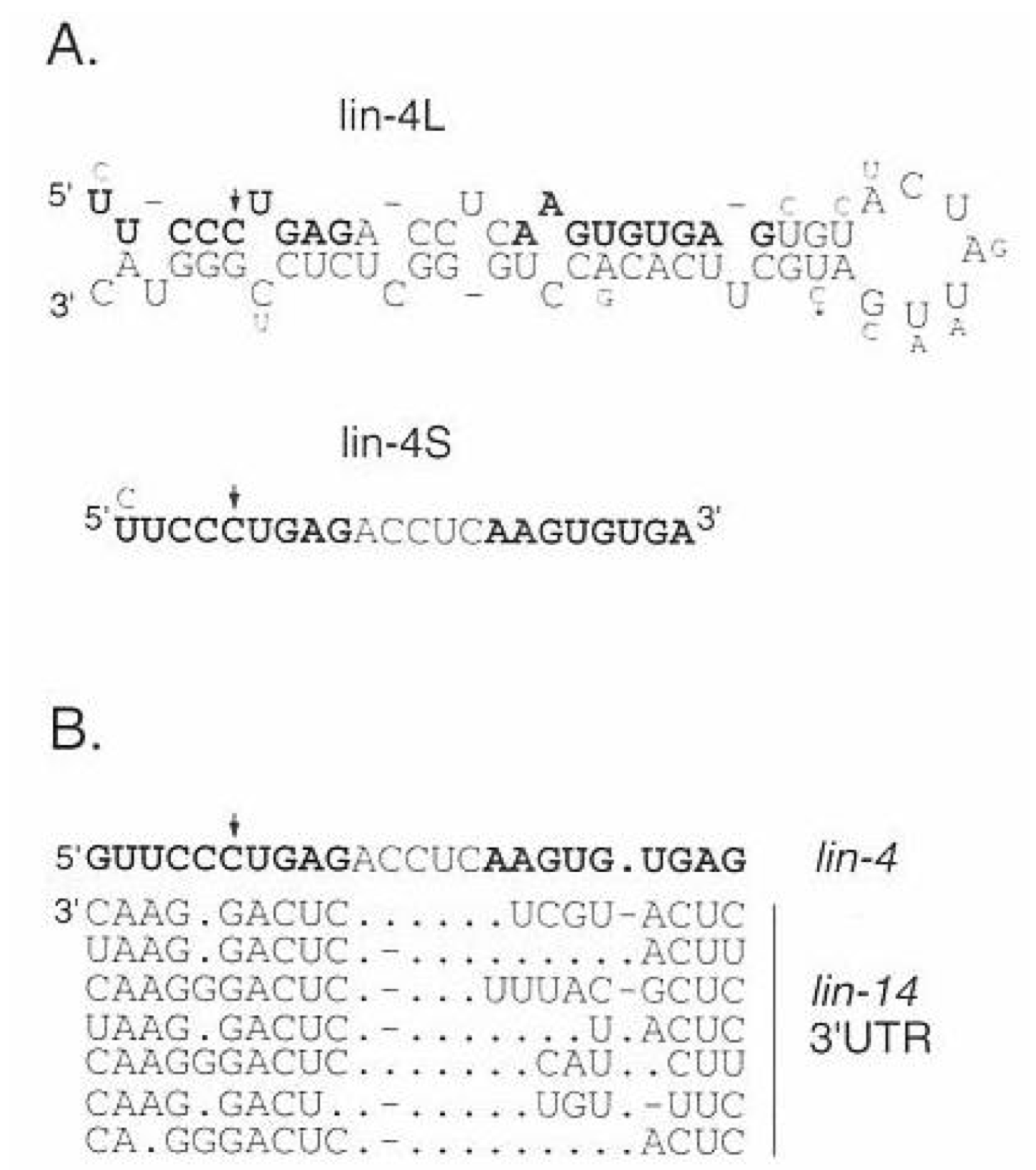
Lin-4 Small vs Large
the smaller lin-4 transcript is the one that is able to hybridize to the 7 regions within the lin-14 3’UTR
the larger lin-4 transcript forms a hairpin structure → it is processed further to make lin-4s
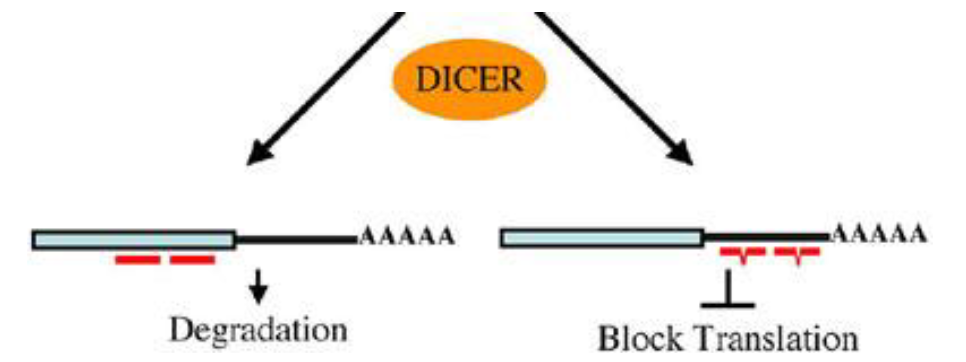
microRNA
specialized RNAs that prevent expression of specific genes thru complementary base pairing (binding is imperfect and usually occurs in the 3’UTR)
first miRNAs (lin-4 and let-7) identified in C. elegans
lin-4 codes for a microRNA
small (21-30 nt) RNAs (lin-4 is 22 nts)
micro-RNAs (miRNAs) and small interfering RNAs (siRNAs)
miRNAS regulate gene expression post transcriptionally (after mRNA is already made)
can reduce mRNA stability and block translation
RNAi (RNA Interference)
A mechanism by which small RNAs suppress translation of mRNAs. 2 types:
siRNA (small interfering RNA): binds to target w/ perfect complementarity to trigger degradation
miRNA: binds to targets w/ imperfect complementarity to block translation without degradation (classic miRNA pathway)
there may be some crossover in their function
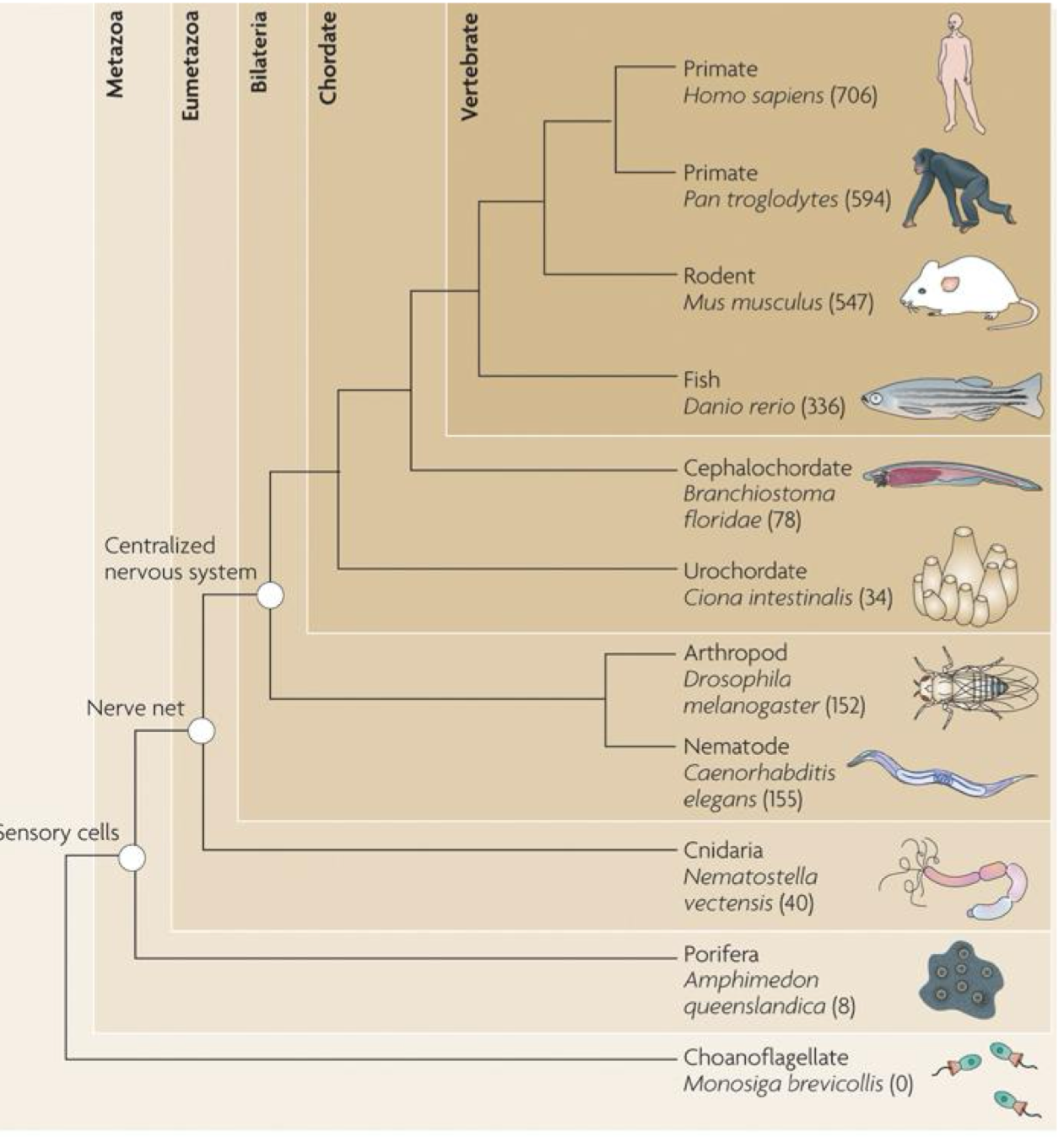
Shared microRNAs perspecies
the identification in C. elegans was the big breakthru, it has homologs in other species (including humans)
but we don’t know yet how many of the targets are shared
over time, we are discovering more and more miRNA genes
Mammalian miRNAs implicated in a wide range of developmental processes
formation of skeleton, brain, teeth, eyes, neurons, muscle, heart, lungs, kidneys, vasculature, liver, pancreas, intestine, skin, fat, breast, ovaries, testes, placenta, thymus
development of hematopoeitic lineages (blood cells; immune cells)
cell functions such as axon sprouting and synapse formation, mitotic spindle orientation
physilogical functions such as blood pressure, lipid metabolism, insulin production, pituitary function, bone resorption, growth of embryos
miRNAs add another layer of regulation, allowing further complexity and fine detailing of gene expression
Primary transcripts contain miRNA
most miRNAs are transribed by RNA pol II, and are thus controlled by TFs that work w/ this pol
most miRNAs are transcribed from noncoding DNA regions that generate short dsRNA hairpins
some miRNA coding sequences lie within introns, some lie outside of protein-coding genes, a few lie within exons
miRNA genes have been subject to duplication and diergence, generating clusters and families of related miRNAs
miRNA families can be grouped together based on their extended seed regions (nt 2-8, the region involved in targeting). Family members often have partially redundant functions
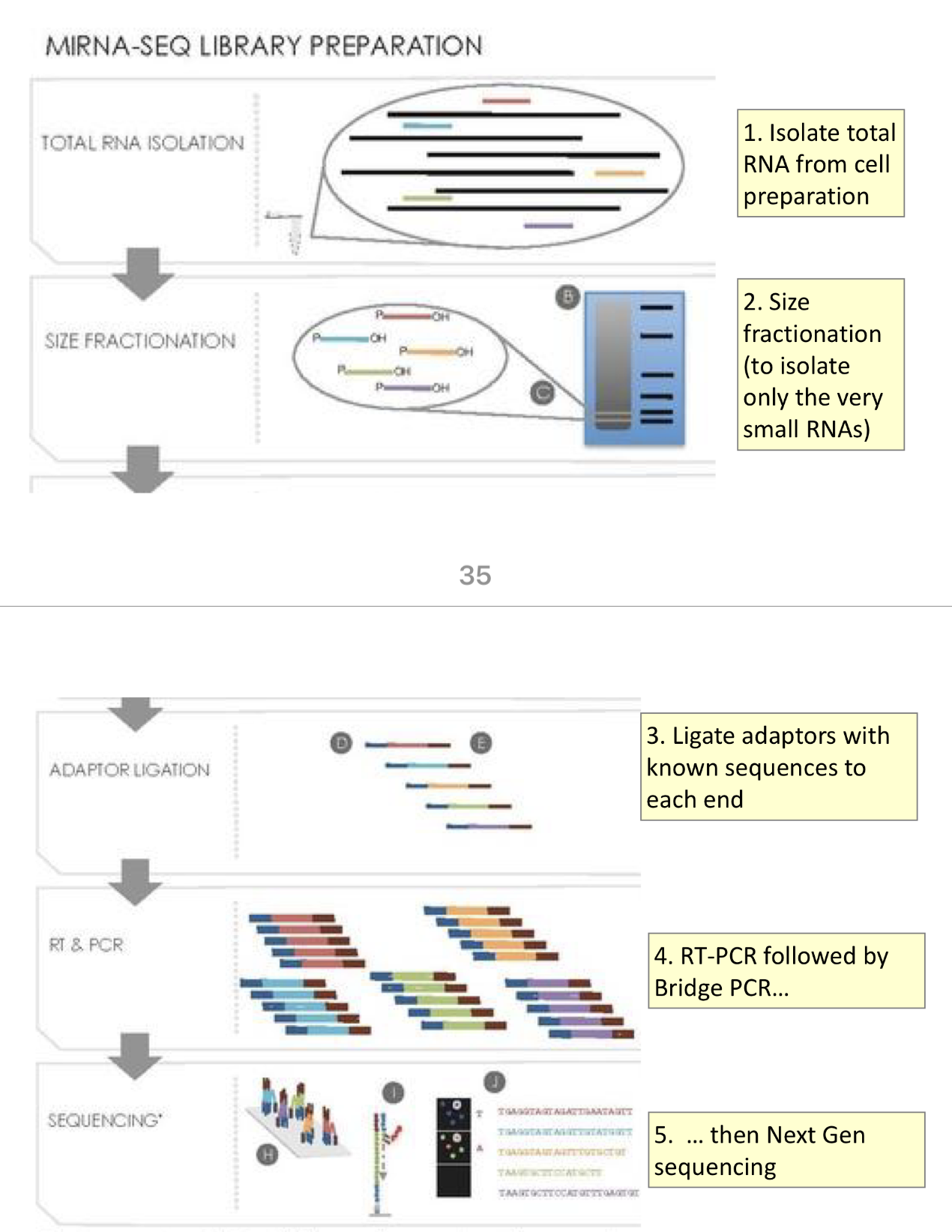
Whole transcriptome approach to identifying and sequencing miRNAs
1) isolate total RNA from cell preparation
2) size fractionation (to isoalte only the very small RNAs)
3) ligate adaptors with known sequences to each end
4) RT-PCR followed by Bridge-PCR
5) Next gen sequencing