mmg 2040 lecture 27 - recombinant protein expression
1/13
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
14 Terms
Define: transcriptome, proteome, translatome, isoelectric focusing
transcriptome: Set of all RNA molecules, including mRNA, rRNA, tRNA, and other non-coding RNA, produced in one or a population of cells
proteome: Total set of proteins encoded by a genome or the total protein complement of an organism
translatome: total set of proteins that have actually been translated and are present in a cell under any particular set of conditions
isoelectric focusing: Technique for separating proteins according to their charge by means of electrophoresis through a pH gradient
high-resolution electrophoretic technique that separates proteins or amphoteric molecules based on their unique isoelectric point —> pH where they carry no net charge
Molecules migrate through a stable pH gradient under an electric field until they reach a zone where the pH equals their pi, stopping because they become neutral
Principle of Isoelectric Focusing
𝒑𝑰 (Isoelectric Point): The specific pH value where a molecule's net charge is zero.
pH Gradient: IEF utilizes a gel or medium with an immobilized, established pH gradient (acidic to basic).
Migration: When an electric field is applied, molecules with a positive net charge move toward the cathode, and negatively charged molecules move toward the anode.
Focusing: As molecules migrate, they reach the pH corresponding to their 𝑝𝐼, lose their net charge, and stop moving. This "focuses" them into sharp bands
Provide reasons for the discrepancy between: proteome and translatome, proteome and transcriptome
Some RNA molecules are non-coding
Alternative splicing →multiple protein products
Levels of mRNA may not correlate with protein levels due to differential rates of mRNA translation or degradation
Regulation of proteins by addition or removal of acetyl, phosphate, AMP, ADP-ribose, or other groups
Regulation of proteins by chemical modification of amino acid residues
Modification by proteolytic cleavage or addition of sugar or lipid residues to give glycoproteins or lipoproteins
Proteins themselves may be degraded and vary greatly in stability
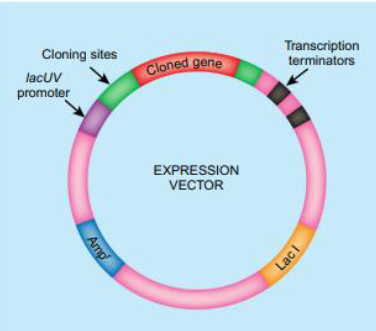
Identify the essential components of an expression vector used for eukaryotic expression systems
Strong promoter (e.g., T7)
Ribosome binding site
Selectable marker
Tag sequence (His-tag, GFP)
PolyA site
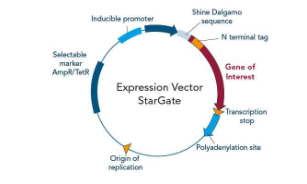
Describe the basics steps in recombinant protein expression
Clone gene into expression vector
Expression Vectors
Strong promoter (e.g., T7)
Ribosome binding site
Selectable marker
Tag sequence (His-tag, GFP)
PolyA site
Introduce into host cell
Bacterial systems - Heat-shock transformation + Electroporation
Yeast systems - Heat-shock transformation + Electroporation
Mammalian systems - Calcium phosphate transfection + Lipid-mediated transfection + Viral Transduction
Induce expression (using lacuv, t7, tet-on/off)
Host produces protein
Control timing of expression
Prevents toxicity
Harvest cells - General Purification Strategy
Lyse cells
Separate proteins
Isolate target protein
Validate purity
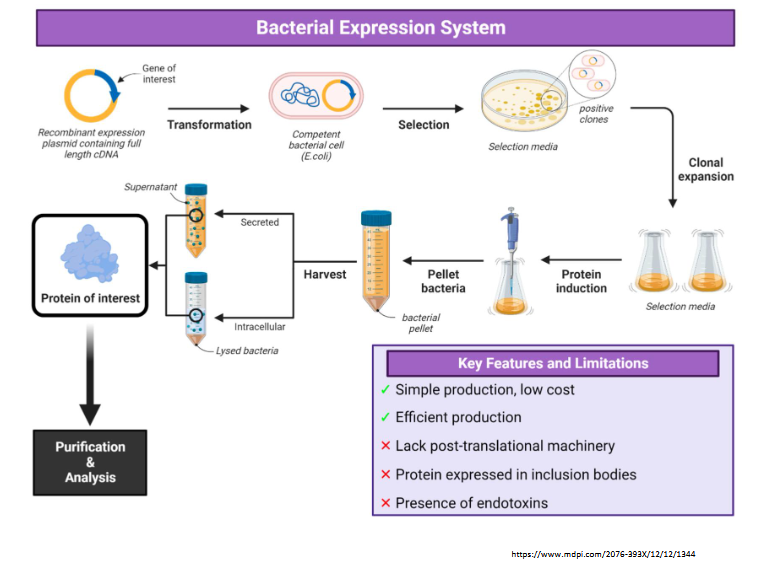
Compare pros/cons of bacterial expression systems
bacterial: simple production, low cost, efficient production
lacks post-translational machinery
protein expressed in inclusion bodies
presence of endotoxins
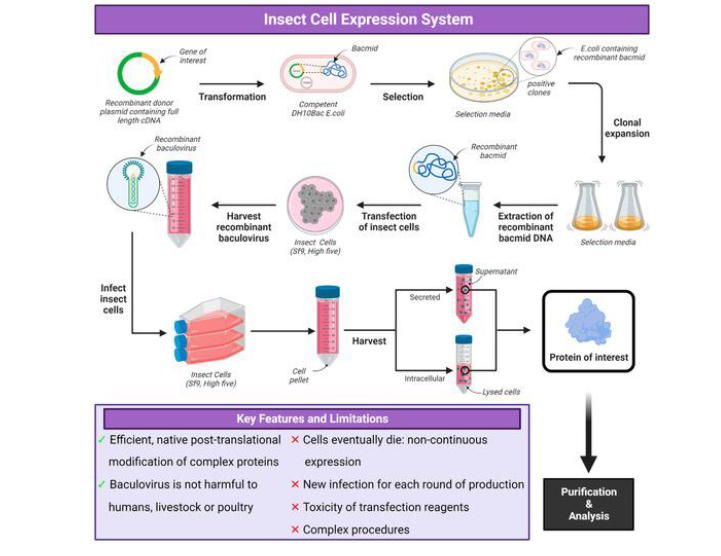
Compare pros/cons of insect expression systems
insect: Efficient, native post-translational modification of complex proteins, baculovirus is not harmful to humans, livestock, or poultry
cells eventually die —> non-continuous expression
new infection for each new round of production
toxicity of transfection agents
complex procedures
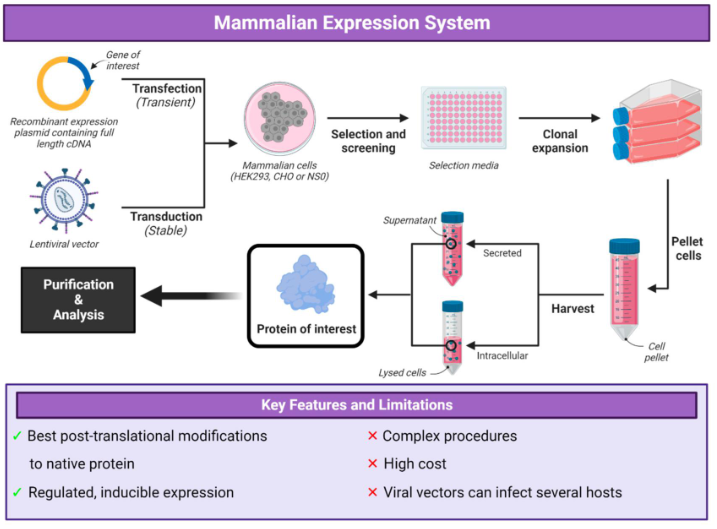
Compare pros/cons of mammalian expression systems
mammalian: best post-translational modifications to native protein, regulation + inducible expression
complex procedures, high cost, viral vectors can infect several hosts
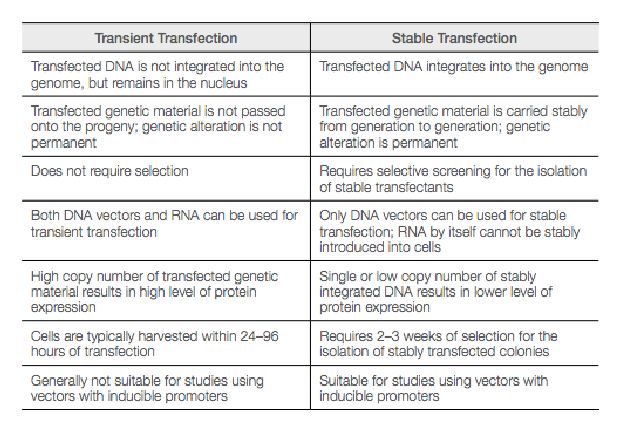
Compare/contrast transient and stable transfections
transient: not integrated in genome but remains in nucleus, not passed onto progeny/genetic alteration is not permanent, does not require selection, high copy number of material results in high level of protein expression, generally not suitable for studies using inducible vectors
both DNA/RNA vectors can be used
harvested 24-96 hrs after transfection
stable: integrated into genome carried stably from gen. to gen./genetic alteration is permanent, requires selective screening for stable transfectants, single/low copy number of integrated
stable vs. transient transfection when entering cells
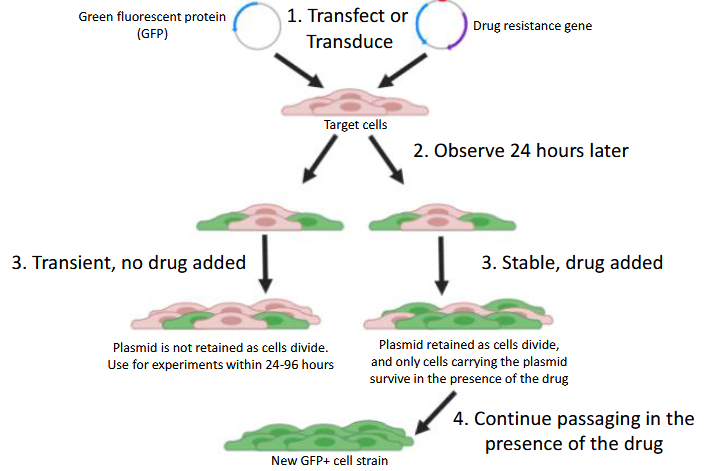
Define transfection and transduction relative to mammalian expression vectors
transfection: Introduction of vectors, without using viruses, into mammalian cells
Transduction: Introduction of vectors into mammalian cells using viral-mediated delivery
Design a basic mammalian expression strategy
Bacterial or mammalian system?
Transient or stable?
Inducible or constitutive?
Viral or non-viral delivery?
a cloning experiment - it is important it is IN FRAME
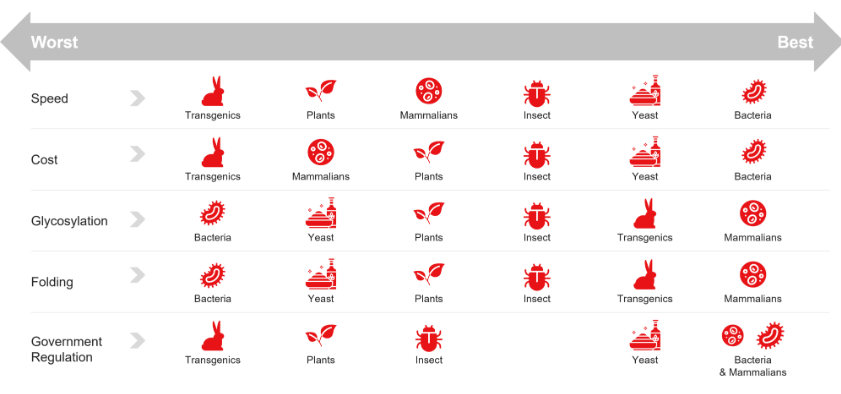
Describe the following inducible promoter systems: lacUV
The lacUV5 promoter is a mutated promoter from the Escherichia coli lac operon very similar to the classical lac promoter,
it requires no additional activators and it drives high levels of gene expression
The lacUV5 promoter sequence conforms more closely to the consensus sequence recognized by bacterial sigma factors than the traditional lac promoter does
Due to this, lacUV5 recruits RNA Polymerase more effectively
leads to higher transcription of target genes
lacUV5 works independently of activator proteins or other cis-regulatory elements
While no activators are required, lacUV5 promoter expression can be regulated by the LacI repressor and can be induced with IPTG, which is an effective inducer of protein expression
Due to this control, the lacUV5 promoter is commonly found on expression plasmids
is used when controllable but high levels of a product are desired
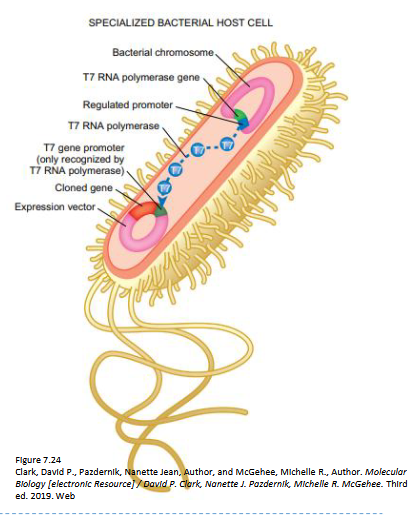
Describe the following inducible promoter systems: T7
T7 expression vectors (commonly pET vectors) enable high-level protein production in E. coli by using T7 RNA polymerase to specifically transcribe a target gene
system is induced by IPTG, which triggers the expression of T7 RNA polymerase from the host genome
recognizes a T7 promoter on the plasmid to drive massive target gene expression
Vector Construction: The target gene is inserted into a plasmid (e.g., pET) downstream of a strong T7 promoter.
Host Cell: The vector is introduced into special E. coli strains that have the gene for T7 RNA polymerase integrated into their chromosome.
Induction: T7 RNA polymerase expression is controlled by an inducible promoter (typically lacUV5). The addition of IPTG relieves repression, producing T7 RNA polymerase.
Transcription: The produced T7 RNA polymerase specifically transcribes the target gene, leading to high-level protein overexpression.
Tight Regulation: To prevent basal expression (leakiness) of potentially toxic proteins, T7 lysozyme (a natural inhibitor) can be expressed, or the system can use Lac repressor binding
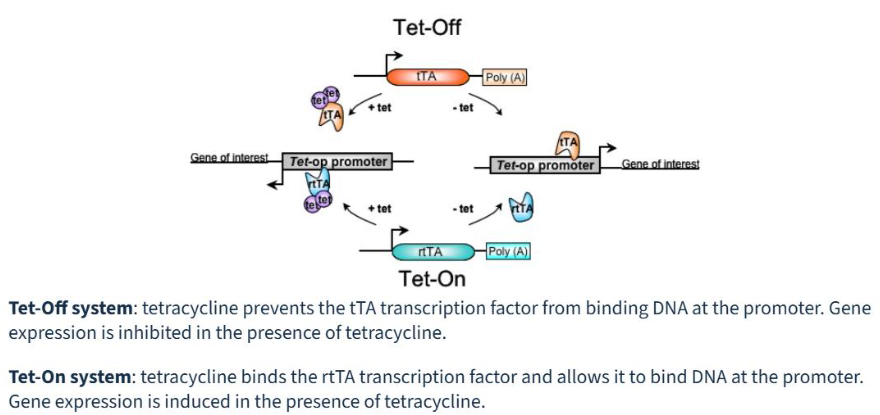
Describe the following inducible promoter systems: Tet-on/Tet-off
Tet-Off and Tet-On systems are inducible gene expression tools that use tetracycline or its derivative doxycycline (Dox) to control gene activity.
both relying on a transactivator protein binding to a Tetracycline Response Element (TRE) promoter
Tet-Off turns expression off when Dox is added
requires tTA (Tet-Off): Tetracycline-controlled Transactivator
The tTA protein binds to the TRE promoter, activating transcription of the target gene
Dox binds to the tTA protein, preventing it from binding to the TRE. Expression turns OFF
Useful for long-term studies where expression is usually needed, then turned off
Tet-On turns expression on when Dox is added
requires rtTA (Tet-On): Reverse Tetracycline-controlled Transactivator
The rtTA protein cannot bind the TRE. Expression is OFF
Dox binds to the rtTA protein, inducing a conformational change that allows it to bind the TRE. Expression turns ON
Often preferred for rapid response to Dox induction