BIO 181L Study Guide Final Exam (GCU - Youngblood)
1/65
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
66 Terms
LAB 7: catalyst
substance that speeds up the rate of a chemical reaction
LAB 7: Enzmes
Proteins which serve as catalysts for certain chemical reactions
LAB 7: energy of activation
The amount of energy that reactants must absorb before a chemical reaction will start
LAB 7: Active site of an enzyme
the region of an enzyme that attaches to a substrate
LAB 7: How do enzymes catalyze reactions?
by lowering the activation energy required for the reaction to occur
LAB 7: What are the seven major properties of enzymes?
Most are proteins.
They are highly specific; most will only react with a certain substrate.
They form an enzyme-substrate complex:
E +S↔ES →P +E
They do not affect the direction of a reaction.
They lower the energy of activation of a reaction.
They are not consumed in the reaction.
They are often highly regulated.
LAB 7: How do inhibitors work?
by preventing reactants from coming together because they have a similar shape to the substrate molecule.
Therefore less substrate molecules can bind to the enzymes so the reaction rate is decreased.
LAB 7: How do different substrates affect the rate of enzyme reactions?
The presence or absence of a specific chemical functional group can decide how well the enzyme works for that reaction.
The ability of the enzyme to work well with certain substrates and not well with other similar substrates is called specificity.
LAB 7: enzyme specificity
The ability of the enzyme to work well with certain substrates and not well with other similar substrates
LAB 7: How does substrate concentration affect the rate of enzyme reactions?
Enzymes are not consumed in the reaction, so small amounts of an enzyme can work well on large amounts of substrate.
Increasing amounts of substrate will plateau the activity of a fixed amount of enzyme, taking longer to convert substrate into product.
Increasing amounts of enzyme will increase the rate of reaction.
LAB 7: How does pH affect the rate of enzyme reactions?
Changes in pH can affect the functional groups of certain amino acids, which change their charge or intermolecular interactions.
This can influence the structure of the enzyme.
Extreme pH levels can denature enzymes, rendering them nonfunctional.
LAB 7: How does temperature affect the rate of enzyme reactions?
Enzymes have optimal temperatures where they work the best.
Temperatures outside of their range can physically change their structure and disrupt their function.
LAB 7: How enzymes are regulated in the body?
By controlling their concentration
By controlling the availability of substrate
By controlling the activity of the enzyme
LAB 7: What is denaturation?
a structural change in a protein that results in a loss of its biological properties
LAB 7: What happens to an enzyme's activity when it is denatured?
structure = function
so when structure is denature, the enzyme looses its function
LAB 7: What conditions will cause an enzyme to become denatured?
change in pH
extreme (hot) temperature change
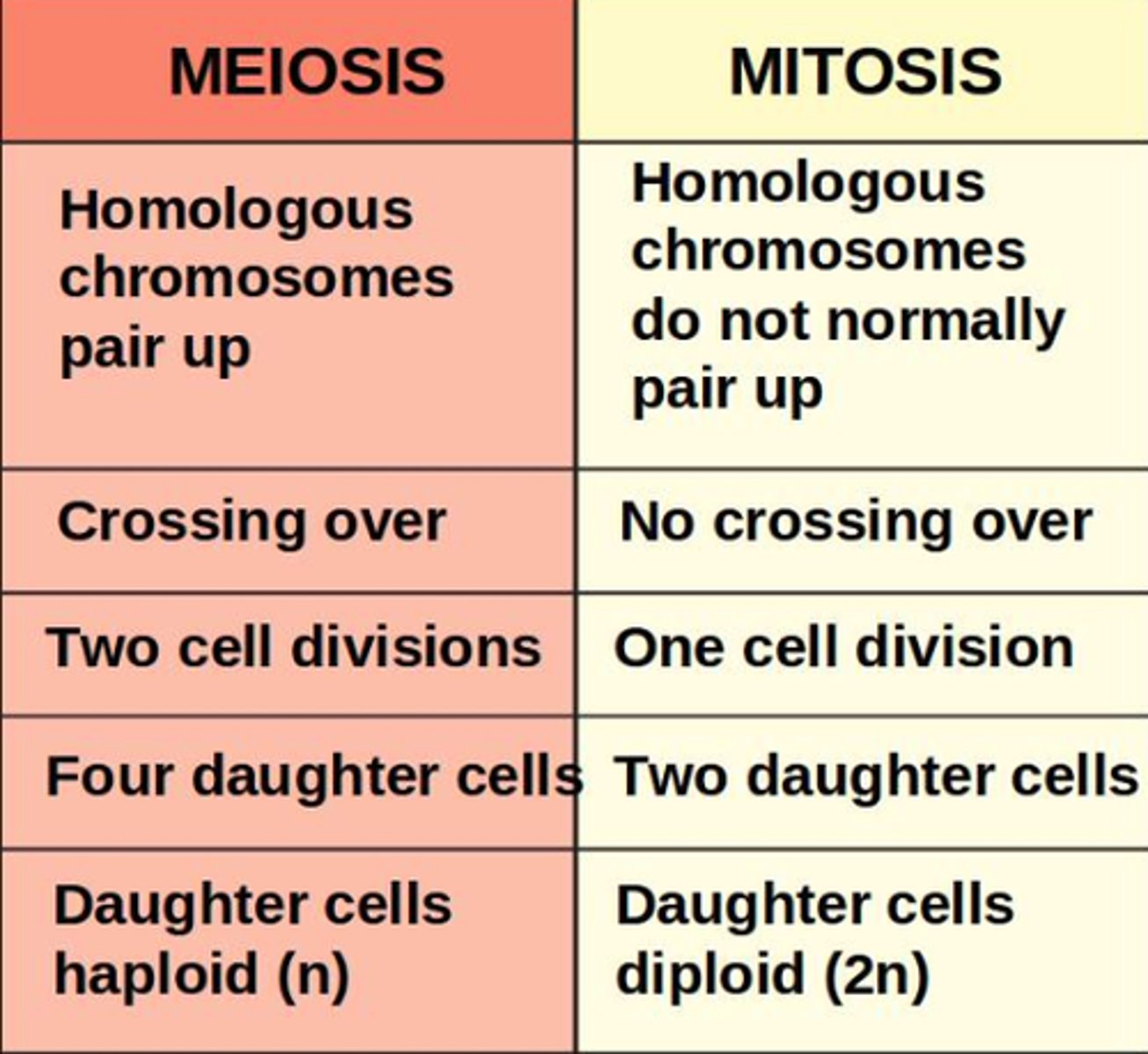
LAB 8: What are the main similarities and differences between mitosis and meiosis?

LAB 8: Prophase
first and longest phase of mitosis, during which the chromosomes become visible and the centrioles separate and take up positions on the opposite sides of the nucleus
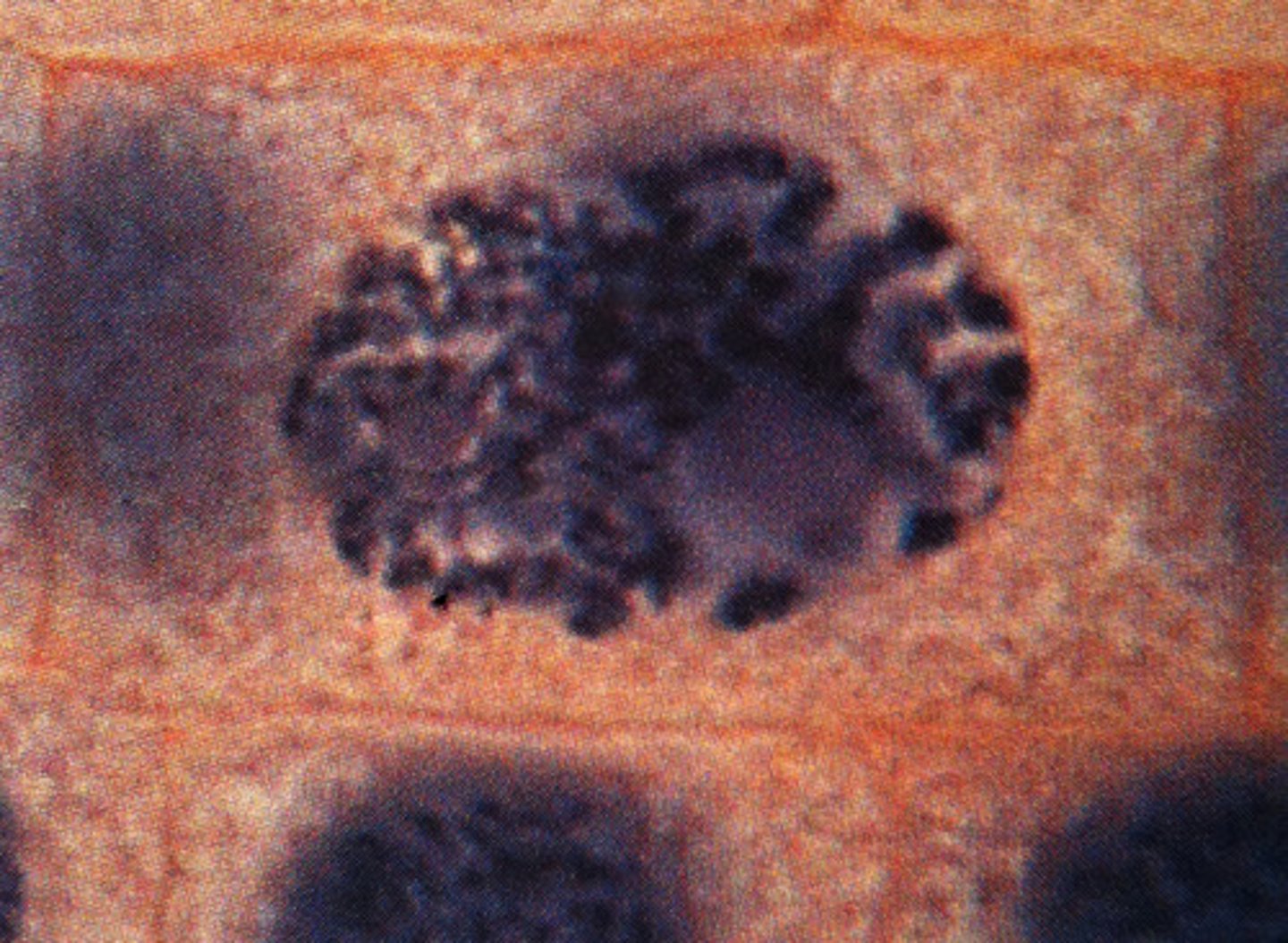
LAB 8: Metaphase
second phase of mitosis, during which the chromosomes line up across the center of the cell
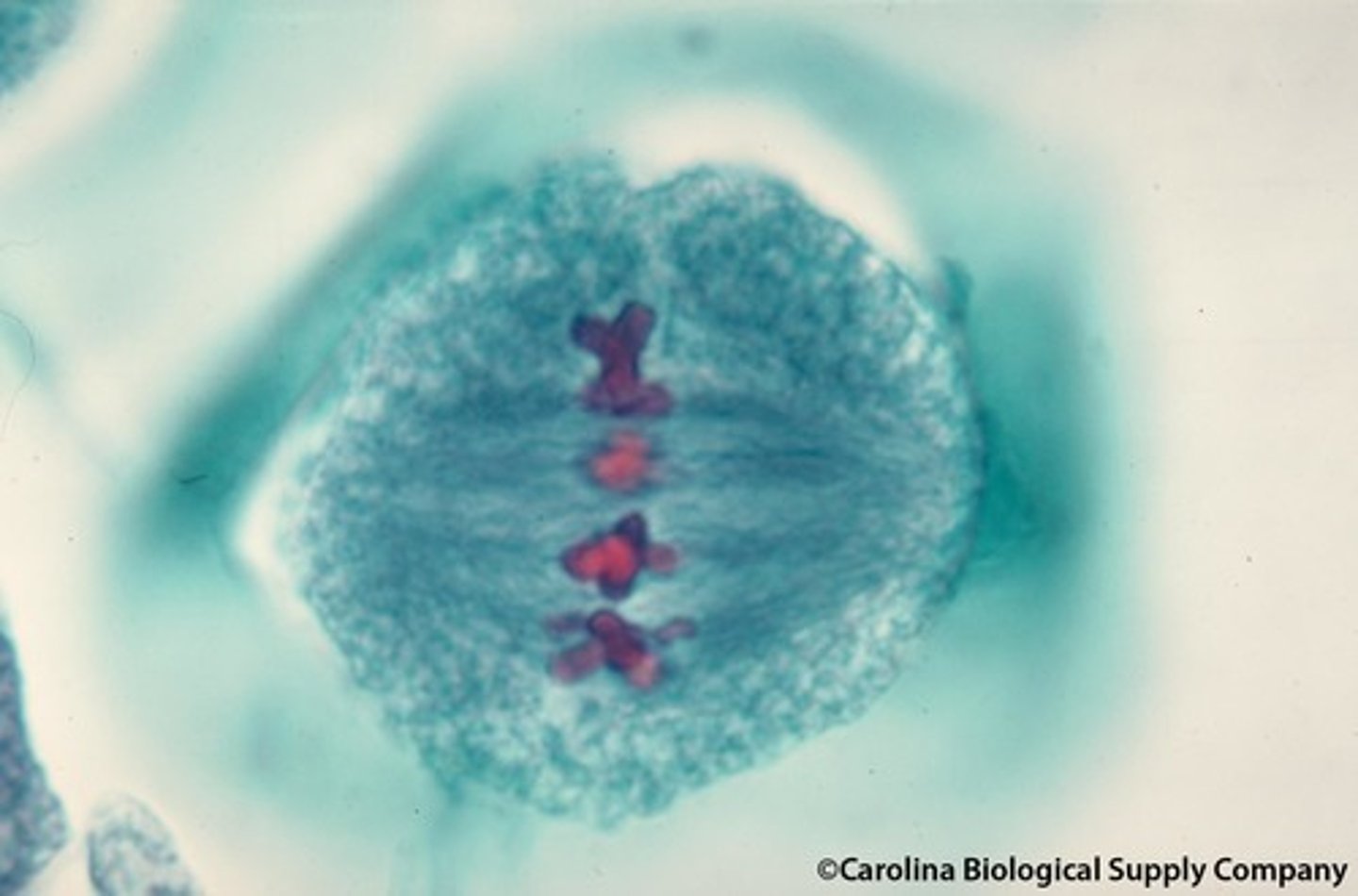
LAB 8: Anaphase
the third phase of mitosis, during which the chromosome pairs separate and move toward opposite poles
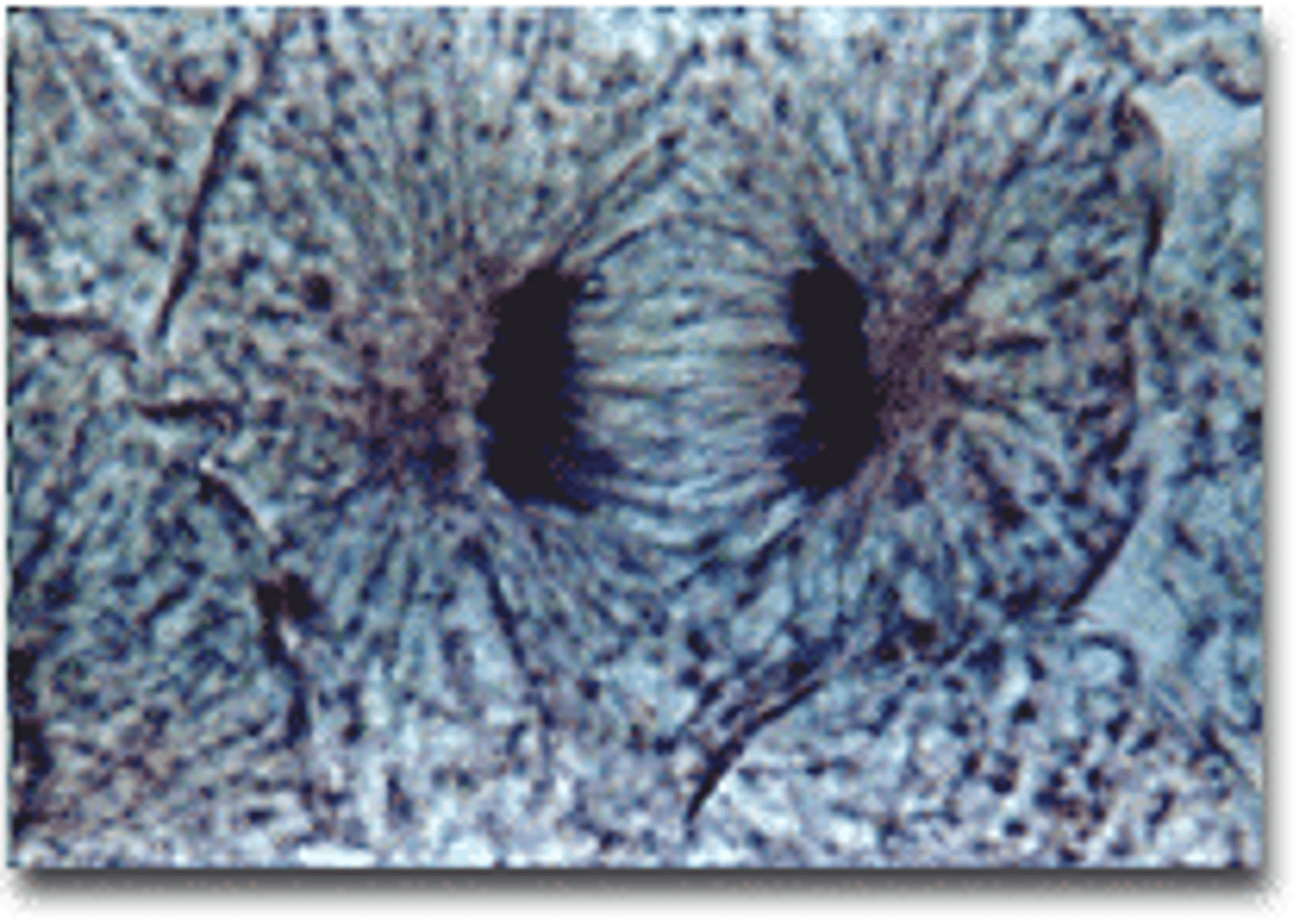
LAB 8: Telophase
the final phase of cell division, between anaphase and interphase, in which the chromatids or chromosomes move to opposite ends of the cell and two nuclei are formed.
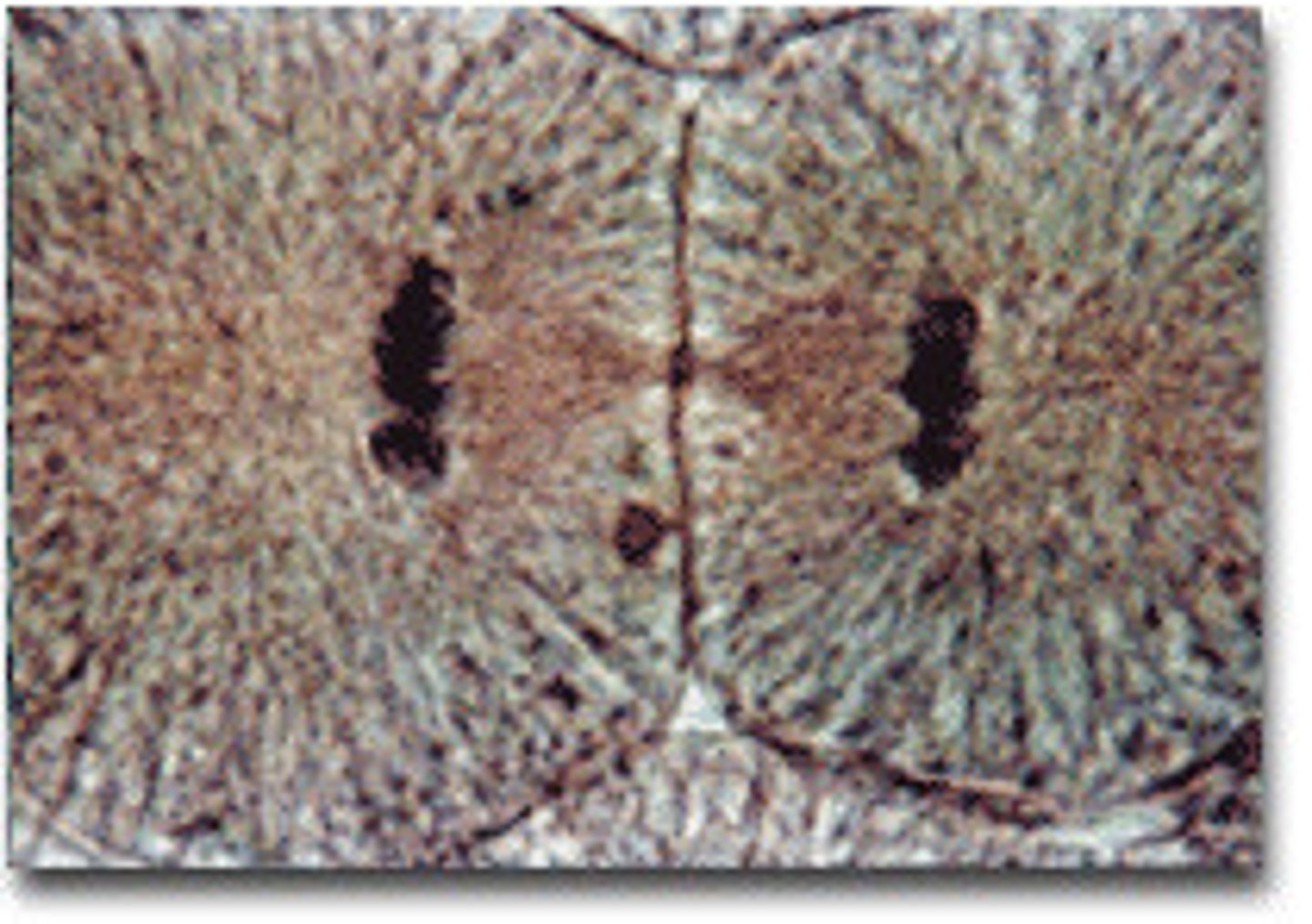
LAB 8: Prophase I (Meiosis)
homologous chromosomes pair up and form tetrads, crossing over occurs

LAB 8: Metaphase I (Meiosis)
homologous pairs (tetrads) align at the equatorial plane and each pair attaches to a separate spindle fiber at the kinetochore
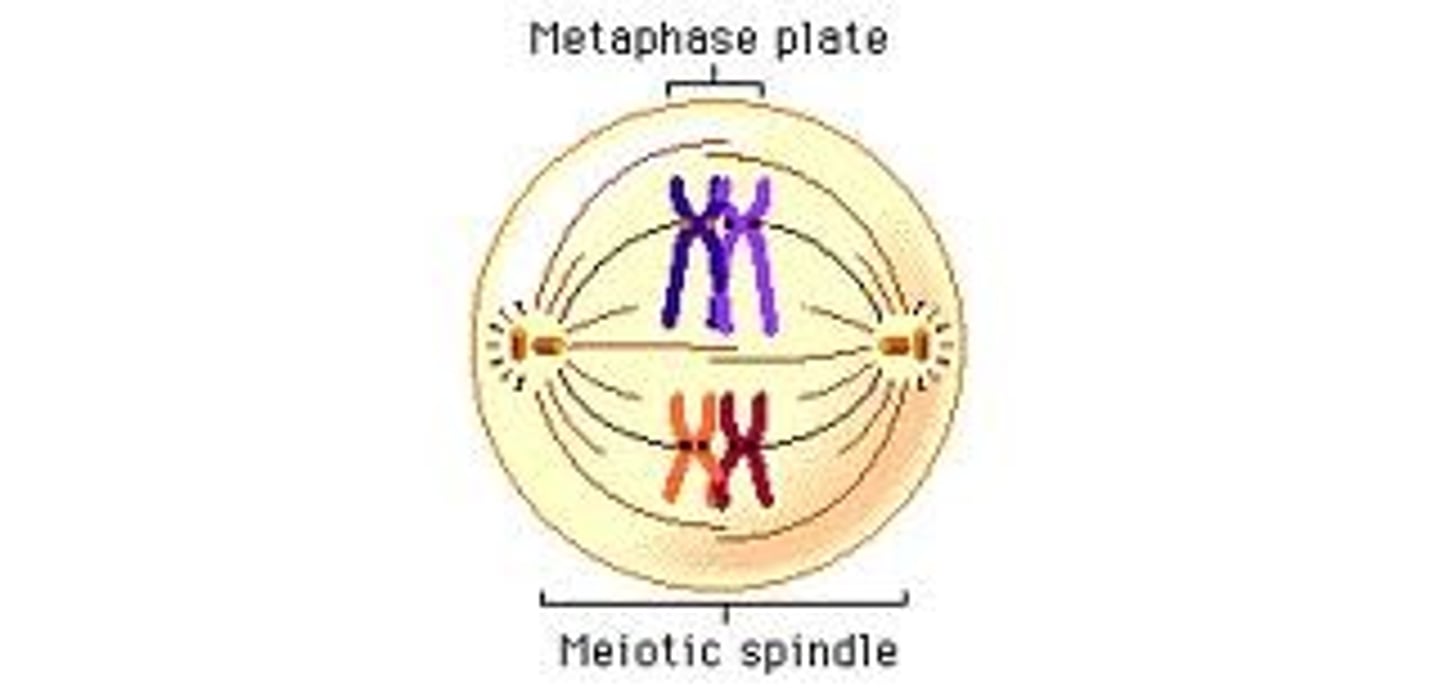
LAB 8: Anaphase I (Meiosis)
Tetrads split up and head to opposite poles
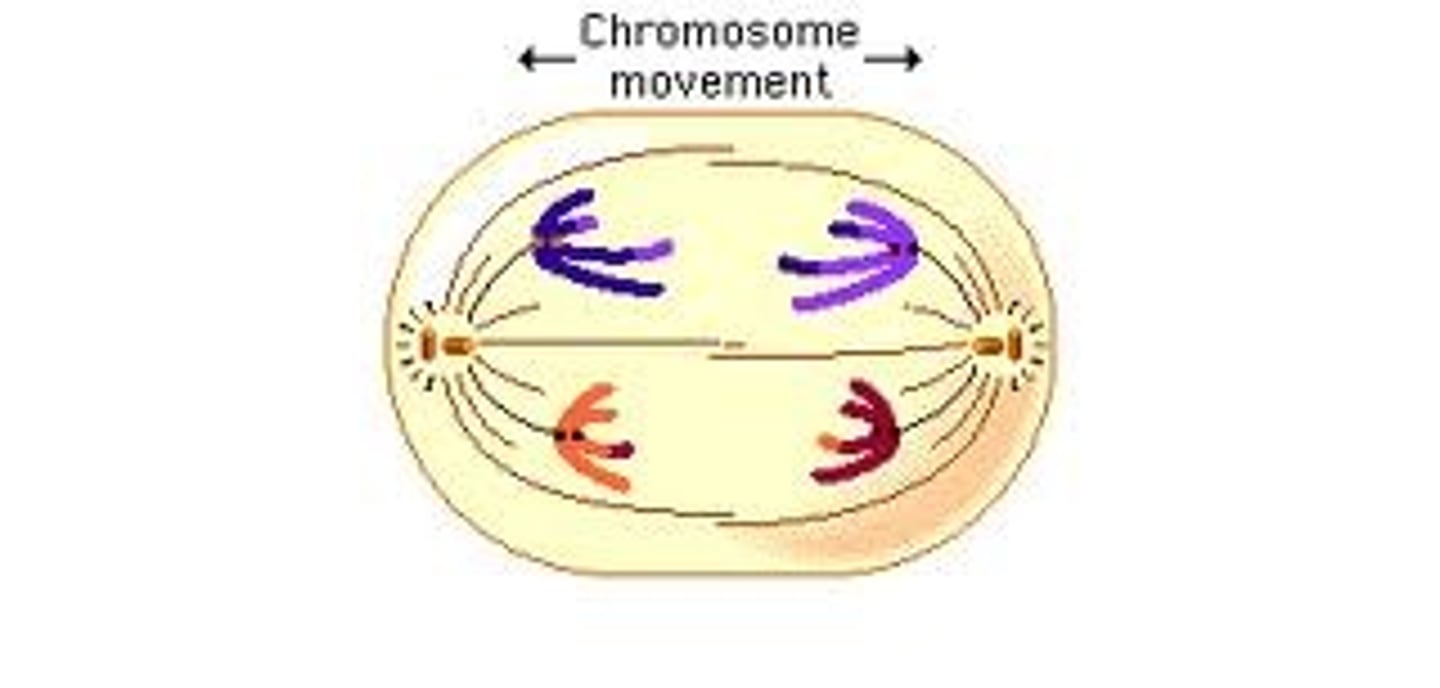
LAB 8: Telophase (mitosis)
1: Chromosomes lengthen and become indistinct chromatin
2: Nuclear envelope forms and nucleolus appears
3: spindle fibers disappear
4: 2 daughter nuclei present
5: Cytokinesis occurs (division of cytoplasm)
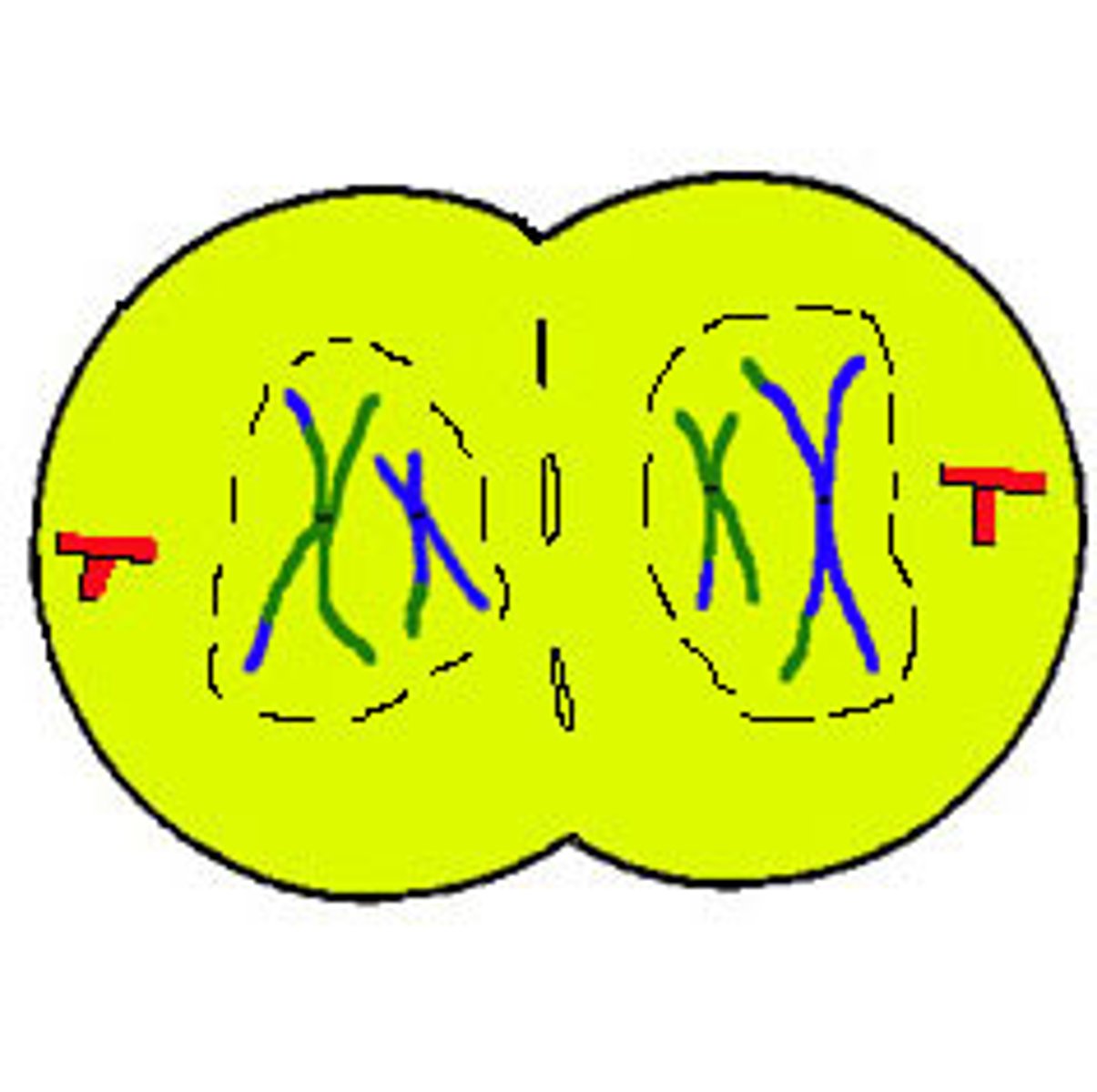
LAB 8: Prophase II
chromosomes condense - nuclear membranes start to dissolve again. Sister chromatids are again joined by a centromere. Spindles start to reform between centrosomes. (pic is wrong in this//disregard)

LAB 8: Metaphase II
Spindles are fully formed again and attach to the centromeres. The chromosomes line up down the middle of the spindles.
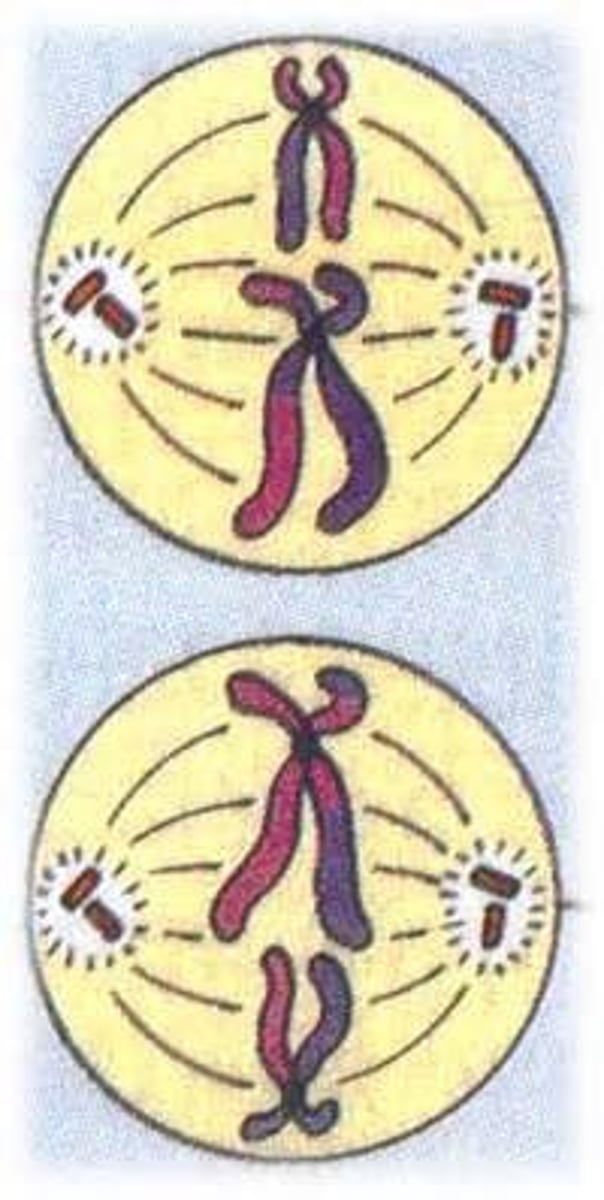
LAB 8: Anaphase II
The spindles pull the sister chromatids apart. Each goes towards a different pole.
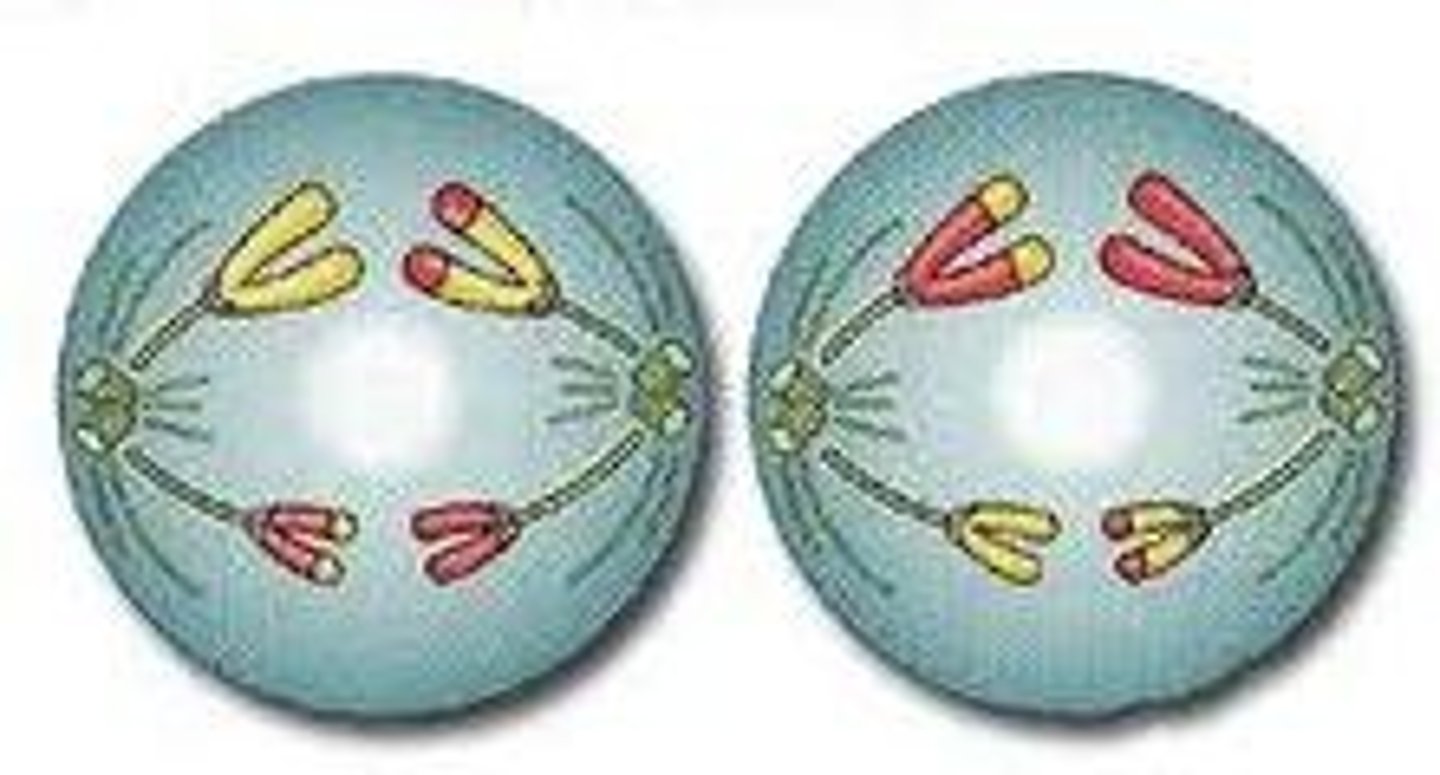

LAB 8: Telophase II
Nuclear membranes start to form around the chromosomes again. A cleavage forms. Cytokinesis occurs and the two diploid cells have now divided into 4 haploid cells. In males this equals 4 sperm. In females this creates 1 egg and 3 polar bodies which are useless in humans.
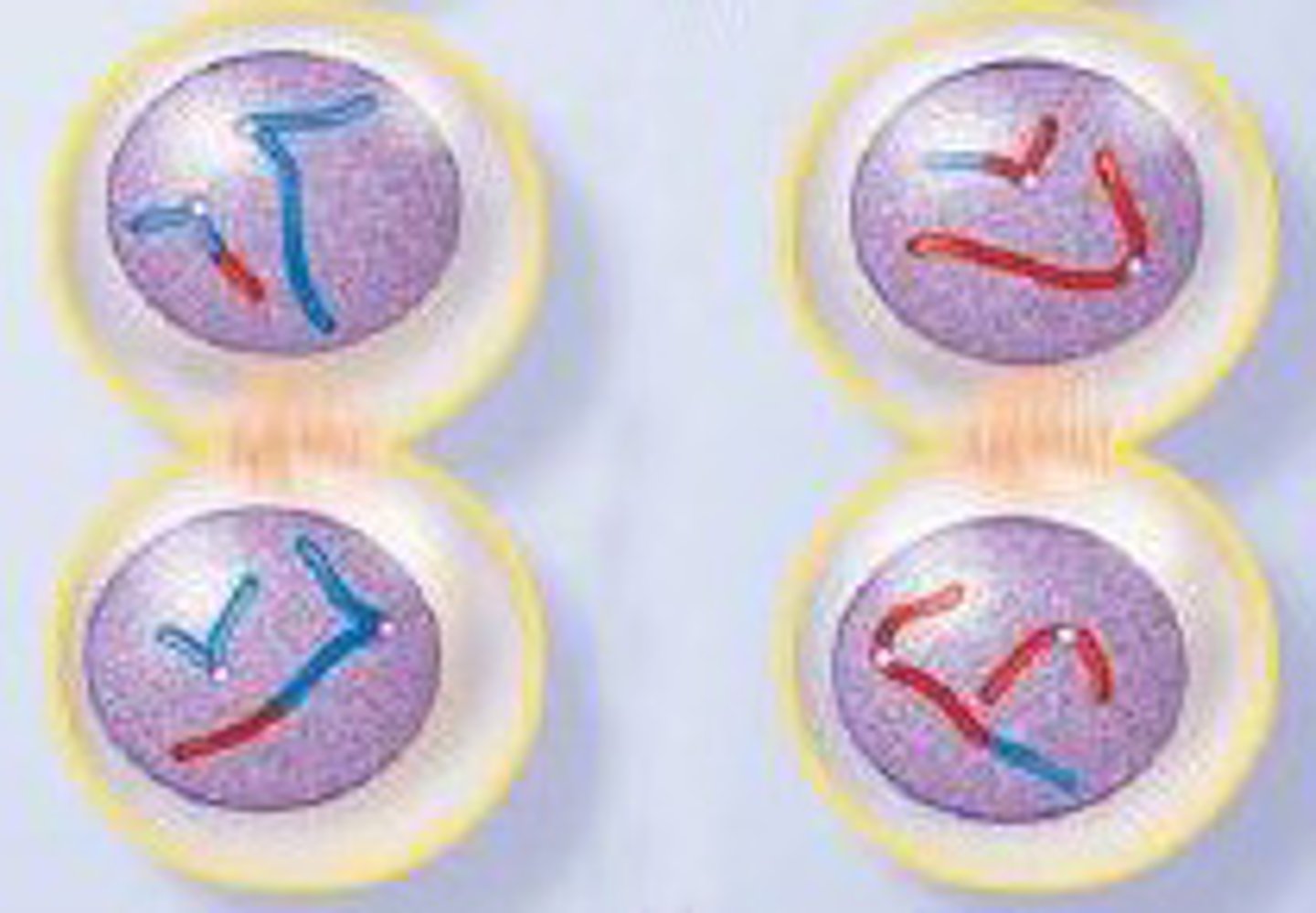
LAB 8: Cytokinesis
Division of the cytoplasm during cell division
LAB 8: What kind of cells does meiosis produce?
sex cells/gametes (haploid)
LAB 8: What kind of cells does mitosis produce?
somatic cells (diploid)
LAB 8: Where does mitosis take place?
somatic cells
LAB 8: Where does meiosis occur?
sex cells (gametes)
LAB 9: What are the three degrees of dominance?
complete dominance, incomplete dominance and codominance
LAB 9: complete dominance
a relationship in which one allele is completely dominant over another
The situation in which the phenotypes of the heterozygote and dominant homozygote are indistinguishable.
LAB 9: complete dominance example
Flower color changes if one allele is present.
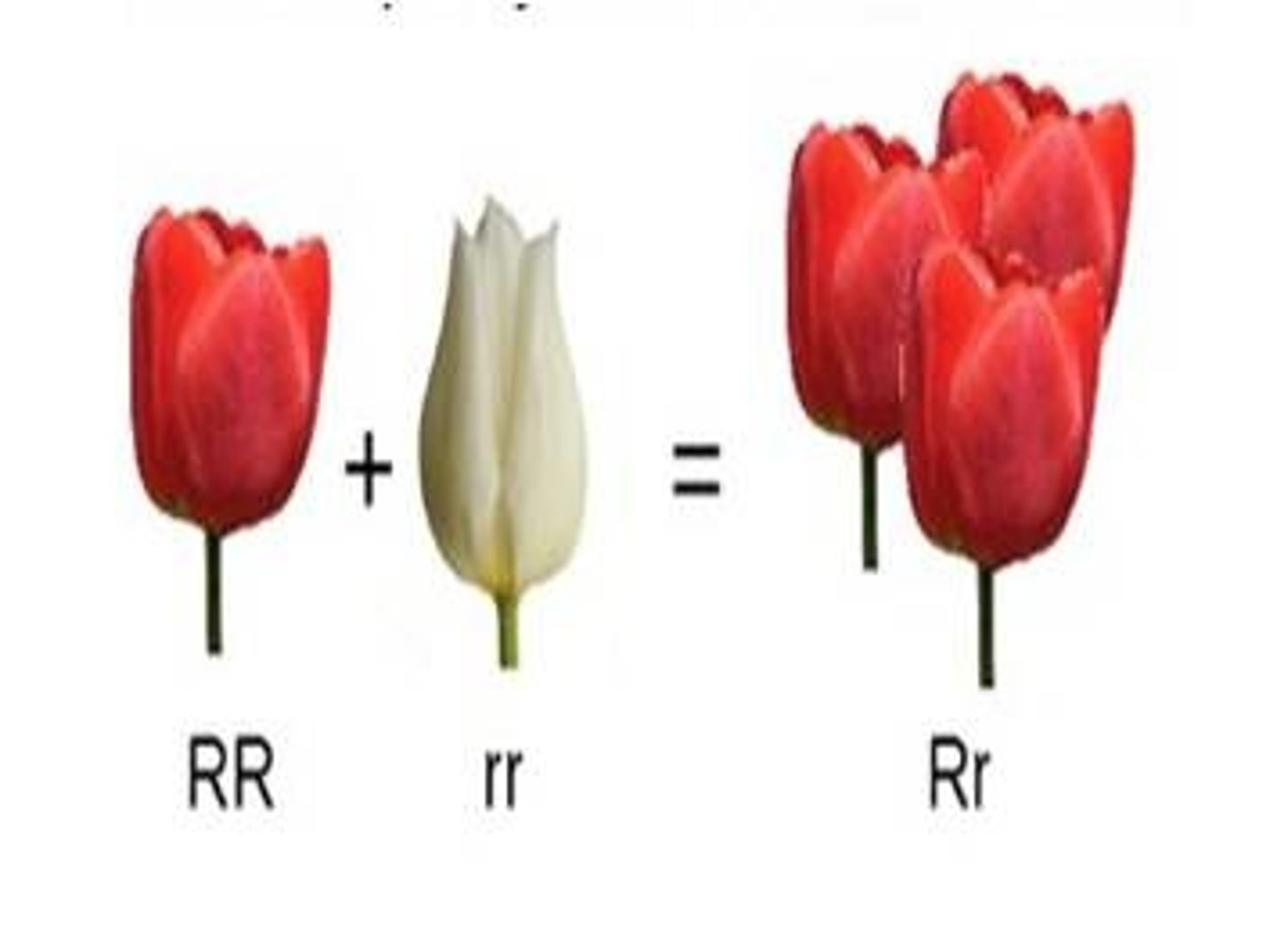
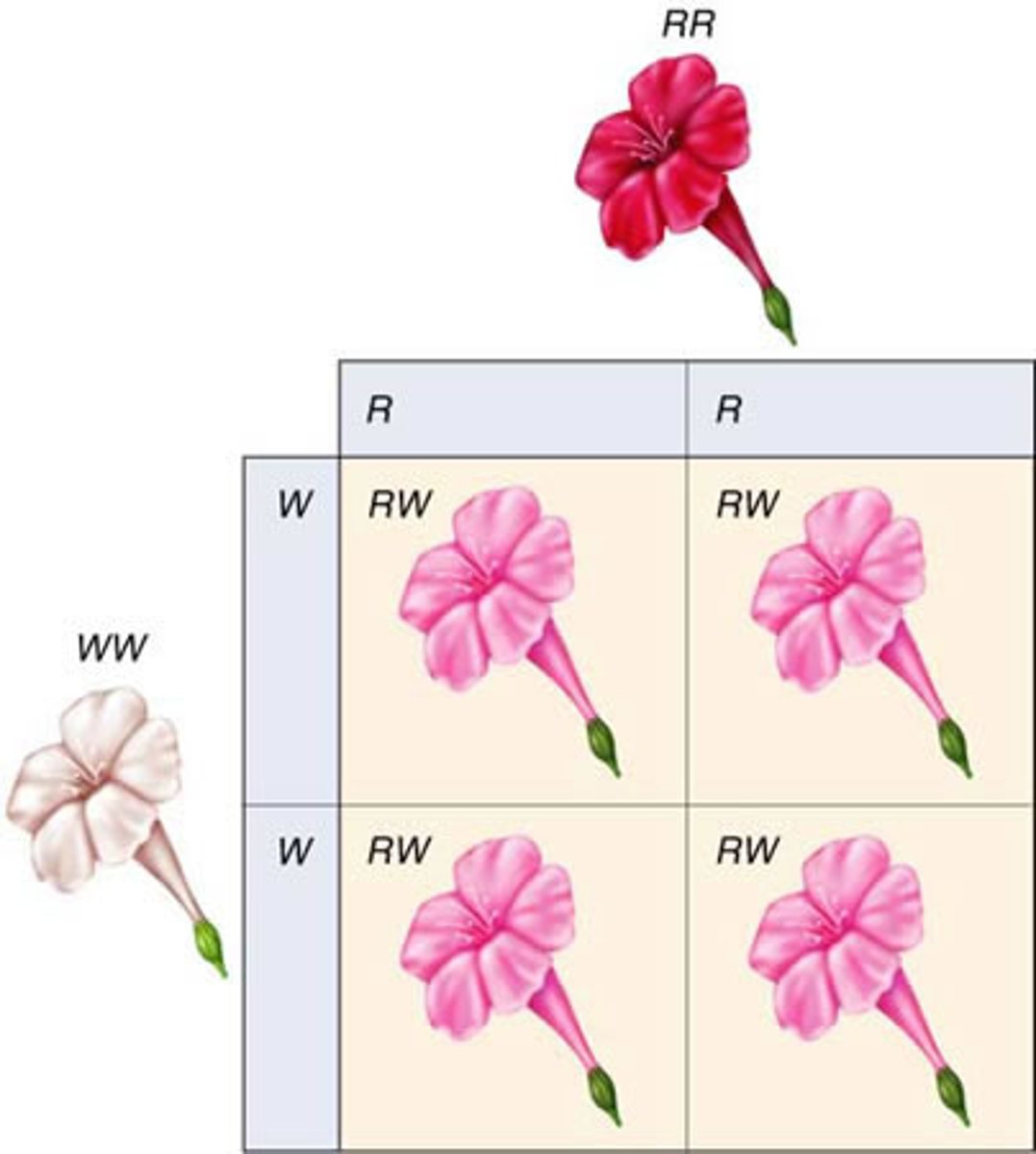
LAB 9: incomplete dominance
Situation in which one allele is not completely dominant over another allele
LAB 9: incomplete dominance example
2+ alleles influence phenotype (red x white = pink)
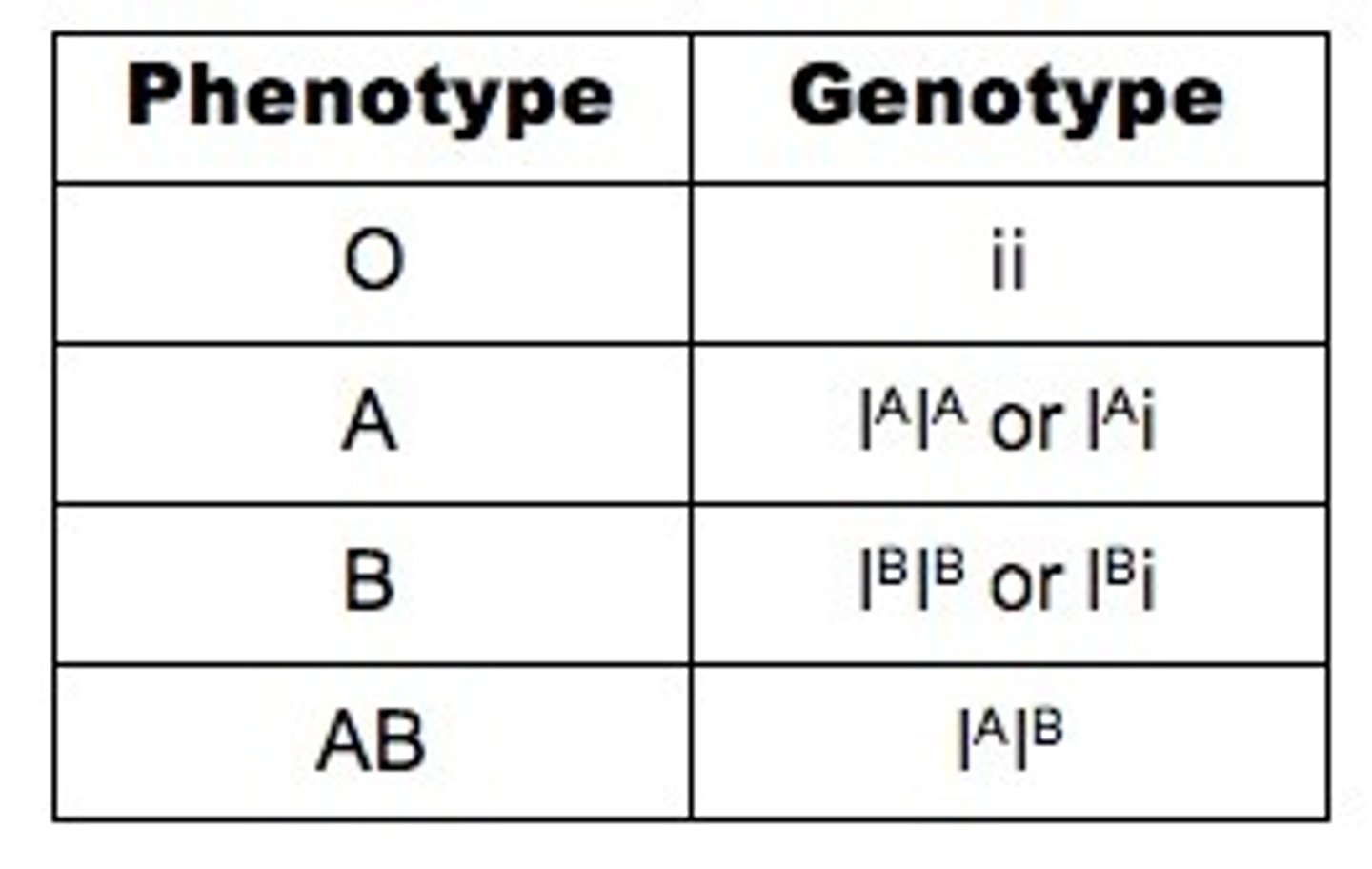
LAB 9: Codominance
situation in which both alleles of a gene contribute to the phenotype of the organism
LAB 9: Codominance example
Blood groups A, B, AB
a1-antitrypsin deficiency
LAB 9: How do you do monohybrid crosses
ex: Yy *yy = ?
You keep it like this and multiply in a box
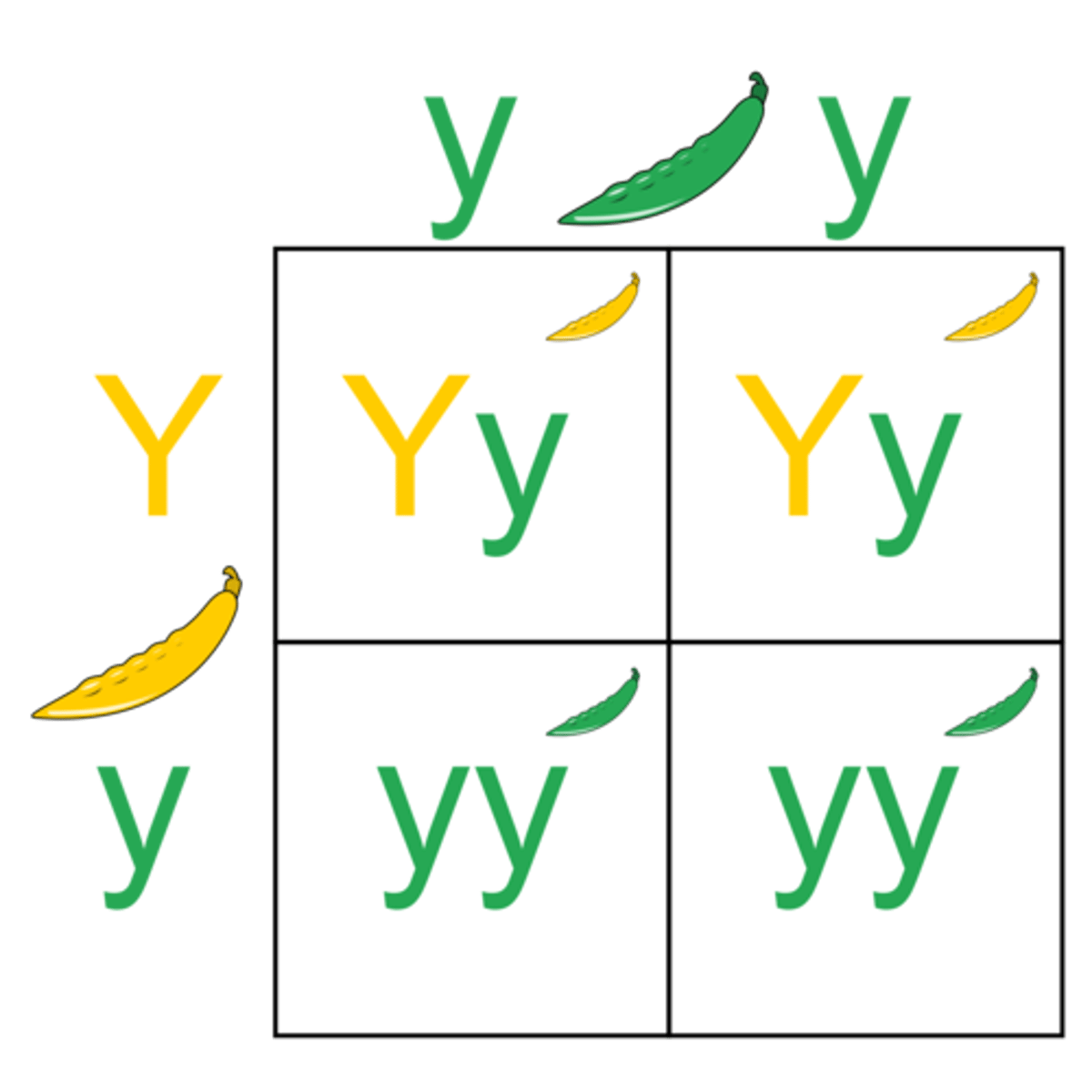
LAB 9: How do you do dihybrid crosses
ex: SsYy * SsYy = ?
You do 12341324 pattern to get what goes in each box
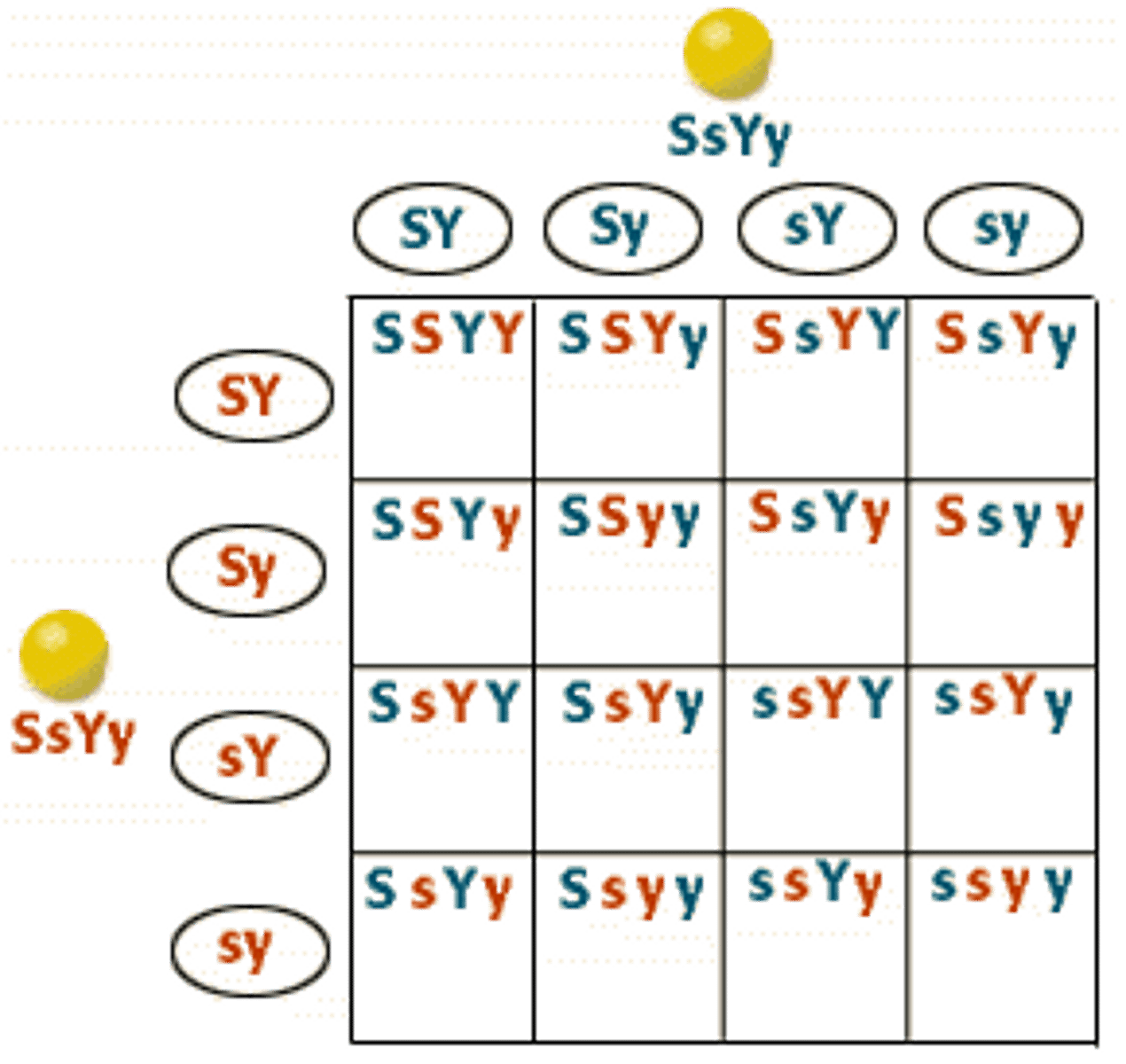
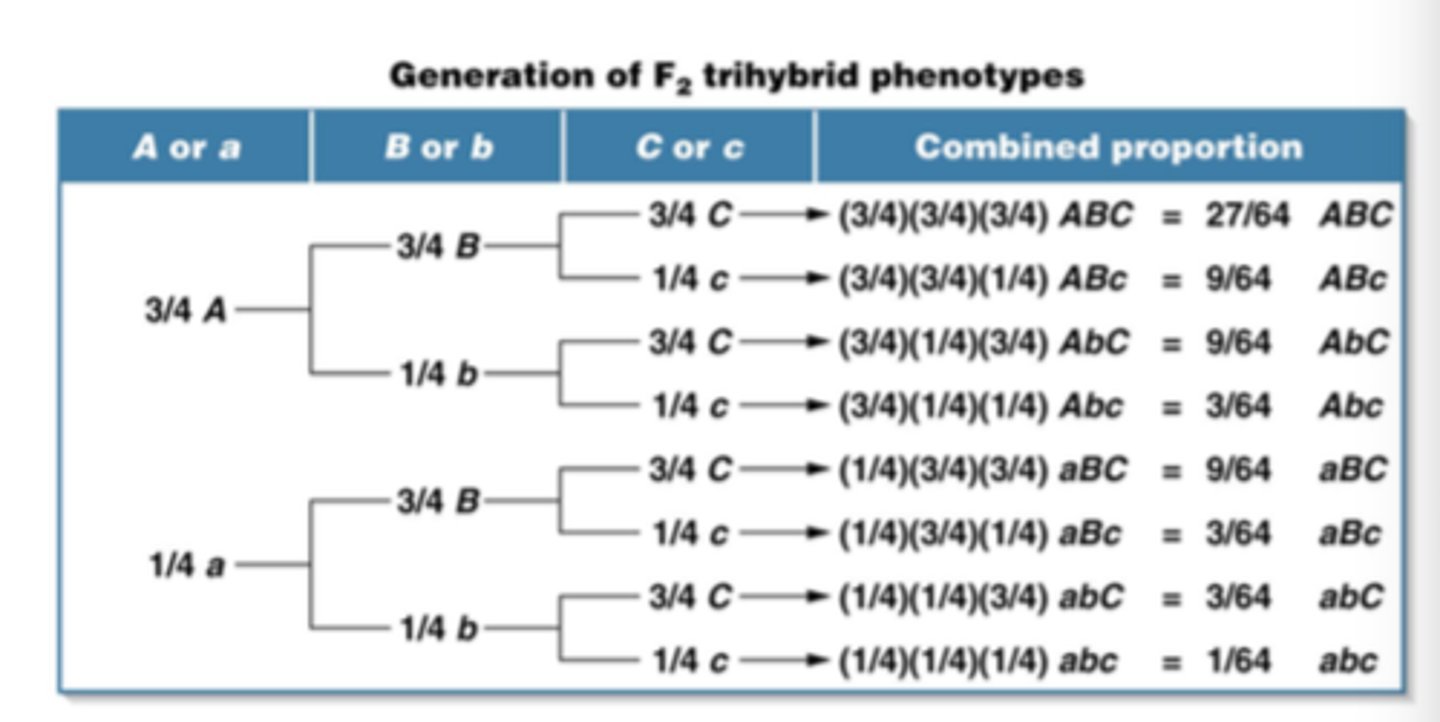
LAB 9: How do you do a fork diagram for dihybrid cross?
conduct a monohybrid for both individual genotypes
find ratio of allele presence in first genotype
multiply by allele presence of second genotype
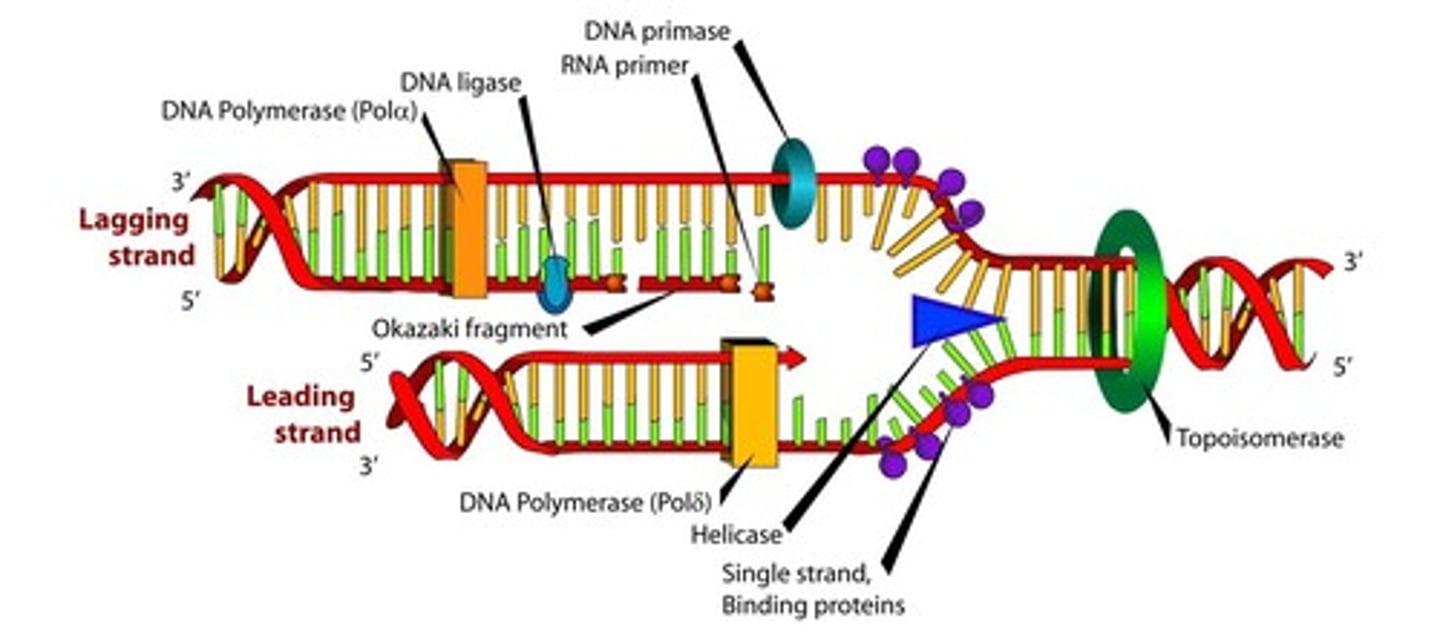
LAB 10a: What are the steps in DNA replication?
1) Helicase- unwinds the parental double helix
2) DNA topoisomerase - upstream of helices alleviating torsional strain
3) Single-strand binding proteins (SSBP) stabilize unwound DNA, aided by DNA gyrase.
4) Primase synthesizes a short RNA primer for DNA polymerase to bind to in the 5' to 3' direction to start replication on each strand.
5) DNA polymerase synthesizes the leading strand in 5' to 3' direction while the lagging strand is made discontinuously by primase making short pieces and then DNA polymerase extending these to make Okazaki fragments.
6) DNA ligase joins the Okazaki fragments together
LAB 10a: Convert an mRNA strand into a peptide, given a genetic code table: G T A C G C G T A T A C C G A C A T T C
mRNA: C A U G C G C A U A U G G C U G U A A G
Codons: AUG-CGC-AUA-UGG-CUG-UAA
Amino Acids: METHIONINE-ARGININE-ISOLEUCINE-TRYPTOPHAN-LEUCINE
LAB 10b: What are the steps of PCR?
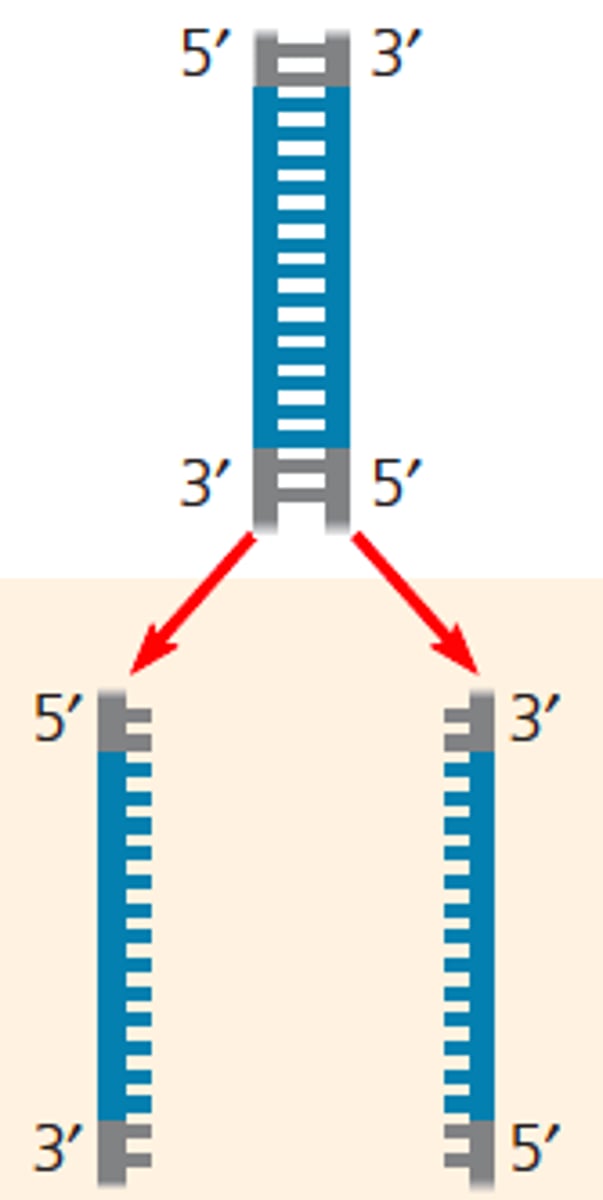
1. Denaturation
2. Annealing
3. Extension
LAB 10b: Denaturation (PCR)
-Step 1
-separation of the two strands of DNA
-Using high temperatures: 94 degree (boiling) to denature hydrogen-bonds between base pairs bases in two strands
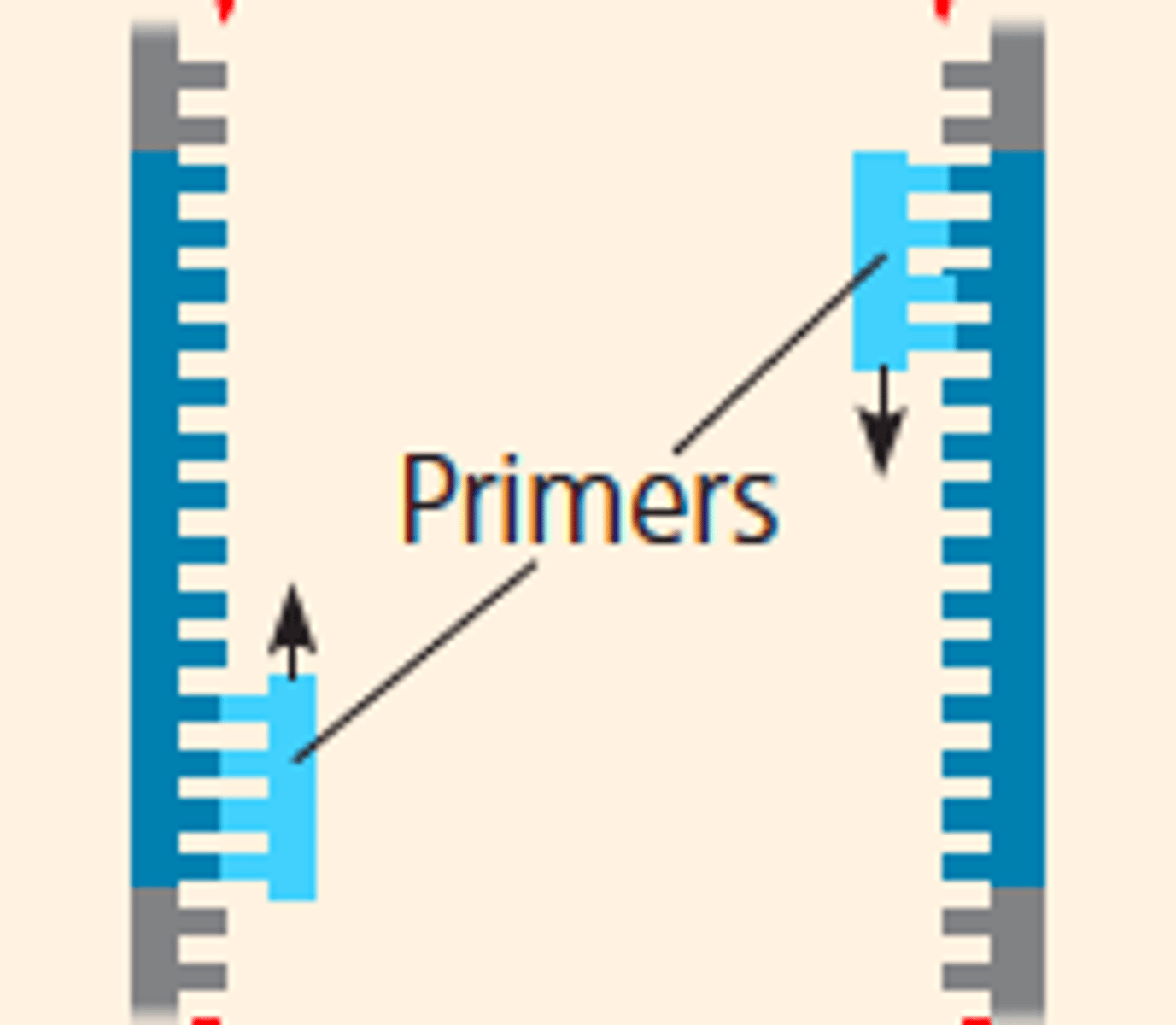
LAB 10b: Annealing (PCR)
-step 2
-DNA primers attach to opposite ends of the target sequence
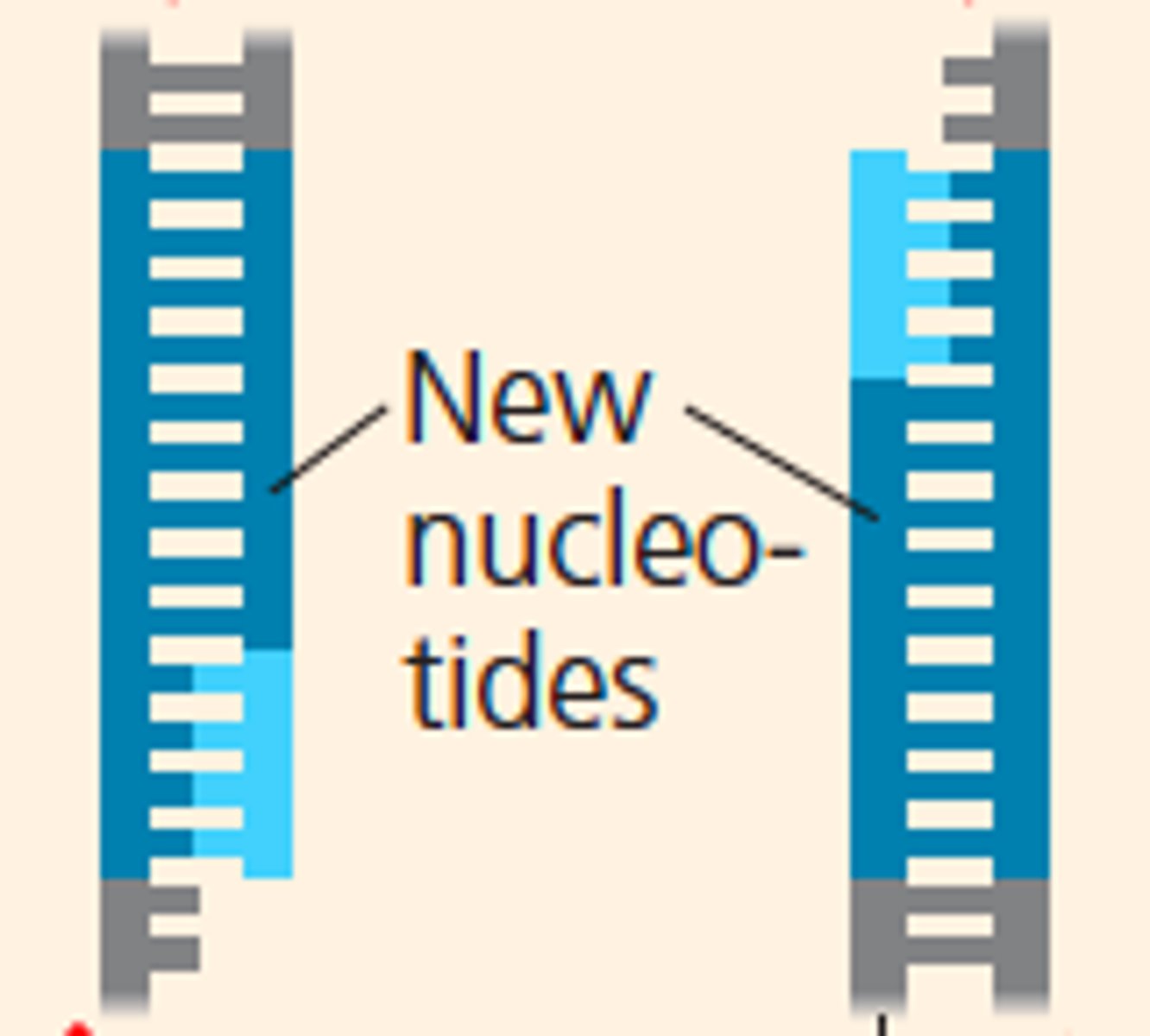
LAB 10b: Extension (PCR)
-step 3
-heating (72 degree C) to increase the rate of replication by DNA polymerase
-DNA polymerase add nucleotides to the end of the primer (3' hydroxyl group)
-It adds: dNTPs: building blocks of DNA (A,G,C,T)
-5'->3' direction of DNA polymerase
-DNA polymerase replicates each strand to produce more DNA
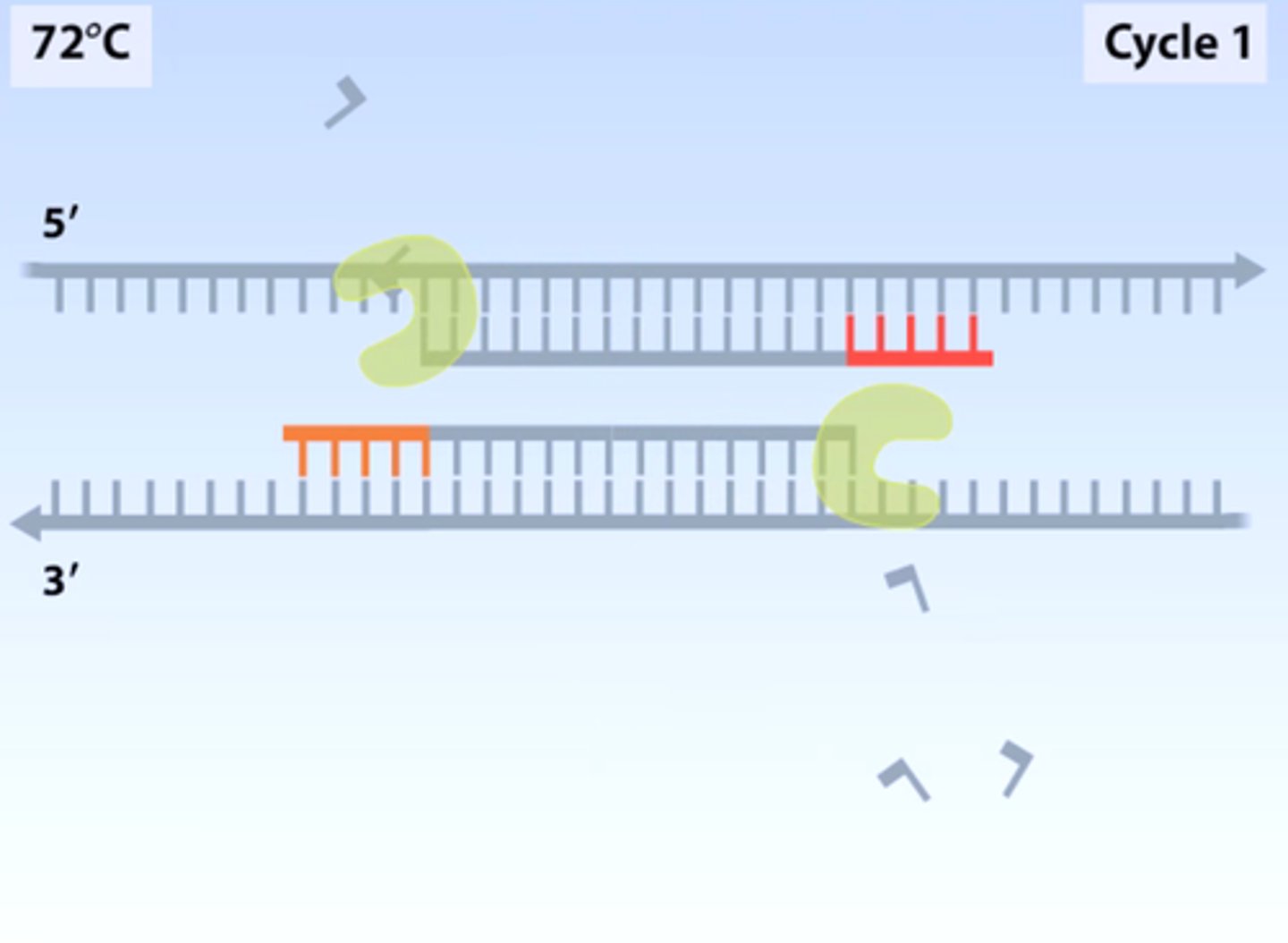
LAB 10b: What is Taq polymerase?
A heat-stable form of DNA polymerase extracted from bacteria that live in hot environments, such as hot springs, that is used during PCR technique
LAB 10b: What are ALL the necessary reagents for PCR?
template DNA, PCR primers, nucleotides, PCR buffer and Taq polymerase
LAB 10b: What is the master mix and why do you need each component?
individual building blocks of DNA (nucleotides, or dNTP's),
a special buffer to maintain optimum pH,
Salts and MgCl2 (Magnesium ions) are cofactors needed for the Taq polymerase to perform optimally.
LAB 10b: What are primers and how do they bind?
TPA-25 gene 'oligonucleotide primers' are designed specifically to anneal at opposite ends of opposite strands of the specific sequence of DNA that is desired to be amplified.
LAB 10b: Alu sequence
-a DNA sequence about 300 base pairs long
-5% of human DNA
LAB 10c: dna ladder
set of known DNA fragments with different sizes in base pairs or kilo bases. these DNA fragments are separated and visualized as DNA bands on a gel. used to determine the size and quantity of DNA fragments
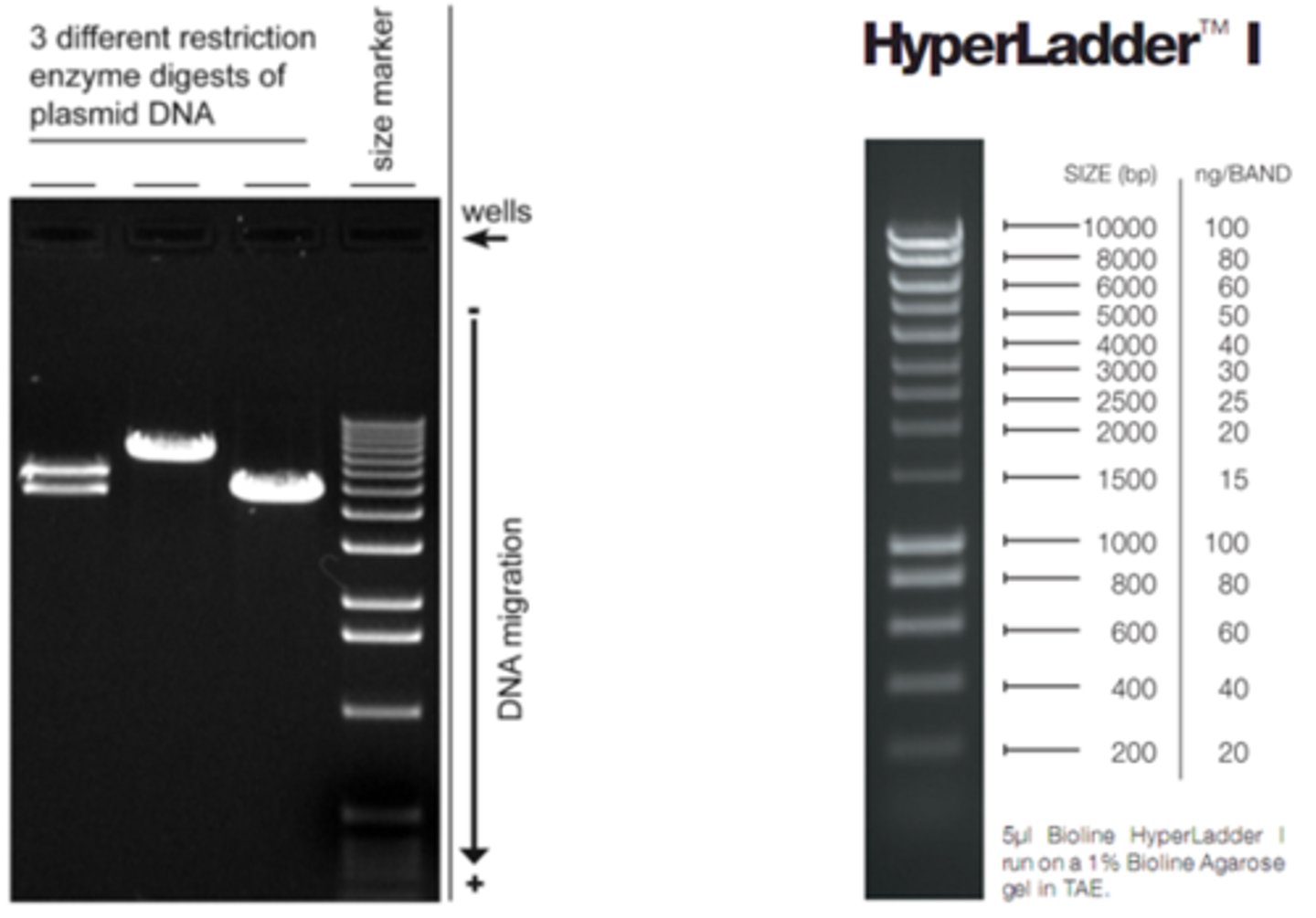
LAB 10b: What were you amplifying: sequence with Alu vs. sequence without?
-- = no insert = homozygous = single band at 136bp for TPA-25(Alu-)
++ = two copies of 300bp Alu insert = homozygous = single band on the gel at 436bp for TPA- 25(Alu+)
-+ = one no insert, one with 300bp Alu insert = heterozygous = two bands on the gel at 136bp and 436bp for TPA-25(Alu)
LAB 10c: Why does the DNA move through the gel during gel electrophoresis?
DNA is negatively charged, therefore, when an electric current is applied to the gel, DNA will migrate towards the positively charged electrode.
LAB 10c: What are the steps of gel electrophoresis?
1. Make the gel
2. Set up the gel apparatus
3. Load the DNA sample into the gel
4. Hook up the electrical current and run the gel
5. Stain the gel and analyze the results
LAB 10c: How are fragments separated in gel electrophoresis?
By length
LAB 10c: What does a band closer to the well mean in gel electrophoresis?
a long strand of DNA
LAB 10c: What does a band further from the well mean in gel electrophoresis?
a shorter strand of DNA
LAB 10c: Why do you need a molecular weight marker / DNA ladder?
Since molecular weight is inversely proportional to migration rate through a gel, the ladder gives an idea of how long a strand is of DNA by comparing it to the marker.
LAB 10c: What is the purpose of the loading dye?
1. Dye increases density of sample
-Sample will sink when added to wel
2. Dye monitors progress of gel in real time
-Separates into three colors
LAB 10c: What is the purpose of the 1X TAE electrophoresis buffer that fills the chamber?
to maintain the pH of the DNA solution to neutral.
LAB 10c: Be able to read a simple gel.
that's on you boo boo