exam 4 genetics review
1/87
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
88 Terms
Transcription
The process of making mRNA from DNA
Translation
Process of making protein from mRNA
Template strand
RNA is transcribed from one DNA strand
Nontemplate strand
Strand usually not transcribed
Ribosomal RNA
Cell type: prokaryotic and eukaryotic
Location of function in eukaryotic cells: Cytoplasm
Function: structural and functional components of the ribosome
Messenger RNA (mRNA)
Cell type: prokaryotic and eukaryotic
Location of function in eukaryotic cells: nucleus and cytoplasm
Function: carries genetic code for proteins
Transfer RNA (tRNA)
Cell type: prokaryotic and eukaryotic
Location of function in eukaryotic cells: Cytoplasm
Function: helps incorporate amino acids into polypeptide chain
The transcription unit
A promoter
RNA-coding sequence
Terminator
3 stages of transcription
Initiation, elongation, termination
Initiation
In which the transcription apparatus assembles
Elongation
In which DNA is threaded through RNA polymerase
Termination
The recognition of the end of the transcription unit
Promoter
Sequences that determine the location of the gene of where to start the process, does not transcribe RNA but has info of where to start polymerzation
Active transcription
Christmas tree like structure
Multiple RNA polymerases
transcribe the same gene simultaneously
Bacterial RNA polymerase
Five subunits made up of the core enzyme:
Two copies of α
Single copy of β
Single copy of β’
Stabilizing enzyme: ω
The sigma factor
Binding to the promoter when transcription starts
Bacterial promoters
Consensus sequences (an example)
Consensus sequences
Sequences that possess considerable similarity
-10 consensus: 10 by upstream of the start site
Pribnow box
5’ TATAAT 3’
3’ ATATTA 5’
-35 consensus sequence: TTGACA
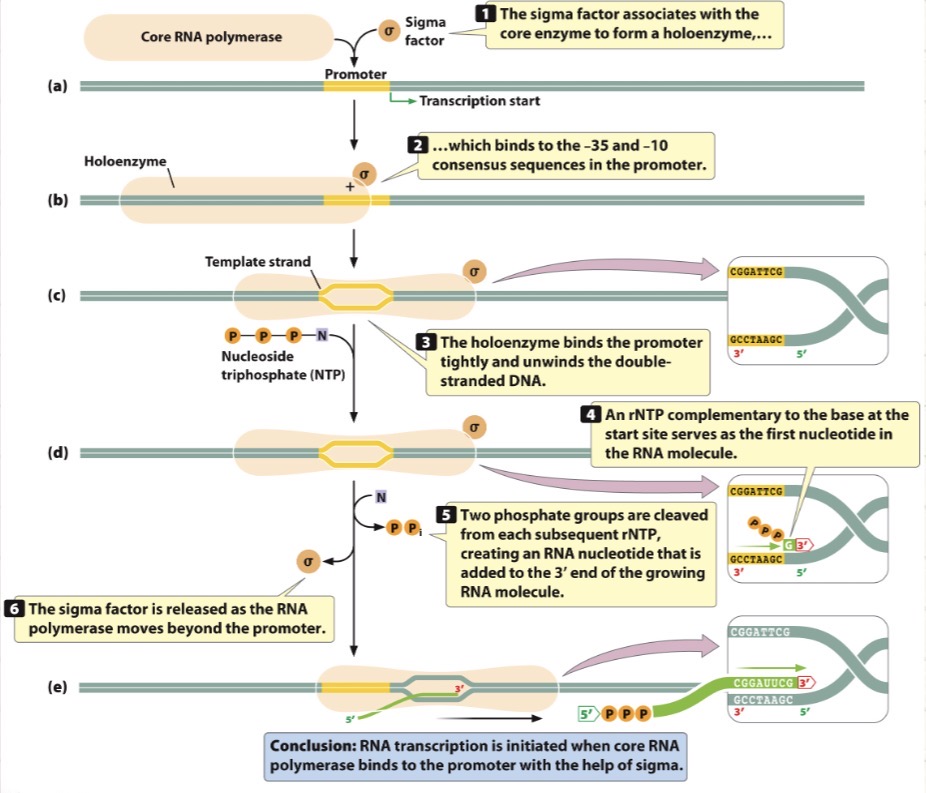
What step of transcription does this represent?
Elongation
Rho-dependent termination
Uses rho factor
Does not require primer
Rho-independent termination
Hairpin structure formed by inverted repeats, followed by a string of uracil’s
Does not need helicase
Promoters of eukaryotic transcription
Basal transcription apparatus
Transcriptional activator proteins
RNA polymerase II
Consensus sequences of RNA splicing
5’ : GU A/G AGU: 5’ splice site
3’ : CAGG
Spliceosome
Five RNA molecules + 300 proteins
Determines where the 5’ and 3’ consensus sequences are and splice them
(Eukaryotic) transcription by RNA polymerase II
Terminated when an exonuclease enzyme
attaches to the cleaved 5’ end of the RNA
Moves down the RNA
Reaches the polymerase enzyme
3 primary regions of mature mRNA
5’ untranslated→ the protein-coding region→ the 3’ translated region
Proteins
Nothing more than chain of amino acids just as DNA is a chain of nucleic acids
Triplets of codons
Sets of three DNA bases
(Why there are silent mutations)
All protein starts with the amino acid…
Methionine (AUG start codon)
Translational termination
Translation is ended once we come upon a stop codon
Translation
We land on the 5’ end of mRNA, scan for the first AUG start building a chain of amino acids until we reach a stop codon
Ribosome
The site of translation and allows information contained within mRNA to be used to make of amino acids
lands on the 5’ end of the mRNA and scans until it finds the first AUG
Each nucleotide have three things in common
A phosphate group, five carbon sugar (called ribose), a base
Phosphodiester bond
The bond holding nucleotide together
Protein synthesis
Requires the ribosome
mRNA are scanned for the first AUG; this lines up the ribosome
Then a tRNA recognizes the codon on one end and results an amino acid with the other
Structural genes
Proteins used in metabolism, biosynthesis, play a structural role
Regulatory genes
That regulatory protein (or RNA molecule) that somehow interact with DNA to alter the expression of other genes
Regulatory protein often directly bind to DNA to promote of inhibit gene expression
Constitutive genes
Genes that don’t need regulation and are always on
(DNA binding proteins) Domains
~ 60-90 amino acids, responsible for binding to DNA, forming hydrogen bonds with DNA
(DNA binding proteins) motif
within the binding domain, a simple structure that fits into the major groove of the DNA
Helix-turn-helix
Location: bacterial regulatory proteins; related motifs in eukaryotic proteins
Characteristics: two alpha helices
Binding site: major groove
Helix-loop-helix
Location: eukaryotic proteins
Characteristics: two alpha helices separated by a loop of amino acids
Binding site: major groove
Zinc finger
Location: eukaryotic regulatory and other proteins
Characteristics: loop of amino acids with zinc at base
Binding site: major groove
A lot of un-transcribed genomic DNA
Made up of regulatory elements
Positive regulation
Naturally gene is off- something must be done to turn it on (activation)
Negative control
Naturally gene is on- something must be done to turn it off (repression)
Operon
Cluster genes of similar function together
Lac operon
Three clustered genes clustered together under control of one promoter
Regulator gene
DNA sequence encoding products that affect the operon function but are not part of the operon (able to bind to DNA & make protein)
Inducible operon
Kept off and must be altered to express
repressible operon
kept on and must be turned off
negative inducible operon
the gene’s natural, unregulated state is on
the regulator is usually made however, so the gene is continually kept off
when signal is received the gene is needed, the repressor is not made or in-activated allowing the gene to expressed
negative repressible operon
also a repressor, natural unregulated state is on
usually not made, so the gene is continuously expressed
when signal is not received the gene is not needed, the repressor is made or activated allowing the gene to repress
positive inducible
natural condition: off
an activator
not usually made so the gene is continually off
when signal is received, the gene is needed, the activator is made or activated allowing the gene to be expressed
positive repressible operon
an activator
naturally off
usually made so the gene in continually expressed
when signal is received the gene is not needed, the activator is no longer made or in-activated allowing the gene to be turned off
all living cells need carbon building block and energy
glucose is the easiest and energetically cheapest sugar to digest
if you offer bacteria glucose and lactose
it will choose glucose (ATP)
If there is no glucose for bacteria to choose
it will choose lactose instead (if offered)
how bacteria make the choice at the molecular level if lactose and glucose are around, keep lac operon off (don’t express)
a negative inducible operon
lactose metabolism
structural genes:
lac Z
lac Y
lac A
regulation of the lac operon
inducer: allolactose
lac I
lac P
lac O
Lac Z
encodes B-galactosidases (breaks down into monomers)
lac Y
encoding permease (transports lactose to enter)
lac A
encoding transacetylase
lac I
repressor encoding gene
lac P
operon promoter
lac O
operon operator
allolactose
breaks down galactose→ glucose & lactose
when there is no lactose around
Lac I is sitting on the operator of the lac operon, blocking the RNA polymerase enzyme from transcribing these genes
the operon is repressed, but wants to be on
steps when lactose enters the environment
little bit of the permase is expressed
allows some to enter the cell
the little bit of B-galactosidase converts some of the lactose to allolactose
allolactose binds to lac I, forcing it to change shape
since shape = function, Lac I can not bind to DNA anymore
triptofan (Trp)
amino acid, keeps gene off when it’s there
makes singular mRNA
Trp characteristics
a negative repressible operon
five structural genes (trp E, D, C, B, A)
Trp E, D, C, B, A
five enzymes together convert chorismate to tryptophan
the trp operon is regulated by
transcription attenuation
the attenuator sequence
in the 5’— region of the leader sequence and can make the ribosome stall
the attenuator, which is part of the leader determines
if transcription will be attenuated at the end of the leader or, if transcription will continue into the genes for Trp synthesis
the leader (after operator)
is 162 nucleotides long includes 1-4 segments
if segments 3 and 4 base pair (transcriptional attenuation)
they form a hairpin structure that is the attenuation signal
if segments 2 and 3 base-pair (transcriptional attenuation)
transcription proceeds and the trp synthetic enzymes are made
somatic mutations
mutations in the DNA of any of your cells except your gametes (random not heritable)
germ line mutation
mutation in the DNA of the gametes
transition (base substitutions)
purine (A or G) changes to other purine (double ring
or pyrimidine (C or T) becomes the other pyrimidine (single ring)
transversion (base substitution)
when purines become pyrimidines or vis versa
expanding nucleotides repeats
the number of copies of a set of nucleotide repeats increases, creating repeating sequence
insertion
addition of one or more nucleotides
deletion
deletion of one or more nucleotides