Biochemistry Two Final
1/82
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
83 Terms
nucleotide
pentose, nucleic base, phosphate group
nucleoside
pentose, nucleis base
nucleic acid
chain of nucleotides
purines
adenine, guanine
pyrimidines
cytosine, uracil, thymine
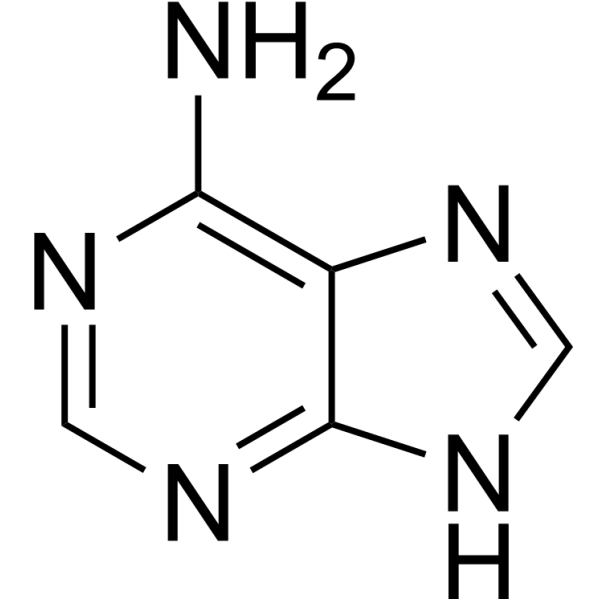
adenine
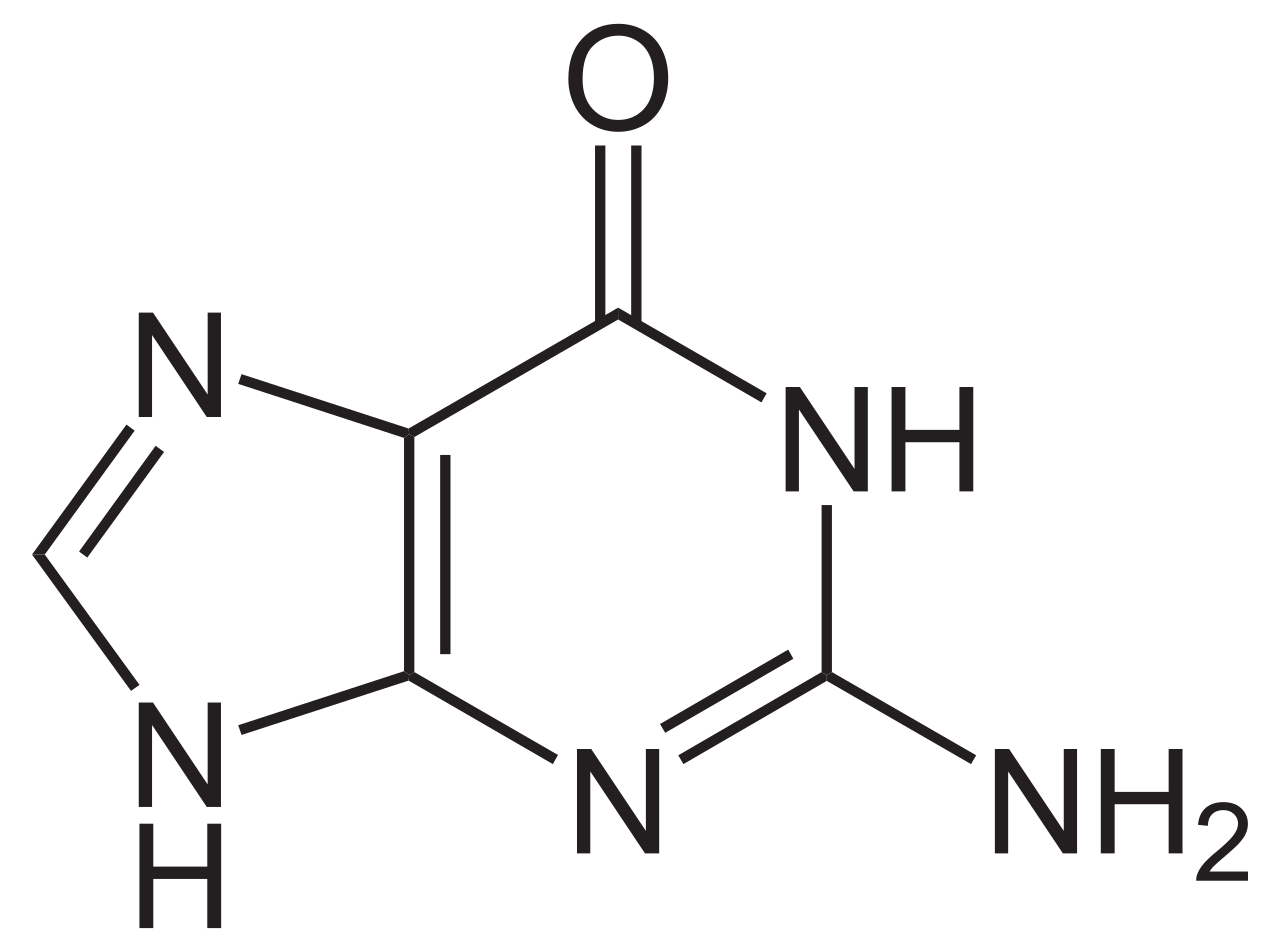
guanine
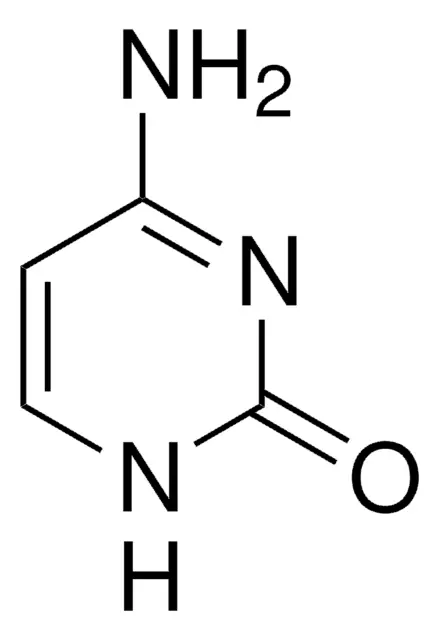
cytosine
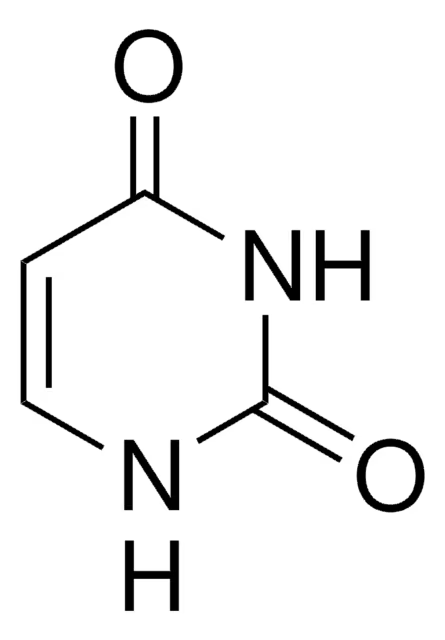
uracil
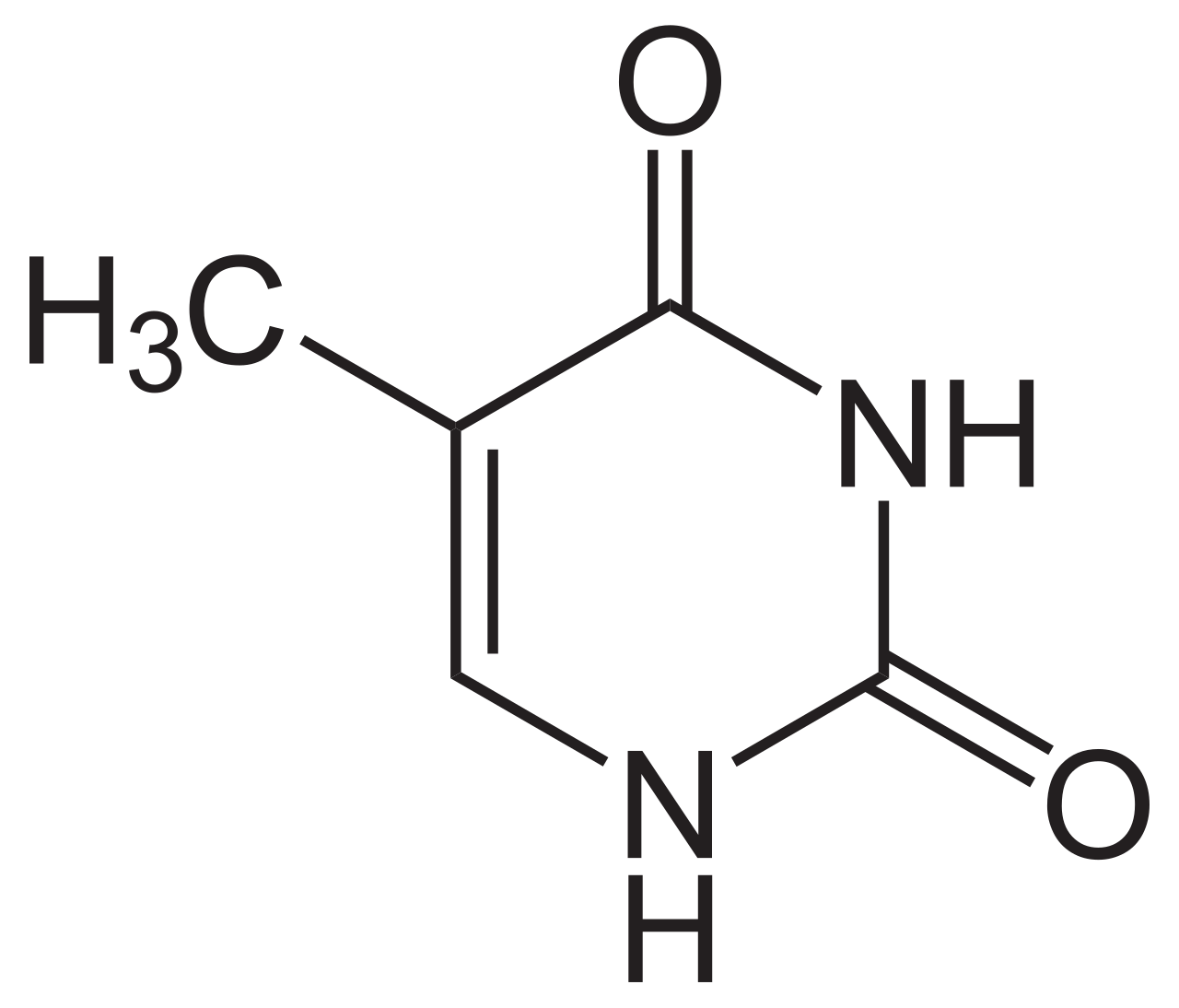
thymine
functional group at 3’ end of nucleic acid
OH group
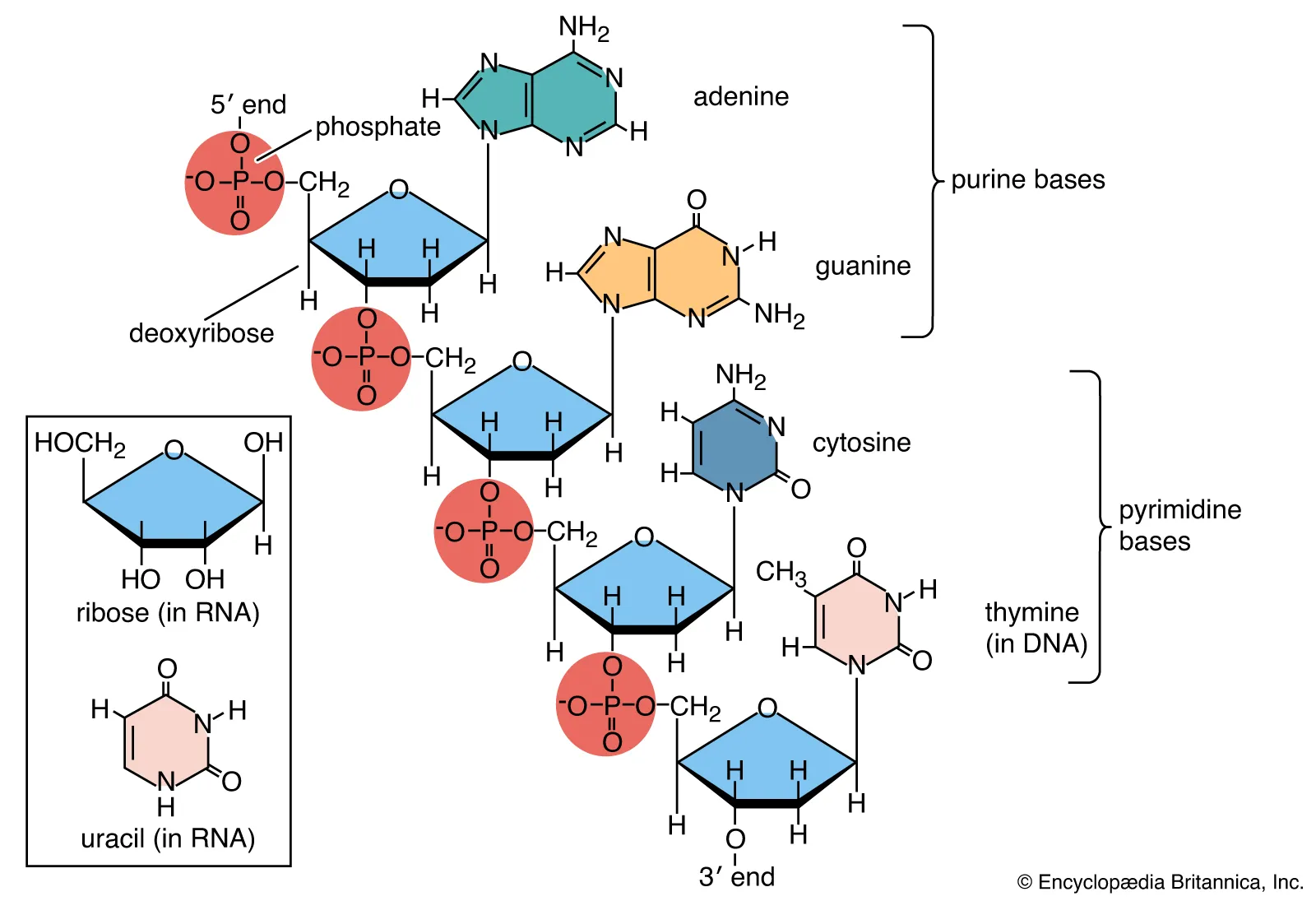
functional group at 5’ end of nucleic acid
phosphate group
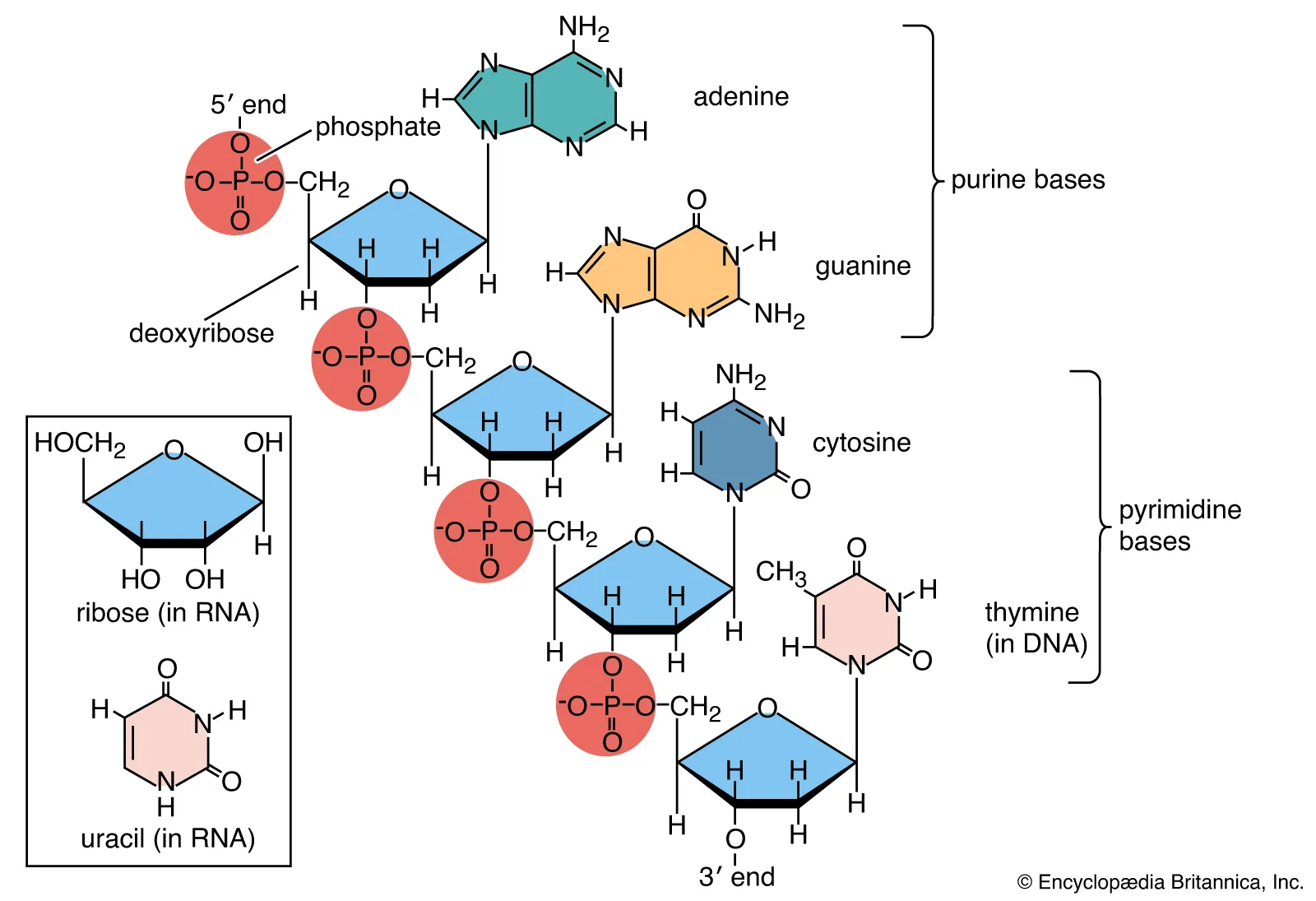
cause of DNA/RNA negative charge
-1 charge on phosphate group
features of Watson and Crick double helix
double helix, bases on interior of helix, phosphodiester backbone, antiparallel strands
stabilizing interactions of DNA helix
H bonds between complementary base pairs, base stacking interactions
base stacking interactions
noncovalent forces between DNA bases within the same stand- include H bonds and pi stacking
DNA vs. RNA comparison
No OH group at 3’ position in DNA, no uracil base in DNA, RNA typically single stranded
denature DNA
break noncovalent interactions
anneal DNA
reform noncovalent interactions
DNA length impact on TM
increased length increases melting temperature
GC content impact on TM
increased GC bps, increased Tm
salt concentration impact on Tm
increased salt concentration, increased melting temp
melting point of DNA using melting curve
melting point is when 50% of DNA is denatured
restriction enzymes
cleave palindromic sites 4-8 bp in length at the phosphodiester backbone
ethidium bromide
used to detect DNA restriction fragments in gels
components needed for PCR reaction mixture
4 dNTPs, template DNA, 2 primer sequences, Mg2+, DNA Polymerase I
impact of increased annealing temperature
higher specificity
effect of increased extension time
less specific gene amplification
characteristics of good DNA primer
18-25 nucleotides, 40-60% GC content, no hairpin structure, no primer dimer, high specificity, annealing temperature less than 5 degrees below Tm but between 50-62 degrees, primers within 5 degrees
3 types of membrane lipids
phospholipid, glycolipid, cholesterol
general form of chemical formula of fatty acid
number of carbons in FA:double bonds (deltadouble bond carbon numbers)
impact of double bonds on melting temp
more double bonds, lower melting temp
impact of FA chain length on melting point
increased chain length, increased melting temp
solubility of peripheral proteins
water soluble
solubility of integral proteins
extracted with detergent
delta G in hydropathy plot
energy associated with transferring AA from a hydrophobic to hydrophillic environment
active transport
ATP directly required, moving up concentration gradient
secondary active transport
uses ATP indirectly to move one molecule down its concentration gradient, then use that energy to move another molecule up its concentration gradient
facilitated diffusion/passive transport
allows movement of polar, large molecules down their concentration gradient
example of facilitated diffusion
K+ ion channel
selectivity filter
helps filter out ions based on size and interactions with amino acid backbone
hydration free energy
energetic cost of dehydrating and ion and replacing interactions with water with interactions with channel amino acids
N domain function
binds ATP and hydrolyses ATP, transfers P to P domain
P domain
phosphorylated by N domain
A domain
dephosphorylates the P domain and restores the P-type ATPase to its original conformation
effector enzyme (like adenylate cyclase)
modulate reactions in response to cellular signals
agonist
binds and mimics natural ligand
antagonist
blocks activity of receptor
inverse agonist
decreases activity
Kd
binding constant, lower means better binding
AKAP5
binding site for the enzymes, effector molecules, signaling molcules present in epinephrine signaling cascase
turn off B2-adrenergic receptor
self inactivation, reduction in ligand concentration, desensitization of receptor, removal of secondary messenger, dephosphorylation of activated enzymes
high phosphoryl transfer potential
high standard free energy of hydrolysis
reasons why ATP hydrolysis drives reactions forward
orthophosphate released is resonance stabilized, electrostatic repulsion decreases with release of orthophosphate, release of orthophosphate increases entropy, hydration stabilizes the released ADP+Pi
creatine phosphate
molecule wiht higher phosphoryl transfer than ATP, meaning it can regenerate ATP
constitutional isomer
different connectivity
stereoisomer
multiple chiral centers reversed
enantiomer
every chiral center is exactly flipped
epimer
only one chiral center reversed
d-isomer of sugar
OH group on right side
L-isomer of sugar
OH group on left side
2 phases of glycolysis
energy investment and payoff phase
first committed step fo glycolysis
#3- PFK. product has to be part of glycolytic pathway
activators of glycolysis in muscle
F-1,6-P (activates pyruvate kinase), AMP (activates PFK)
inhibitors of glycolysis in muscle
ATP (PFK and pyruvate kinase), G-6-P (hexokinase)
inhibitors of glycolysis in liver
citrate, glucokinase, phosphorylation (pyruvate kinase)
activators of glycolysis in liver
F-2,6-P (PFK)
location of glycolysis
cytoplasm
location of CAC
matrix
location of electron transfer chain
inner nitochondrial membrane
location of oxidative phosphorylation
inner mitochondrial membrane
E1 of PDC
pyruvate dehydrogenase
E2 of PDC
dihydrolipoyl transacetylase
E3 of PDC
dihydrolipoyl dehydrogenase
E1 prosthetic group
TPP
E2 prothetic group
lipoamide swinging arm
E3 prosthetic group
FAD+
reaction catalyzed at E1
oxidative decarboxylation of pyruvate
reaction catalyzed at E2
transfer of acetyl group to CoA
reaction catalyzed at E3
regeneration of the oxidized form of the lipoamide
prosthetic group
covalently (or very tightly) attached group to an enzyme
cofactors of PDC
NAD+and CoA