lecture 26: positional cloning to locate a disease locus
1/20
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
21 Terms
positional cloning
object is to identify disease-causing genes by genetic linkage to polymorphic loci
narrows down focus of where disease is coming from
strategy of positional cloning
same as linkage analysis using two phenotypes, except
one gene tracked by phenotype (disease phenotype)
the other by DNA genotype (gene variant)
use microarrays to simultaneously analyze millions of two-point crosses, each one a test for linkage between a disease locus and a DNA maker (if they segregate tg, your getting closer to finding disease causing)
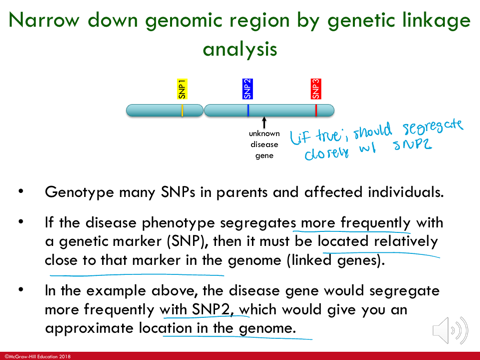
narrow down genomic region by genetic linkage analysis
genotype many SIPs in parents and affected individuals
if the disease phenotype segregates more frequently with a genetic marker (SNP), then it must be located relatively close to that marker in the genome (linked genes)
in the example, the disease gene would segregate more frequently with SNP2, which would give you an approximate location in the genome
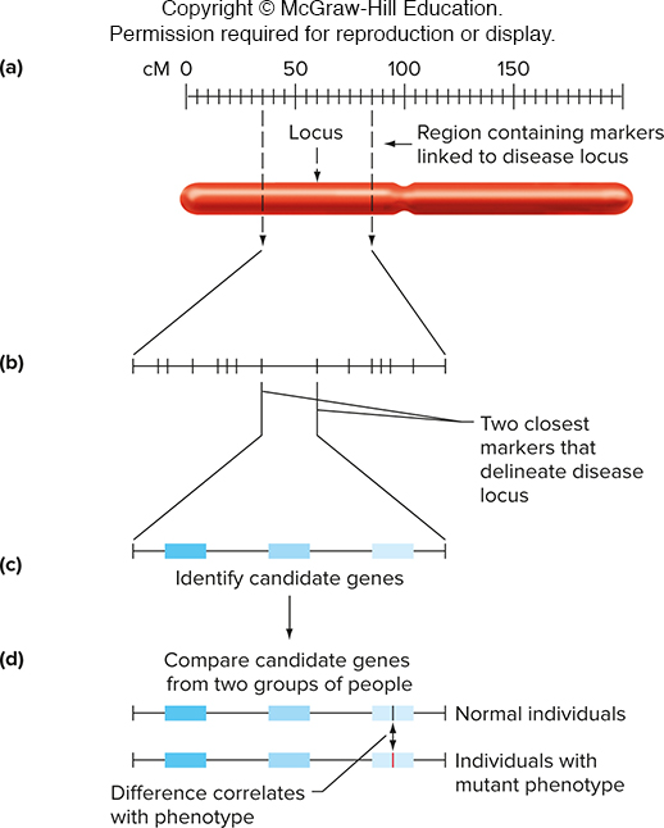
steps of positional cloning
region of interest narrowed by finding closely linked DNA markers
candidate genes are located in the region of interest.
sequence and expression of candidate genes are determined in normal and diseased individuals
Neurofibromatosis is an example of positional cloning!!
what is it
autosomal, dominantly inherited
causes proliferation of nerve tissue
positional cloning example determines whether or not a SNP is linked to the neurofibromatosis gene.
children in generation III are in effect the result of a testcross
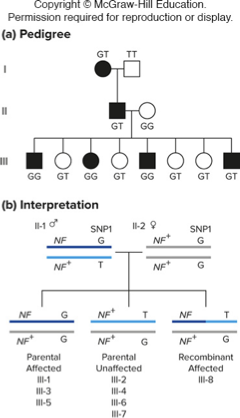
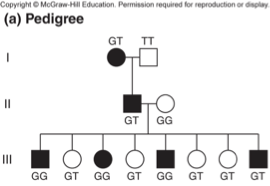
Breakdown of pedigree information #1
Gene 1: NF gene
Black squares and circles indicate affected individuals (disease phenotype)
Gene 2: SNP
Letters indicate genotype at a particular SNP
(G or T allele)
If you look in generation III:
•II-1 parent is heterozygous for NF gene and heterozygous for SNP.
•II-2 parent is homozygous for wildtype NF gene and homozygous for G allele, so this is essentially a test cross!
•Phenotype and genotype determined by parent II-1
•3 out of 4 affected individuals (mutant NF gene) are GG. Since this is more than 50%, this suggests that the NF gene and the SNP might be linked
•Wildtype NF gene is on the same chromosome as T allele since all unaffected are GT
•Mutant NF gene is on the same chromosome as G allele since most affected are GG (parentals most abundant)
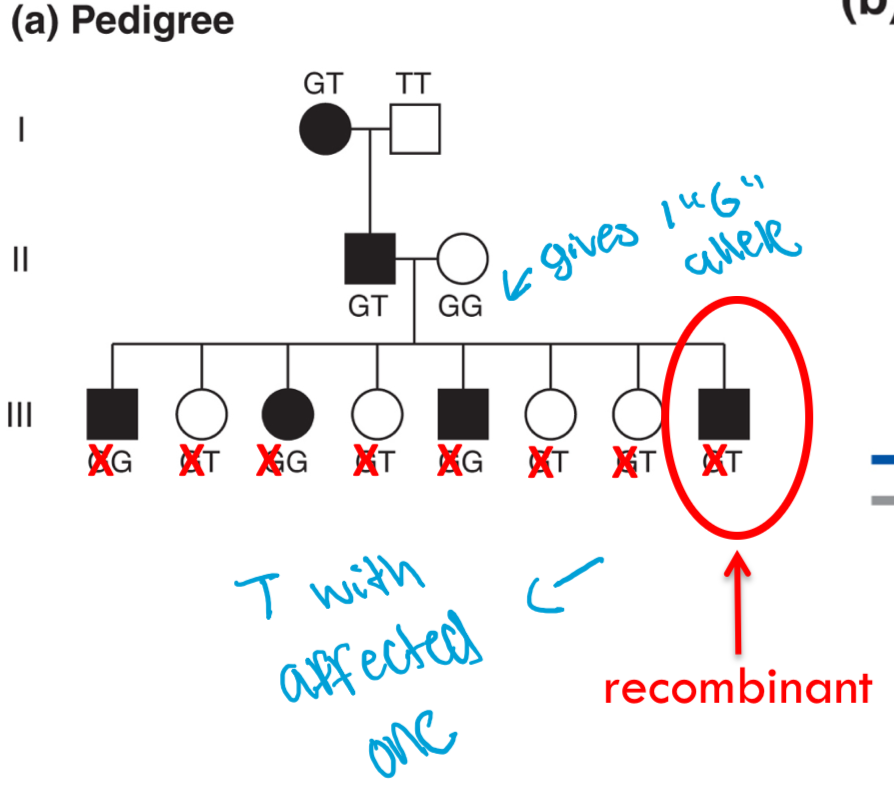
what does these results reflect?
because the G allele segregates with the disease NF allele more frequently, the g ALLELE MUST BE LOCATED on the same chromosome as the the disease allele in parent II-1
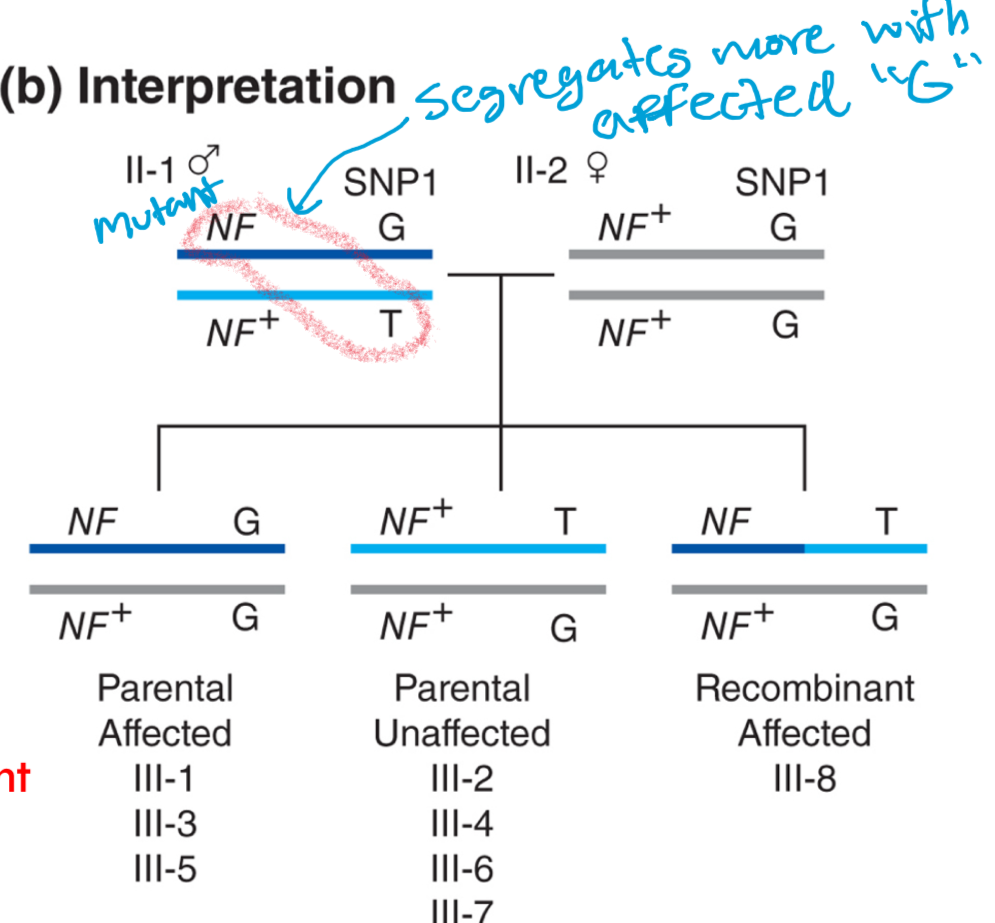
positional cloning limitations:
configuration of alleles not always known
not all matinas are informative:
always want one of the parents to be heterozygous for BOTH alleles (disease allele & gene variant)
most informative crosses will have the mate be homozygous for both alleles (a test cross!)
difficult to obtain sufficient pedigree data in humans
hard to tell w/ only 1 or 2 offspring
1 parent is homozygous for both, 1 parent is hetero for 1 and homo for other
NON-INFORMATIVE
1 parent is heterozygous for both, 1 parent is hetero for 1 and homo for the other
SEMI-INFORMATIVE
The Lod score is a statistical tool
for studying linkage #1 (for a small data set)
TRUE
use a statistical test called a lod score to determine if data are sufficient to conclude with confidence that a disease gene and a marker are linked.
linked lod score is…
lod>3
not enough info to determine if linked lod score is…
lod<3
If you don’t have enough information from one mating, you can analyze other matings in the same pedigree or a different pedigree and add the lod scores together
Remember the first law of logarithms:
log(A*B)= logA + logB
In the previous example, if you had data from two more matings with the same results, the total Lod score would be:
Lod= 1.1 + 1.1 + 1.1 = 3.3
..
allelic heterogeneity
disease caused by different mutations in the same gene
individuals with certain alleles may respond to drug treatment, while others do not
compound heterozygote
individuals with different mutant alleles of the same gene
locus heterogeneity
disease caused by mutation in one of two or more different genes
mutations in different genes tht cause same disease
sequencing of an entire genome become routine at $2,000
sequencing the whole-exome (limited to expressed parts of the genome) is less expensive
true
The gene, XIAP, was identified as causing Nic’s disease by sequencing entire genome through many combinations.
TRUE
Nic’s XIAP gene had a missense mutation that changed a single amino acid that was completely conserved among humans, frogs, flies, and other species
In contrast, XLPD-causing mutations are nonsense and frameshift mutations
this mutation causes his condition
pinpointing a disease gene may point to treatment, how?
XIAP causes another disease (XLPD) that results in production of too many white blood cells.
this is treated with bone marrow transplant…and BMT improves his symptho’s!