BIO 81 Lab Practicum Exam 2
1/103
Earn XP
Description and Tags
Labs 9-16
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
104 Terms
What is paper chromatography?
A technique used to separate substances in a mixture based on their movement up a piece of paper via capillary action.
Which factors determine how far a pigment travels?
Solubility in the solvent, molecular size, and intermolecular interactions (like hydrogen bonds) with the paper fibers.
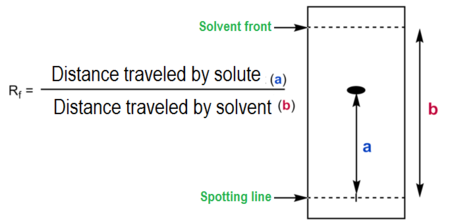
What is the formula for the Retardation factor (Rf)? What is it used for?
A dimensionless ratio in chromatography representing the distance a compound travels divided by the distance traveled by the solvent front,
typically ranging from 0 to 1
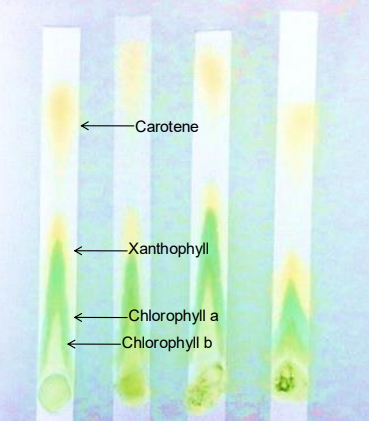
Identify the four pigments separated in this lab from furthest to shortest distance traveled.
Beta carotene: Yellow to yellow-orange; travels furthest because it is highly soluble and forms no H-bonds with the paper.
Xanthophyll: Yellow; contains oxygen and forms some H-bonds, making it less mobile than carotene.
Chlorophyll a: Bright green to blue-green.
Chlorophyll b: Yellow-green to olive-green; travels the shortest distance because it binds most tightly to the paper
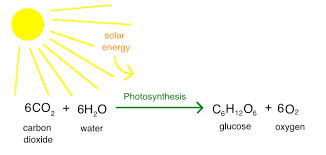
What is the overall chemical equation for photosynthesis?
What are the two stages of photosynthesis and where do they occur?
Light Reactions: Occur in the thylakoids; capture sunlight to produce ATP and NADPH.
Calvin Cycle (Dark Reactions): Occurs in the stroma; uses ATP and NADPH to convert CO2 into sugar.
What is the role of DPIP in the photosynthesis experiment?
It acts as an electron acceptor, replacing NADPH in the light reactions.
How does photosynthesis affect spectrometer absorbance readings?
As photosynthesis progresses and blue DPIP is reduced to colorless, the absorbance value decreases.
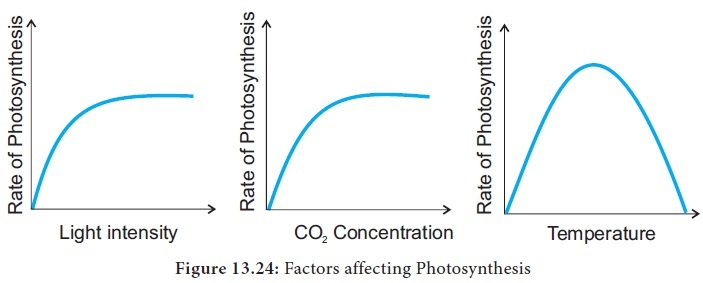
What are some variables that affect the rate of photosynthesis?
Light Intensity:
As light intensity increases, the rate of photosynthesis increases because more energy is available for the light-dependent stage. However, this rate will plateau (saturate) at a certain point, where another factor becomes limiting.
Carbon Dioxide (CO2) Concentration:
(CO2) is a raw material for photosynthesis. Increasing the concentration increases the rate of photosynthesis until the process hits a saturation point.
Temperature:
Photosynthesis is enzyme-driven, and its rate increases as temperature rises up to an optimal point, usually around 35 degrees C or higher, depending on the plant. Beyond this optimum, enzymes begin to denature, causing the rate of photosynthesis to decrease.
Water Availability:
Water is essential for the light-dependent reaction. A lack of water causes stomata to close to conserve water, which reduces the inflow of (CO2) and can stop the process in extreme cases.
How do you interpret color changes in DPIP?
Oxidized DPIP: Blue
Reduced DPIP: Colorless
Result: A change from blue to colorless indicates that the light reactions are occurring
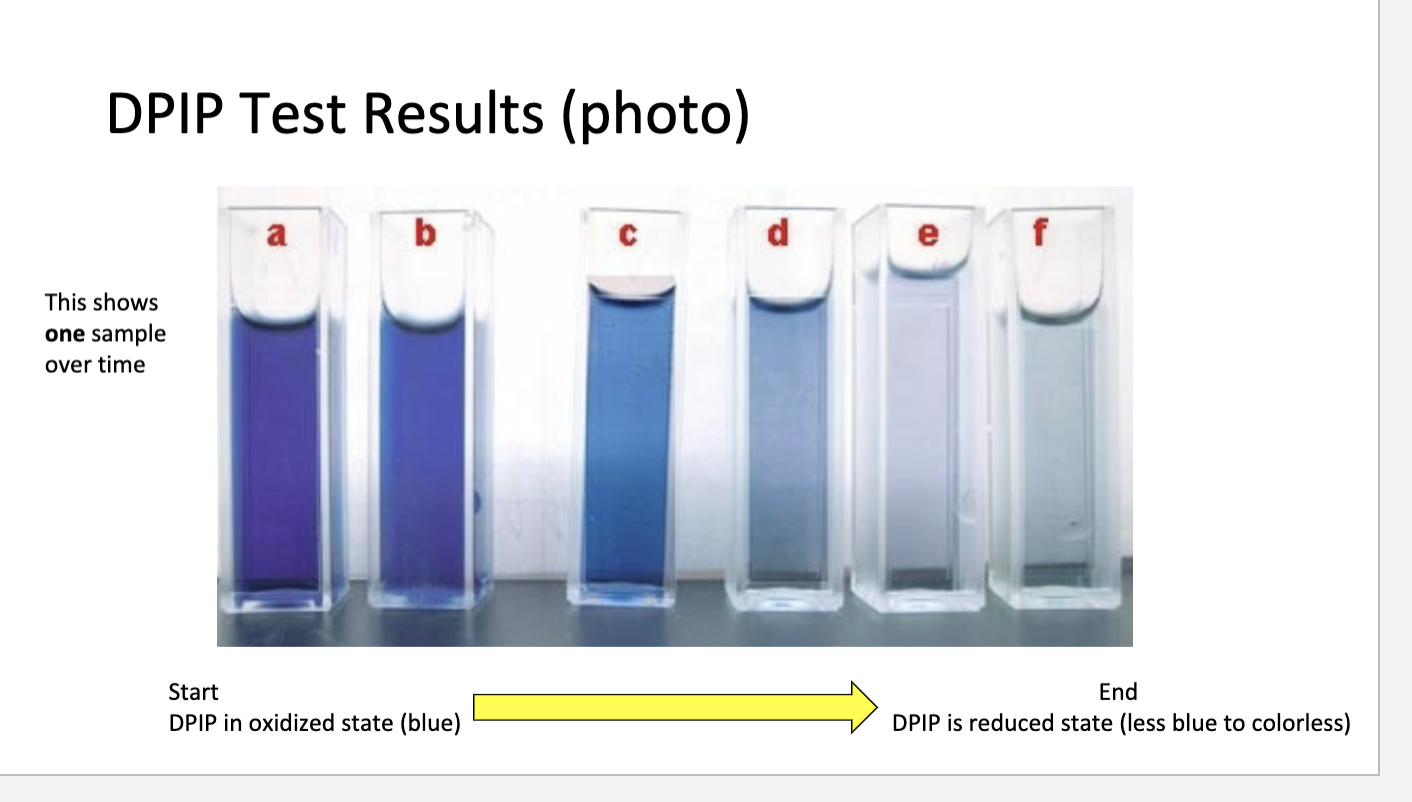

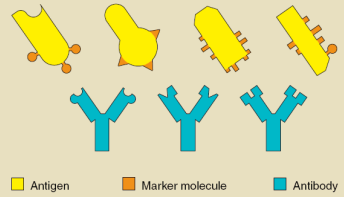
Antigen vs. Antibody:
Antigen:
Molecules from an invading entity (like a virus or bacteria) that trigger an immune response
Antibody:
Y-shaped proteins made by B cells that bind specifically to antigens to mark them for destruction
Antibodies bind to antigens very specifically and target invaders for destruction
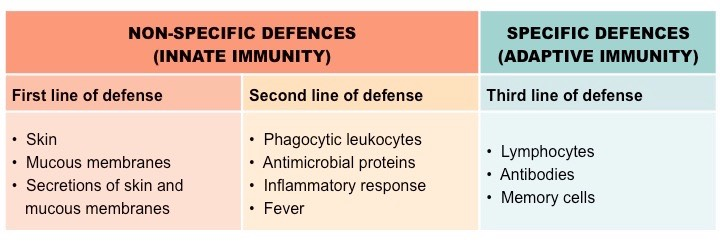
Nonspecific vs. Specific Defense:
Nonspecific:
Physical barriers and general immune responses that act regardless of the invader type.
Specific:
Tailored responses (B and T cells) triggered by specific antigens

What is the purpose of an Indirect ELISA?
ELISA - Enzyme-Linked Immunosorbent Assay
To detect the presence of specific antibodies in a patient's blood, indicating they have been infected and launched an immune response.
Define the key components of the ELISA used in this lab:
Primary Antibody:
The antibodies found in a patient's serum that bind to the target antigen.
e.g., a viral protein
Secondary Antibody:
An anti-human antibody conjugated to an enzyme; it binds to the primary antibody.
e.g., HRP or AP
Chromogen Substrate:
A chemical that changes color if the enzyme (on the secondary antibody) is present.
e.g., TMB
What are the basic steps of the ELISA procedure?
Add Antigen to the wells.
Add Patient Sample (contains primary antibodies if positive).
Add Secondary Antibody (with enzyme).
Add Chromogen and observe for color change.
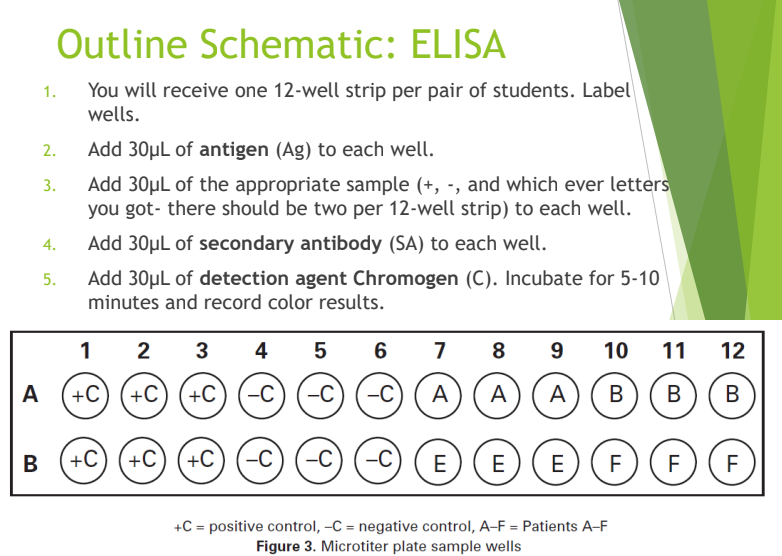

How do you interpret ELISA results?
Color change: Positive result; the patient has antibodies and was likely infected.
No color change: Negative result; the patient has no antibodies against that antigen
Why are samples tested in triplicate in the ELISA lab?
To ensure reproducibility and to help identify potential contamination or false positives.
Primary function of kidneys
To filter waste from the blood to be excreted as urine.
Normal components of urine
Water, nitrogen compounds, salts, and small amounts of protein and glucose.
Significance of glucose in urine
High levels may indicate diabetes.
Significance of protein in urine
High levels may indicate kidney damage.
Causes of acidic urine (low pH)
Respiratory problems, dehydration, starvation, or a high protein diet.
Causes of basic urine (high pH)
Kidney disease, urinary tract infections (UTI), or a diet rich in vegetables/dairy.
What does green urine indicate?
Cause: Bacterial infection, green food dyes (asparagus), or certain diuretics
What does orange urine indicate?
Cause: Bilirubin from obstructive jaundice or certain drugs/carrots.
What does red/brown urine indicate?
Cause: Hemoglobin in the urine (various causes) or eating beets.
What does brown-black urine indicate?
Cause: Melanin pigment from melanoma (rare) or huge quantities of rhubarb.
Specific Gravity
A measure of urine density relative to water; normal range is typically 1.010–1.026.
Hydrometer
The tool used to measure the specific gravity of a urine sample.
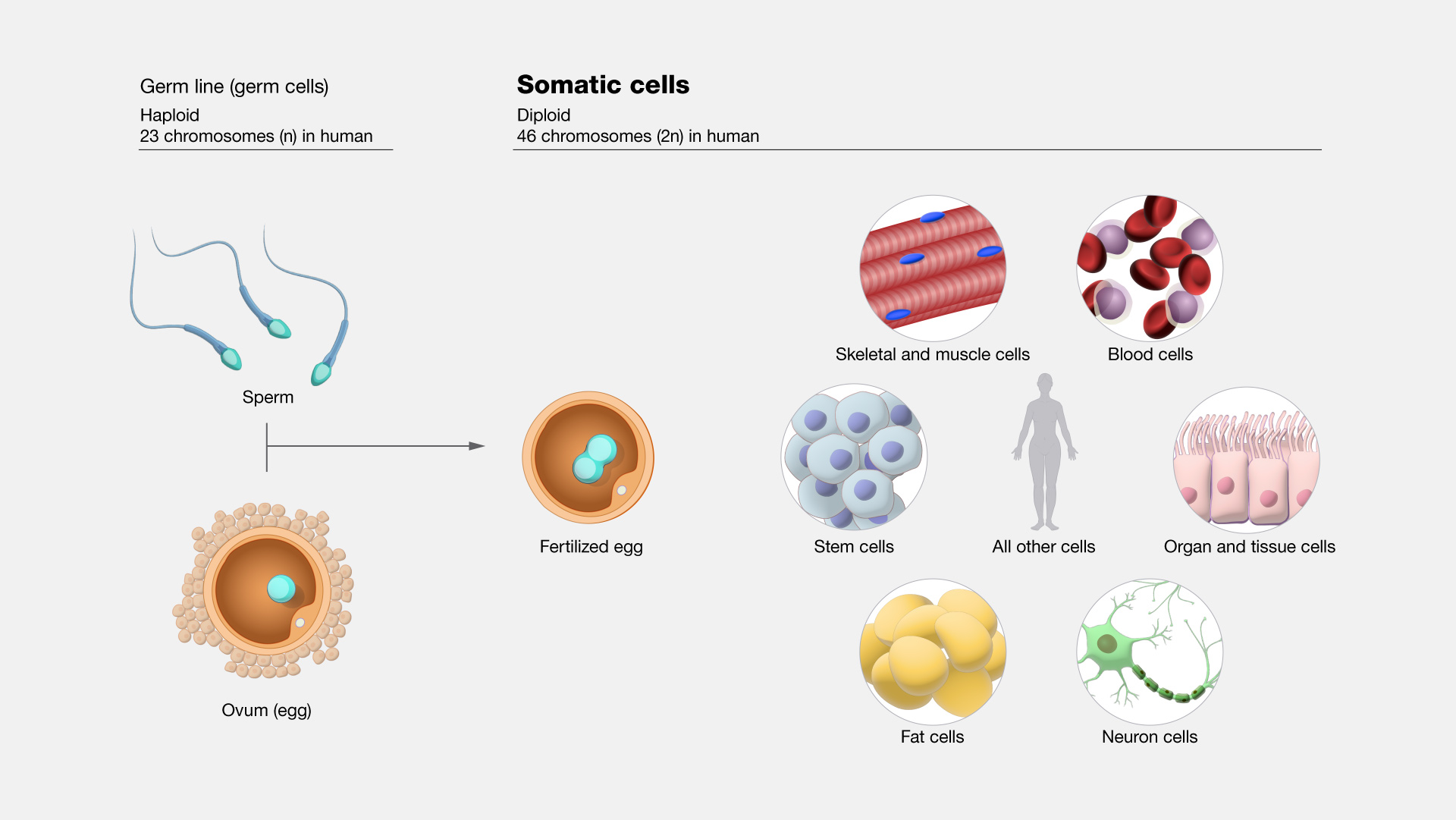
Somatic vs. Germ Cells
Somatic cells
body cells that divide via mitosis
Germ cells
produce gametes via meiosis
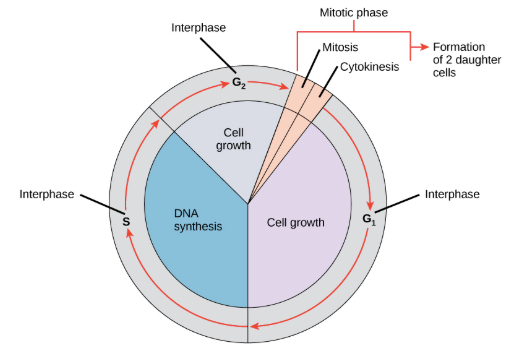
Interphase
The longest part of the cell cycle where the cell grows and replicates its DNA (S phase)
Prophase I (Mitosis)
Nuclear membrane breaks down, chromatin condenses, spindle forms and attaches to kinetochores
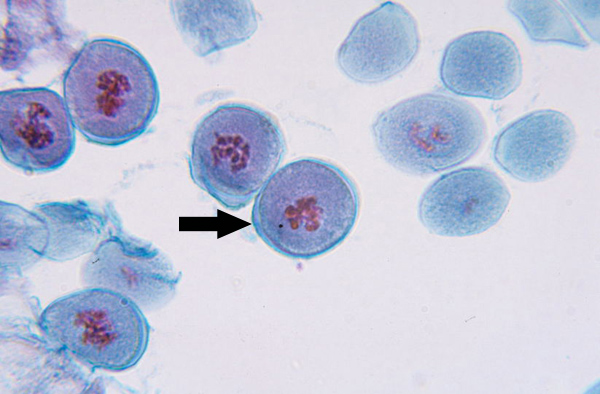
Metaphase I (Mitosis)
Microtubules align homologous chromosome pairs along metaphase plate
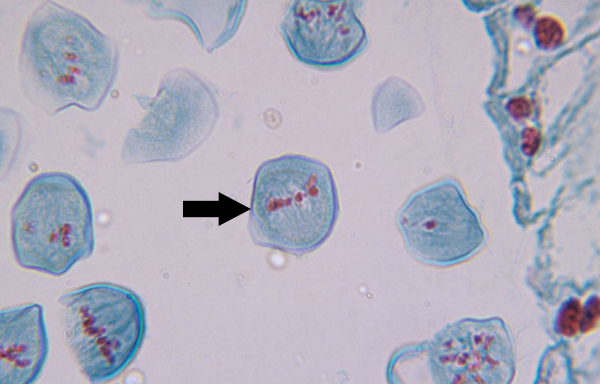
Anaphase I (Mitosis)
Kinetochore microtubules shorten, pulling homologous pairs to opposite poles, polar microtubules elongate, lengthening dividing cell
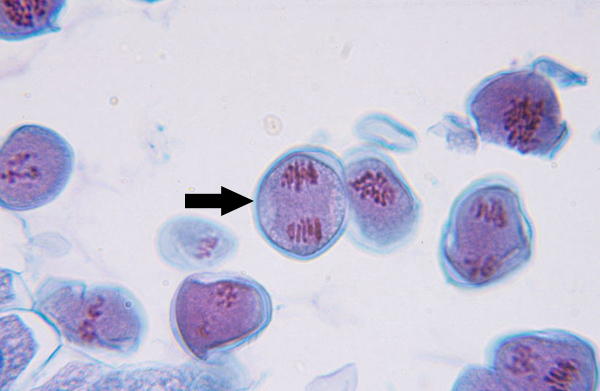
Telophase I (Mitosis)
Nuclear envelopes reform around the two sets of chromosomes, and the spindle disappears
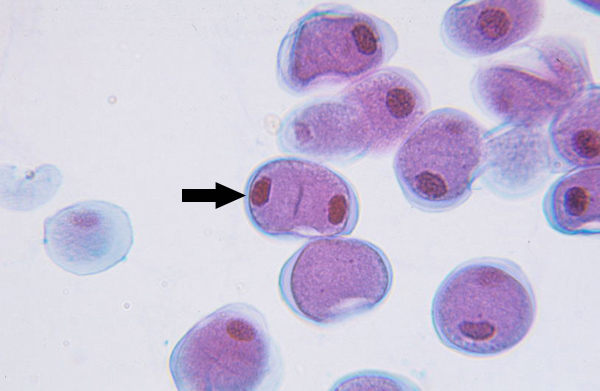
Cytokinesis I (meiosis)
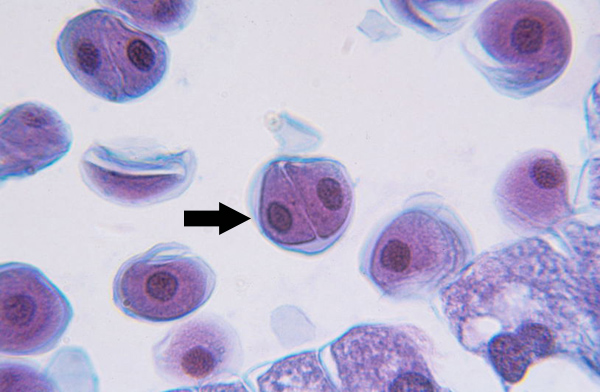
Prophase II (meiosis)
Nuclear membrane breaks down, chromatin condenses, mitotic spindle forms and attaches to kinetochores
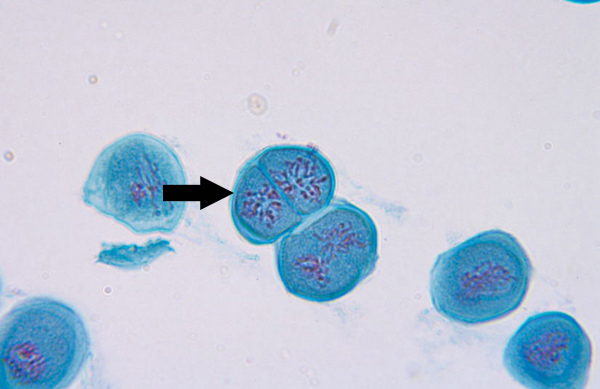
Metaphase II (meiosis)
Microtubules align chromosomes along metaphase plate
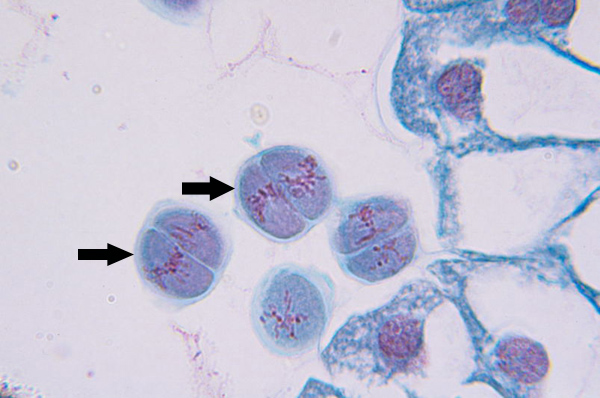
Anaphase II (meiosis)
Kinetochore microtubules shorten, pulling sister chromatids to opposite poles, polar microtubules elongate, lengthening dividing cell
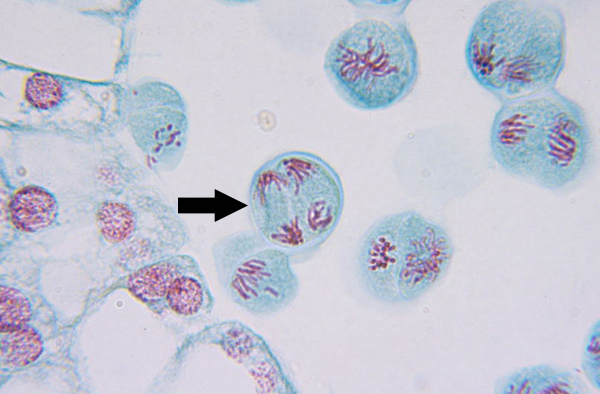
Telophase II (meiosis)
Nuclear membrane reforms, and chromatin decondenses, and cell plate begins to form
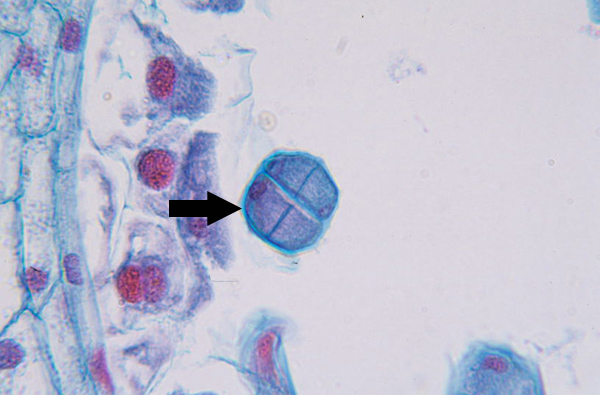
Cytokinesis II (meiosis)
Completed tetrad
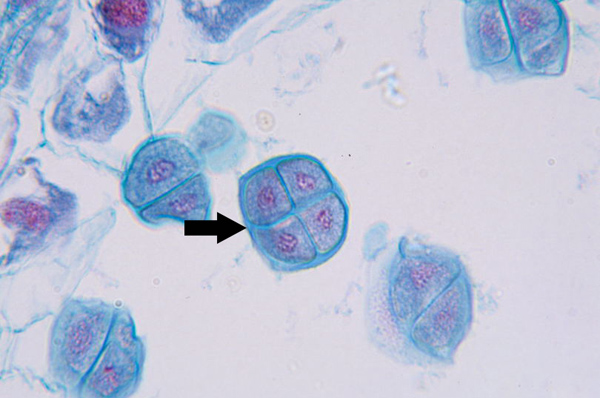
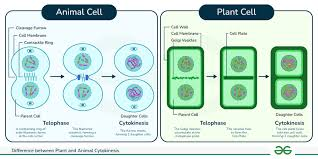
Cytokinesis in Plants vs. Animals
Animals form a cleavage furrow; plants form a cell plate due to their rigid cell wall.
Meiosis I unique features
Homologous chromosomes pair into tetrads, and crossing over occurs during Prophase I to increase genetic variation.
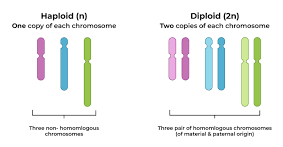
Haploid vs. Diploid
Diploid (2n) cells have two sets of chromosomes
Haploid (n) gametes (egg/sperm) have only one set.
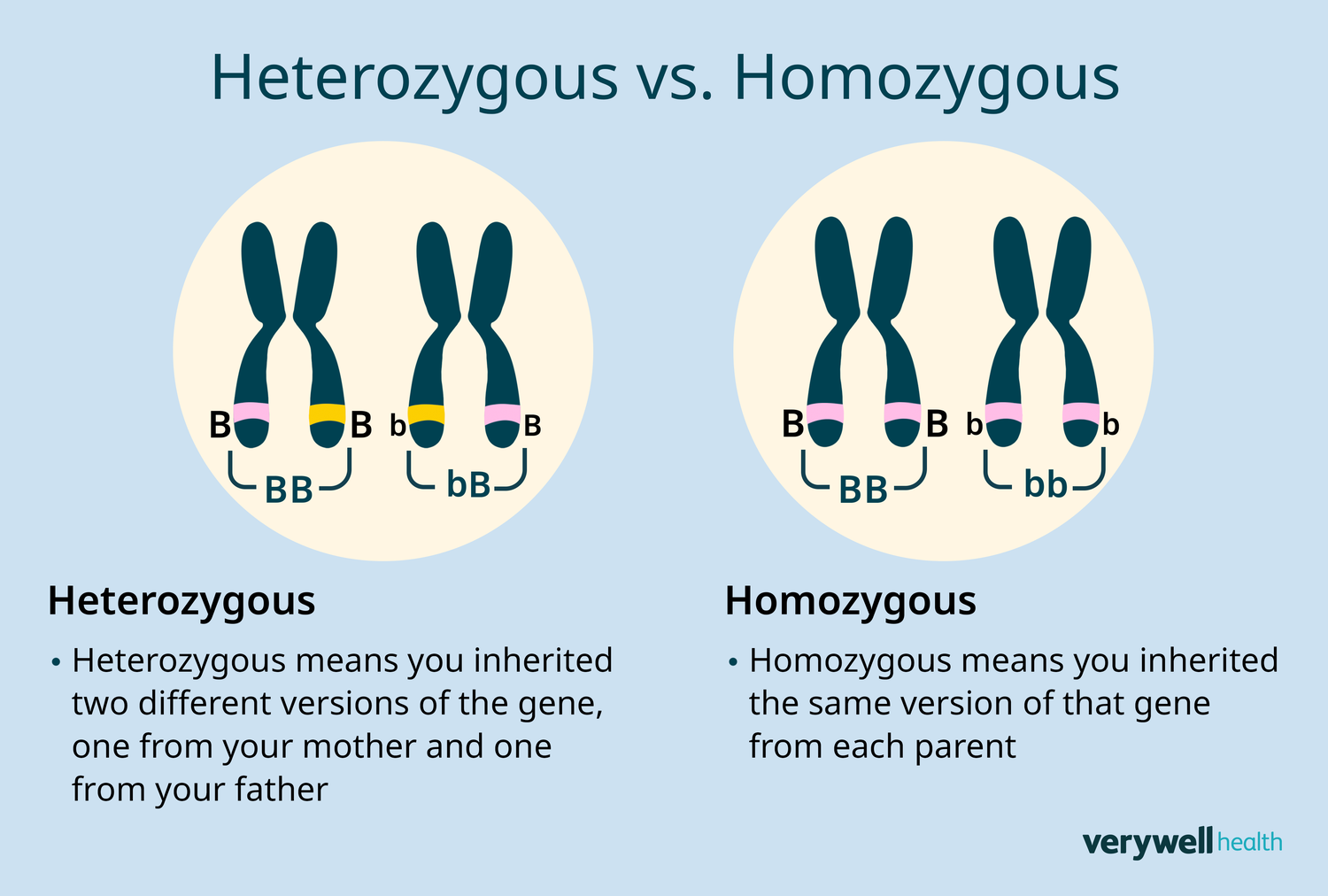
Heterozygous vs. Homozygous
Heterozygous (e.g., Tt) has two different alleles; homozygous (e.g., TT or tt) has two identical alleles.
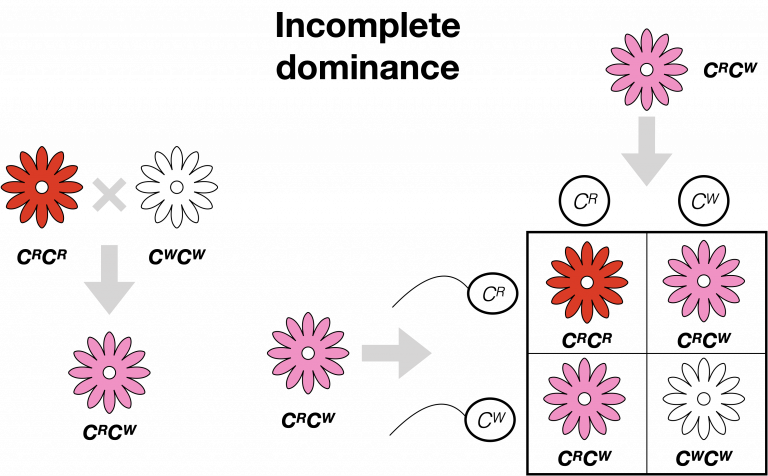
Incomplete Dominance
A form of inheritance where the offspring show an intermediate phenotype (e.g., black and white parents producing grey offspring).
Epistasis
When one gene (control gene) must be present for another gene to express its trait (e.g., purple color needs an "E" gene to show).
Test Cross
Mating an individual with a homozygous recessive parent to determine the unknown individual's genotype.
Linked Genes
Genes located on the same chromosome that are likely to be inherited together; identified when parental genotypes appear more frequently than recombinants.
Map Units
A measure of distance between genes on a chromosome based on how often crossing over occurs between them.
What is genetic transformation?
It is a process where a cell takes up and expresses a new piece of genetic material (DNA) from another organism, resulting in a "change caused by genes".
What organism is used as the host for transformation in the transformation pGLO lab?
Escherichia coli (E. coli), because it is a small, single-celled organism that reproduces quickly.
What is a plasmid?
A small, circular piece of DNA that is capable of self-replicating and often contains genes beneficial to bacterial survival.
What are the four key components of the pGLO plasmid?
GFP Gene: Encodes the Green Fluorescent Protein.
bla (or AmpR): Provides resistance to the antibiotic ampicillin.
araC: A regulator gene that turns on the GFP gene in the presence of arabinose.
ori: The origin of replication
What is the purpose of CaCl2 and "Heat Shock" in the procedure of the pGLO transformation lab?
CaCl2 (transformation solution) makes the cells "competent" to take up DNA, and the heat shock (42°C for 50 seconds) creates pores in the cell wall to allow the plasmid to enter.
Why is arabinose added to one of the plates in the pGLO lab?
Arabinose is a naturally occurring five-carbon sugar (aldopentose) found extensively in plant cell walls, often derived from sources like gum arabic, sugar beet pulp, and corn fiber. It is widely used as a low-calorie sweetener, a metabolic health supplement to inhibit sucrase activity, and a crucial tool in molecular biology for inducing gene expression in bacteria
acts as a switch to turn on (regulate) the expression of the GFP gene. Without it, the bacteria will have the gene but will not glow.
Predicted Results – Which plate(s) will show growth in the pGLO lab?
+pGLO LB/amp: Growth (white colonies).
+pGLO LB/amp/ara: Growth (fluorescent green colonies).
-pGLO LB: Growth (a "lawn" of bacteria)
Predicted Results – Which plate will glow under UV light in the pGLO lab?
Only the +pGLO LB/amp/ara plate, because it contains the GFP gene, the antibiotic resistance to survive, and the arabinose to trigger expression.
What is transformation, and what are the steps to perform E. coli transformation with pGLO plasmid?
Transformation is a genetic engineering process where bacteria uptake foreign DNA, such as a plasmid, changing their traits.
E. coli transformation with pGLO introduces a plasmid containing GFP (green fluorescent protein) and ampicillin resistance, enabling bacteria to grow on antibiotic plates and glow under UV light in the presence of arabinose sugar.
Key Steps to Perform E. coli Transformation (pGLO):
Prepare Cells (+pGLO/-pGLO): Label two microtubes "+pGLO" and "-pGLO" (control). Add \(250 \mu L\) of \(CaCl_{2}\) transformation solution to each and place on ice.
Add Bacteria and Plasmid: Pick a single colony of E. coli from a starter plate and mix into both tubes. Add the pGLO plasmid DNA specifically to the "+pGLO" tube.Incubate on Ice:
Incubate both tubes on ice for 10 minutes.
Heat Shock: Transfer the tubes to a \(42^{\circ}C\) water bath for exactly 50 seconds to make the membrane porous, then quickly return them to ice for 2 minutes.
Recovery: Add \(250 \mu L\) of LB (Luria-Bertani) nutrient broth to each tube and incubate at room temperature for 10 minutes.
Plating: Spread the samples onto LB agar plates (some containing ampicillin and arabinose).
Incubation: Incubate the plates upside down at \(37^{\circ}C\) for 24 hours.
Expected Results:
-pGLO on LB/Amp: No growth (bacteria killed by ampicillin).
+pGLO on LB/Amp: Bacterial colonies grow (resistant to ampicillin).
+pGLO on LB/Amp/Ara: Colonies grow and glow bright green under UV light (GFP expressed due to arabinose).
Features of a plasmid vector and why they are important?
Key Features of a Plasmid Vector
Origin of Replication (Ori): This is a specific DNA sequence where replication initiates, allowing the plasmid to replicate independently of the host's chromosomal DNA.
Importance: It dictates the plasmid's copy number (how many copies are present in a cell) and ensures the plasmid is passed on to daughter cells.
Selectable Marker (e.g., Antibiotic Resistance): A gene that allows for the selection of bacteria containing the plasmid, such as genes for resistance to antibiotics like ampicillin.
Importance: It ensures only bacteria holding the plasmid survive during the selection process, allowing for the identification of transformed cells.
Multiple Cloning Site (MCS) or Polylinker: A short, custom-designed region containing several unique recognition sites for different restriction enzymes.
Importance: It allows researchers to easily cut the plasmid and insert foreign DNA at a specific location, making it versatile for various cloning projects.
Promoter Region: A regulatory sequence that controls the transcription of the inserted gene, often driving expression.
Importance: It enables the host cell to transcribe the inserted gene into mRNA, which is essential if the goal is to produce a protein from that gene.
Small Size: Engineered to be small enough for efficient manipulation and high-copy amplification.
Importance: Smaller plasmids are easier to work with in the lab, easier to transfer into cells (transformation), and less likely to break (shear) during handling.
Summary of Importance
Gene Cloning: They act as vehicles (vectors) to carry and insert DNA sequences into cells.
Protein Production: They can contain promoters that drive high-level expression of proteins for therapeutic or industrial use.
Stable Maintenance: Their ability to replicate autonomously ensures they are passed down through generations of bacteria, providing a stable system for research.
Easy Manipulation: The small size and known sequence make them straightforward to modify.
How to analyze transformation results from plates and calculate transformation efficiency
1.Analyzing Transformation Plates
After an overnight incubation (usually 16–18 hours at \(37^{\circ}\text{C}\)), analyze the plates for efficiency and control check: [1, 2, 3]
Counting: Count the number of discrete colonies on the experimental plates (e.g., LB + antibiotic + DNA).
Tip: If there are too many, use a marker to dot each colony on the back of the plate to ensure accuracy.
Range: Ideal plates have 30–300 colonies for accurate counting.
Controls Check:
Positive Control (Cells + Known Plasmid): Should show high density growth, indicating competent cells worked, according to NEB.
Negative Control (Cells + No DNA): Should show no growth, confirming the antibiotic selection is effective and the cells were not contaminated.
2. Calculating Transformation Efficiency
The transformation efficiency represents the total number of bacteria transformed by \(1\ \mu\text{g}\) of DNA.
Key Data Required:
Colonies (CFU): Counted number of colonies.
Volume Plated (\(\mu \)L): Volume of suspension added to the plate.
Total Recovery Volume (\(\mu \)L): Total volume of cells + DNA + outgrowth medium (e.g., SOC/LB).
Amount of DNA : Mass of plasmid added, calculated as V*C (volume in muL x Concentration in mug/muL)
![<p><strong>1.Analyzing Transformation Plates</strong></p><p>After an overnight incubation (usually 16–18 hours at \(37^{\circ}\text{C}\)), analyze the plates for efficiency and control check: [1, 2, 3]</p><ul><li><p><span><strong>Counting:</strong> Count the number of discrete colonies on the experimental plates (e.g., LB + antibiotic + DNA).</span></p><ul><li><p><span><em>Tip:</em> If there are too many, use a marker to dot each colony on the back of the plate to ensure accuracy.</span></p></li><li><p><span><em>Range:</em> Ideal plates have 30–300 colonies for accurate counting.</span></p></li></ul></li><li><p><span><strong>Controls Check:</strong></span></p><ul><li><p><span><strong>Positive Control (Cells + Known Plasmid):</strong> Should show high density growth, indicating competent cells worked, according to NEB.</span></p></li><li><p><span><strong>Negative Control (Cells + No DNA):</strong> Should show no growth, confirming the antibiotic selection is effective and the cells were not contaminated.</span> </p></li></ul></li></ul><p><strong>2. Calculating Transformation Efficiency</strong></p><p>The transformation efficiency represents the total number of bacteria transformed by \(1\ \mu\text{g}\) of DNA. </p><p><strong>Key Data Required:</strong></p><ul><li><p><span><strong>Colonies (CFU):</strong> Counted number of colonies.</span></p></li><li><p><span><strong>Volume Plated (\(\mu \)L):</strong> Volume of suspension added to the plate.</span></p></li><li><p><span><strong>Total Recovery Volume (\(\mu \)L):</strong> Total volume of cells + DNA + outgrowth medium (e.g., SOC/LB).</span></p></li><li><p><span><strong>Amount of DNA :</strong> Mass of plasmid added, calculated as V*C (volume in muL x Concentration in mug/muL)</span></p></li></ul><p></p>](https://assets.knowt.com/user-attachments/6192f4e6-23bd-485c-b888-38f2fc17ba22.png)
What are restriction enzymes?
Special proteins (endonucleases) that recognize and cut DNA at specific sequences called restriction sites
What does PCR stand for and what is its goal?
Polymerase Chain Reaction; it is used to amplify (make many copies of) specific regions of DNA
What are the three steps of a PCR cycle?
1. Denaturation (94–96°C): Heat separates the double-stranded DNA.
2. Annealing (~68°C): Primers attach to the target DNA sequences.
3. Extension (~72°C): DNA polymerase (Taq) adds nucleotides to create a new strand.
What are STRs (Short Tandem Repeats)?
Non-coding regions of DNA with repeating units 2–6 base pairs long. The number of repeats varies between individuals, creating a unique profile.
How does gel electrophoresis separate DNA fragments?
It uses an electrical current to move negatively charged DNA toward the positive electrode (anode). The agarose gel matrix acts as a sieve, allowing smaller fragments to move faster and further than larger ones.
What is a DNA Marker (or Ladder)?
A set of DNA fragments of known sizes used as a reference to determine the size of unknown experimental bands in base pairs (bp).
How is DNA visualized on a gel?
Since DNA is colorless, it must be stained with dyes like Fast Blue or Ethidium Bromide and viewed under a transilluminator (white or UV light).
What is CODIS?
The Combined DNA Index System, a database maintained by the FBI that uses 13 standard STR loci for human identification.
Looking at the pGLO plasmid map, what is the role of the araC region?
The araC gene acts as a regulatory "switch". In the presence of the sugar arabinose, the araC protein promotes the binding of RNA polymerase, turning on the expression of the GFP gene so the bacteria will glow.
You observe a plate labeled +pGLO LB/amp that has many white colonies but does not glow under UV light. Is this result expected? Why?
Yes, this is expected. The bacteria have the amp resistance gene and can grow in the presence of ampicillin, but because there is no arabinose on this plate, the GFP gene is not "turned on," and they will not glow.
(Plate Analysis) You observe a plate labeled -pGLO LB/amp with zero growth. Why?
These bacteria did not receive the pGLO plasmid (the "selectable marker" for ampicillin resistance). Therefore, they are susceptible to the antibiotic ampicillin and cannot grow.
(Plate Analysis) What visual difference do you expect between a "lawn" and "colonies"?
Lawn: A continuous, thick layer of bacteria covering the agar surface (seen on the -pGLO LB control plate where growth is unchecked).
Colonies: Distinct, individual circular spots of growth, each originating from a single transformed cell.
(Gel Analysis) On an agarose gel, in which direction does DNA migrate and why?
DNA is negatively charged and migrates toward the positive electrode (the red anode). On a gel image, the wells are at the top (negative/black end), and DNA moves toward the bottom.
(Gel Analysis) Which DNA fragments will be found closest to the wells at the top of the gel?
The largest (longest) fragments. Because they move more slowly through the agarose matrix, they travel the shortest distance from the wells
How do you determine the size of an unknown DNA band on a gel?
By comparing its migration distance to a DNA Ladder (Marker) in an adjacent lane. The ladder contains fragments of known sizes (e.g., a 200 bp ladder) that serve as a visual reference
(PCR Analysis) Looking at a diagram of the PCR cycle, what happens during the "Annealing" step (~68°C)?
DNA primers attach (anneal) to the specific target sequences on the single-stranded DNA. This provides a starting point for DNA polymerase to begin replication.
(Case Analysis) How do you visually identify a "match" in a DNA fingerprinting gel?
You look for identical banding patterns. A suspect's DNA profile must have bands that align exactly in size and position with the DNA fragments found at the crime scene.
(STR Analysis) In an STR profile, what is the visual difference between a homozygous and a heterozygous individual at one locus?
Homozygous: Shows only one band because both inherited alleles (maternal and paternal) are the same size.
Heterozygous: Shows two distinct bands because the alleles inherited from each parent are of different sizes.
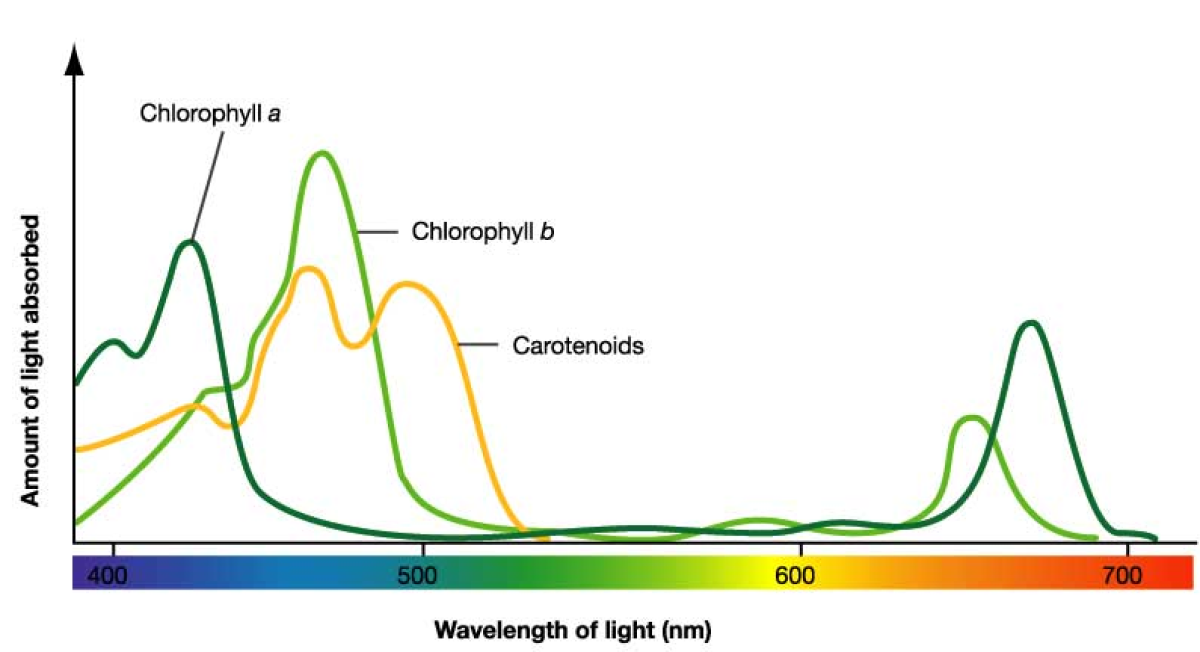
Which colors/wavelengths do chlorophylls absorb most effectively?
Blue-violet and red wavelengths; they reflect green, which is why plants appear green.
What is a Zygote?
The single cell formed when an egg and sperm fuse together.
Define Cleavage.
The process where cells undergo rapid mitosis to form more cells.
What is a Blastula?
A hollow ball of cells formed as cells rearrange themselves after cleavage.
Describe Gastrulation.
A stage where cells reorganize and migrate (often invaginating to form a tube), leading to the formation of germ layers.
What is Organogenesis?
The process by which specialized tissues and organs form.
Name the 6 early developmental stages of a starfish in order.
1. Unfertilized egg (haploid),
2. Fertilized egg (diploid/zygote),
3. Cleavage,
4. Morula (solid ball of cells),
5. Blastula (hollow ball),
6. Gastrula.
What are the three germ layers formed during mid-development?
Ectoderm (outer), Mesoderm (middle), and Endoderm (inner).
What structures arise from the Ectoderm?
The outer surface (epidermis) and the central nervous system (neurons of the brain/peripheral system).
What structures arise from the Mesoderm?
The notochord, skeletal muscle cells, kidney tubules, and red blood cells.
What are Somites?
Groups of mesoderm cells that develop into specific structures like the cartilage of vertebrae/ribs, back muscles, and skin of the back.
What structures arise from the Endoderm?
Internal structures like the pharynx, respiratory tube, pancreas, thyroid, and lung cells.
What is the purpose of the Amnion?
It surrounds the embryo, forming the amniotic sac.
What is the function of the Yolk Sac in humans?
It serves as a source of germ cells and blood cells.
What does the Chorion contribute to?
It contributes to the formation of the placenta.
What structures does the Allantois form?
The umbilical cord and the urinary bladder.
What is PTC?
Phenylthiocarbamide, a bitter-tasting chemical not normally found in food.
What is the difference between Phenotype and Genotype in the PTC lab?
Phenotype is the observable trait (taster or non-taster), while genotype is the specific genetic makeup (e.g., TT, Tt, or tt).
Which gene is responsible for PTC tasting?
The TAS2R38 gene, which encodes a G protein-coupled receptor (GPCR)
Where are taste buds located?
Inside bumps on the tongue called papillae.
What are the 5 steps performed in the PTC lab to determine genotype?
1. Taste PTC paper,
2. Extract DNA (from cheek cells),
3. PCR amplification,
4. Restriction digest (RFLP),
5. Gel electrophoresis.