Antigen Recognition Genetics
1/19
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
20 Terms
Describe the difference in antigen recognition between B/T Cells
B vs T:
Antigen Types:
B:
Outside the cell
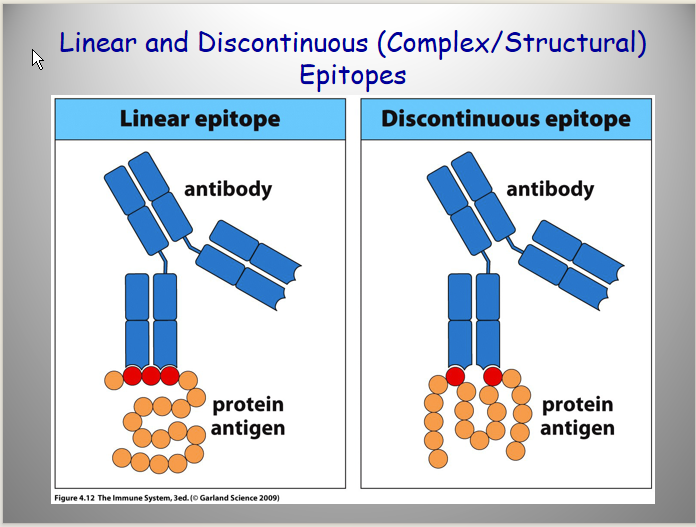
linear peptides or complex 3D; or any kind of molecules
T:
Primarily protein antigens
linear peptides
Presentation:
T cells must be presented an antigen by APCs on MHC
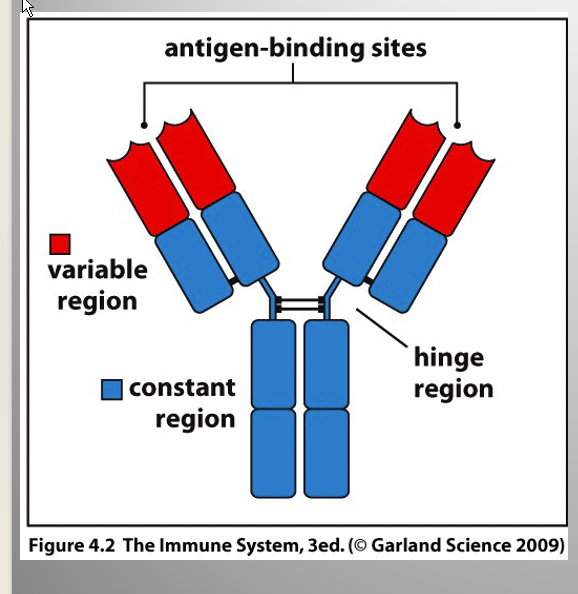
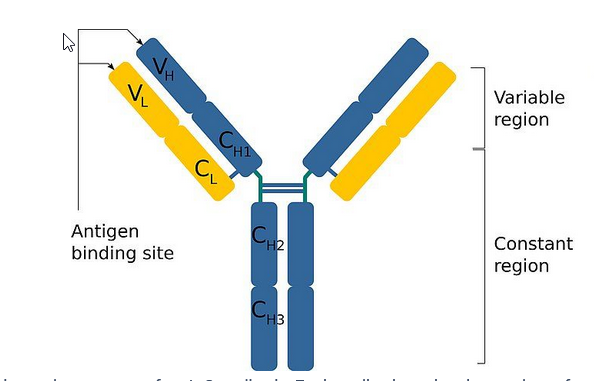
Describe the structures of antibodies polypeptides
Describe the different regions
How is an antibody isotype class determined?
Define:
Isotypes
Allotypes
Idiotypes
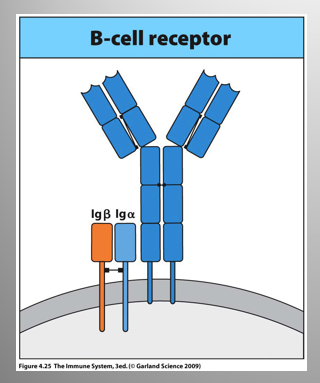
What is the difference between a B Cell Receptor (BCR) and the secreted Ig (sIg)
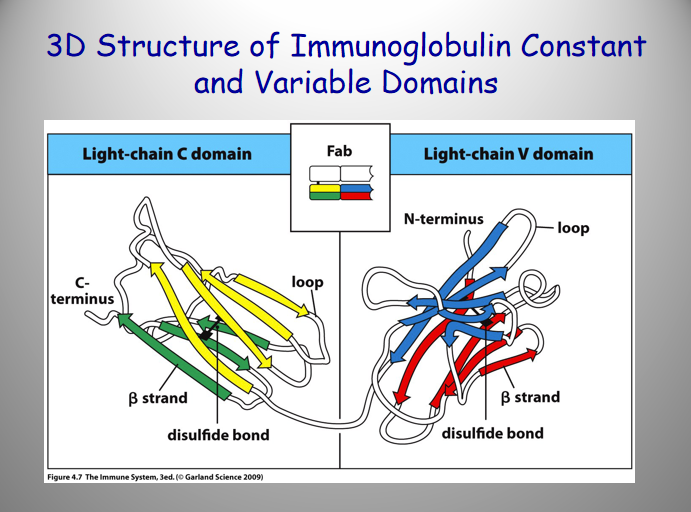
Regions:
Variable Region:
higher degree of variation in amino acid sequence
[] @ N terminal end
Constant Region:
limited degree of variation btw dif. antibodies
Hinge region:
Flexible
relatively unstructured portion in the middle
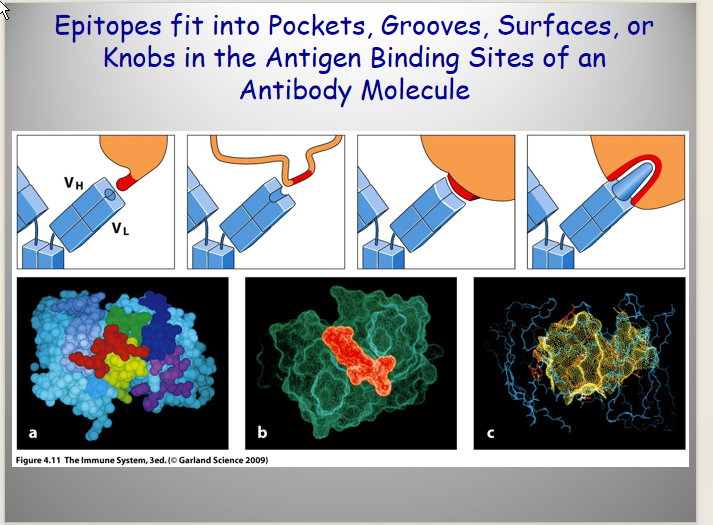
Antigen binding site:
formed by V regions of L and H chains
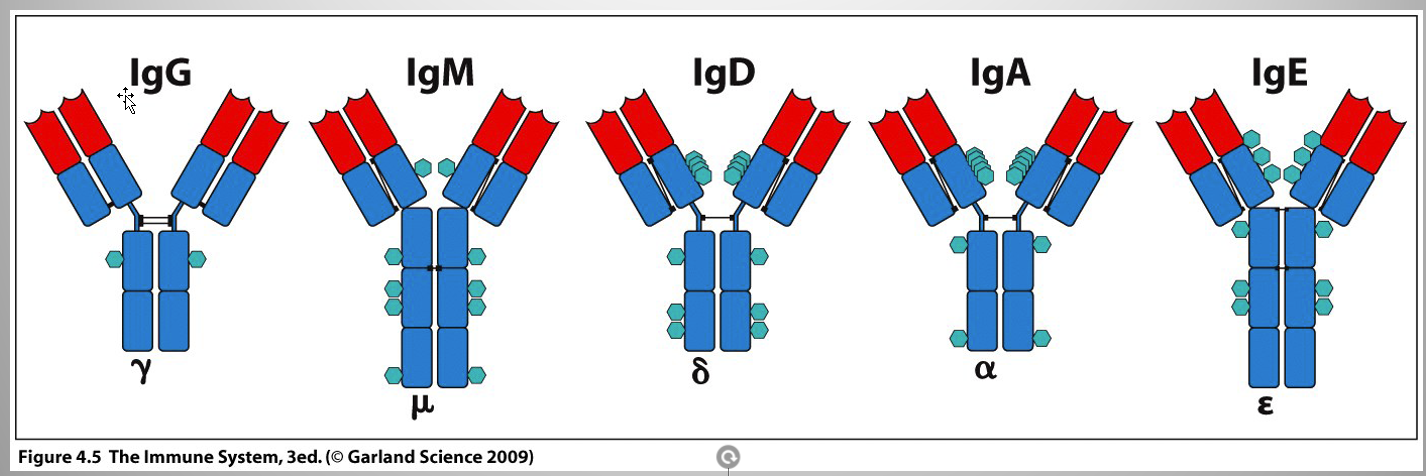
Determination of Ig(antibody) isotypes
Dependent on Heavy chain C regions
α, δ, γ: 3 domains
μ, ε: 4 C domains
extra domain = elongated hinge region
For Light Chains:
Either Kappa or Lambda in all types
No funcitonal differences
Definitions:
Isotypes:
same thing as antibody classes (IgM, IgD, IgG, IgE, and IgA)
Allotypes:
genetically determined differences in antibodies between people
Idiotypes
Antibodies of different specificity found within the same individual due to the diversity of the Ig V region
BCR vs sIg:
same specificity
Difference:
one is membrane-bound; other = secreted.
BCR signaling = mediated by two small proteins w/ long cytoplasmic tails: Iga and Igb
AKA CD79a and b.
Describe the V domain
Framework region
Hypervariable region
V Domain:
Framework Regions:
Flanks the hypervariable regions
Much less variable
Hypervariable regions:
Regions w/in V domain where it is super variable
AKA CDRs (Complementarity-determining regions)
b/c provide a binding surface that is complementary to that of antigen.
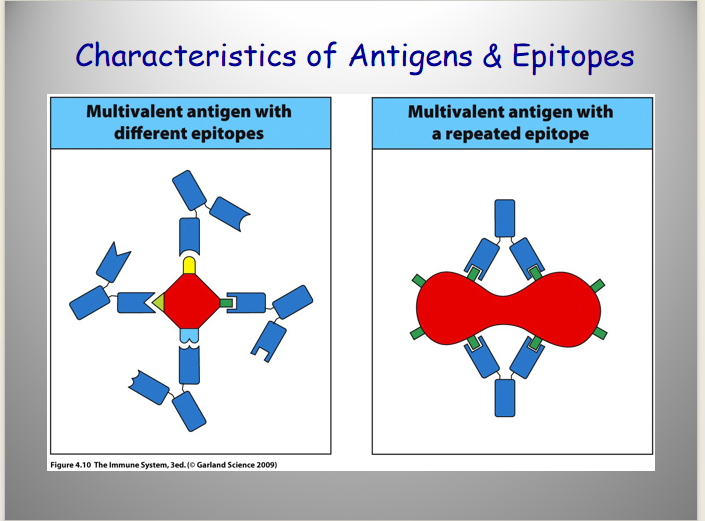
Describe the Characteristics of B Cells:
Antigen Specificity
Antigen Repertoire
B Cells Characteristics:
Antigen Specificity:
each B cell is specific for only one epitope of an antigen.
Antibody Repertoire:
number of different antibodies/B cells recognizing different epitopes that an individual can generate.
Describe how the mechanisms to generate B cell Diversity
List the Different Domains
Describe Rearrangement/Recombination Mechanism:
What is RSS? Components?
what is the 12/23 rule
What does RAG do?
Describe Junctional Diversity
3 components?
Resolving
End result
The Different Domains:
V = variable region gene segment
D = diversity region gene segment (only in Ig heavy chain genes)
J = joining region gene segment
C = constant region gene segment
Rearrangement/Recombination Mechanism:
RSS: (Recombination Signal Sequences)
Special DNA motifs that directs recombination site
Components:
Heptamer(7) region:
Always adj. to Variable Region’s Coding Sequence
Nonamer(9) Region:
Spacer Region:
Separates Heptamer and Nonamer regions
Either 12 or 23 bps
12/23 Rule:
RSS w/ 12bp spacer interacts only w/ RSS w/ 23bp spacers
ensure gene segments are joined in correct order
RAG1 & RAG2 (Recombinase Activating Genes 1 & 2)
Mediate V-(D)-J Joining
Rags attach to the RSS of the V/J regions
They join together and Cut the DNA
The VJ segments gets joined together (Coding Joint) → Retained in genome
everything originally between the V/J gets jointed together (Signal Joint)→ floats away
Junctional Diversity
3 Components:
Junctional flexibility:
RAGs induced double-strand cuts → creates DNA Hairpin
P-nucleotide addition:
Another enzyme (Artemis, act. by DNA-PK) cuts near end of RSS → allows the Non-coding Strand (bottom strand) to join the Coding Strand (top strand)
N-nucleotide addition:
TdT (Terminal deoxy-nucleotidyl transferase) (lymphocyte specific) → adds random # of nucleotides @ end of RSS (after P addition)
Resolving the Junction:
Pairing of Strands
Unpaired nucleotides = removed by exonuclease
Gaps are filled by DNA synthesis and ligation to form coding joint
End Result:
Some = non-productive
accidentally created “stop” codons or introducing frameshifts in the J region
Some = Productive
will generate increased diversity in the third hypervariable region (CDR3).
Describe what happens if there is Absent or Defective RAG Proteins
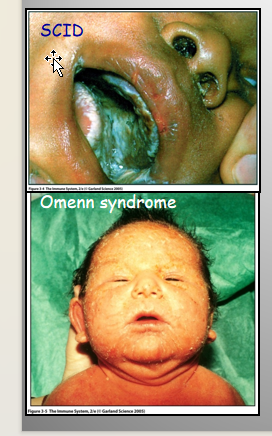
Absent or Defective RAG Proteins:
SCID
T and B cells are absent
Omen syndrome
Chronic inflammation from auto-reactive T cells
Treatment: Bone Marrow Transplant
Describe Allelic Exclusion
Describe Clonal Selection
Allelic Exclusion
After productive rearrangement of one heavy chain and one light chain gene → recomb. of other allele does not proceed.
result in a B cell becoming monospecific to only one antigen
Clonal Selection
B cell binds its cognate antigen → signaling effect → proliferation to generate enough effector cells to fight
Describe the naive B cell
which Ig does it express? how?
Difference between Secreted vs membrane Ig
Describe class Switchin
Occurs when?
Describe the difference in the primary vs secondary immune response; Difference between the two Igs?
Describe the genetic contents for the Ig heavy chain
Mechanism?
Describe Somatic Hypermutation:
Mediated by?
Mech
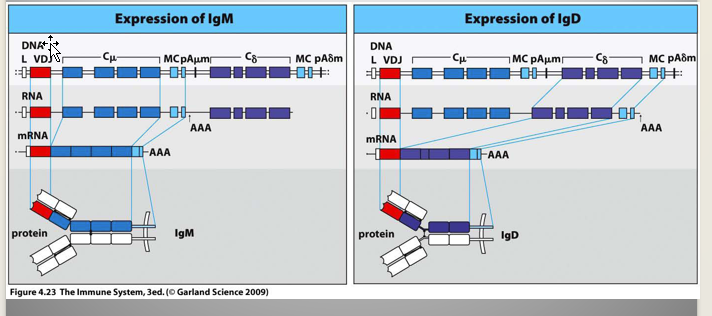
Naive B Cells:
Express both Surface IgM & IgD
Expression of both isotype accomplished by alternative mRNA splicing
***NOTE: The constant region for IgM and IgD is close together; so after rearrangement; that’s why they’re both expressed together***
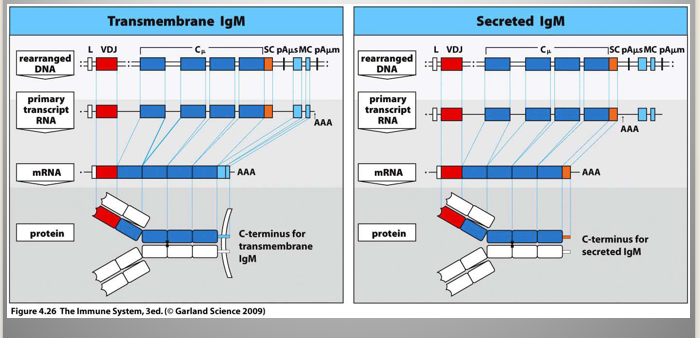
Secreted vs membrane Ig:
same selectivity; only genetic difference is the C-terminus
Class Switching
Occurs AFTER Antigen Exposure
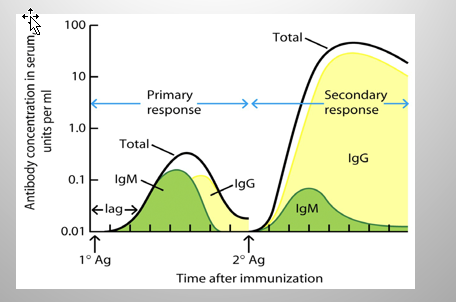
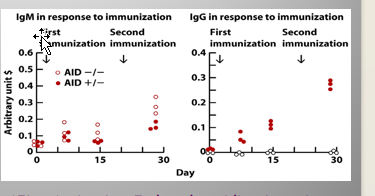
Primary vs Secondary Response:
Primary: Mostly IgM
Secondary: Mostly IgG
Difference:
IgG = same specificity/heavy chain VDJ and light chain VJ gene segments as IgM
ONLY THE CONSTANT REGION HAS CHANGED!!
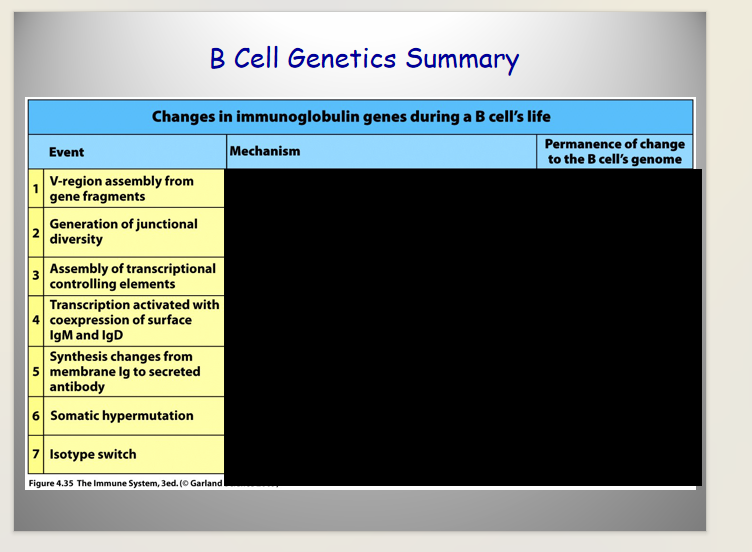
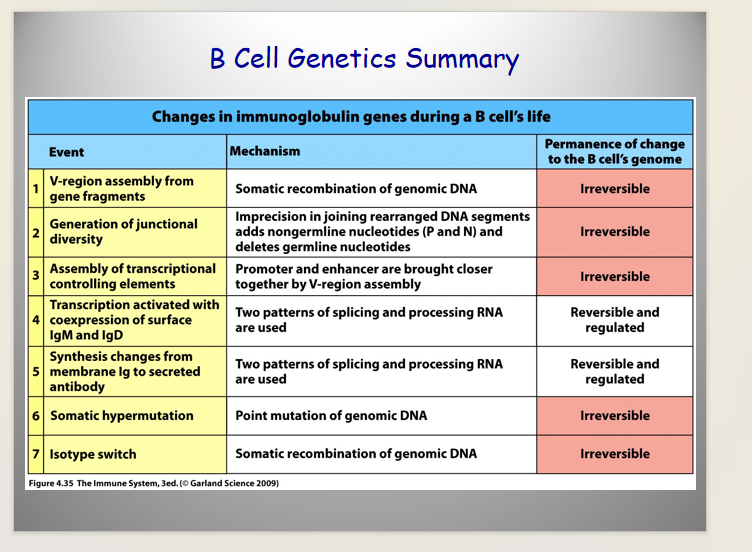
Genetics of Ig heavy Chain
has Multiple Constant Domain Regions (determining different types of heavy chains)
ORDER matters in isotype switching
Mech:
Mediated by AID (Activation-Induced cytidine deaminase)
AID binds to 2 Switch regions
Areas located before each specific constant region
One = orgiinal; the other = the new isotype it wants to become
The aid brings the Switch region together →
cut everything in the middle
only the new constant region remains (along its switch region; the switch region of the Original is also gone)
After Switching = no switching back (b/c that region is lost)
(you can still switch if you have some other constant regions after the New one though)
Somatic Hypermutation:
Mediated also by AID
Mech: Introducing Point Mutation ONLY IN V REGION → alters Ab affinity
C + AID → U + repair → A,C,G, or T


Describe the CD3 -T Cell Receptor (TCR) Complex
Function
Deficiencies:
CD3γ or CD3ε
CD3δ / CD3ζ
Function
CD3 complex proteins transmit signals to cell after TCR binds peptide/MHC
Deficiencies:
CD3γ or CD3ε deficiency
Low TCR numbers
Impaired signal transduction
CD3δ / CD3ζ deficiency
T cells absent
Normal B cells
NK cells: impaired cytolytic function
What are teh two types of T Cell Receptor
Describe γδ TCells
Percentage?
Location?
Interact w/
Recognize/Triggered by?
γδ vs αβ?
γδ Chains?
Two types:
γδ
αβ
γδ Tcells:
1-5% of circulating T cells
Dominant in gut mucosa, epithelial tissues and skin epidermis
Interact with nonclassical MHC class 1-like CD1 receptors
May recognize microbial phospholipid antigens
May be triggered by “danger” signals
γδ TCRs = less diversity than αβ TCRs
Fewer V gene segments
Most γδ T cells express same γδ TCR chains.
Restricted TCR = pattern recognition receptor
Draw out how an γδ or αβ cell can be made
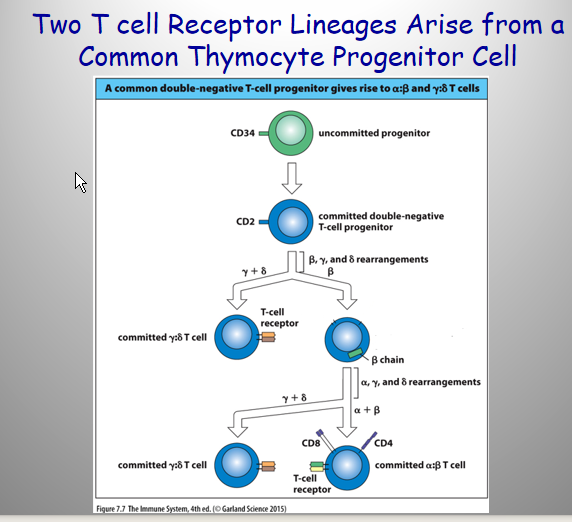
Describe the Mechanism to make either an γδ or αβ cell
Mech:
Thymocytes Try to Correctly Rearrange a b-Chain (Similar to the Ig Heavy Chain) (V joined by D/J)
Rearrangement of the a-Chain Gene Only Occurs in Pre-T cells
IF FAIL → γδ = default
If SUCCESS → no going back to γδ
Compare and Contrast T and B cell receptors
Similarites
T vs B:
membrane/Secreted
Ag Binding
Both Has:
Variable and constant regions
Complementary determining regions (CDRs)
Gene rearrangements → variability
Associated membrane-bound signaling molecules
T vs B:
Membrane/Secreted:
T: Membrane-bound (receptor) only
B: Membrane-bound or secreted
Ag binding:
T: No further changes
Only recognize antigen in MHCs
B: Class Switching/Somatic hypermutation