Population Subdivision
1/31
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
32 Terms
population subdivision entails an inbreeding like effect
decrease in the proportion of heterozygous genotypes
a sub-structured population has 3 levels of complexity
individual organisms (I)
subpopulations (S)
total population (T)
HI
the heterozygosity of an individual in a population
Hs
the expected heterozygosity of an individual in an equivalent random mating subpopulation
HT
the expected heterozygosity of an individual in an equivalent random mating total population
HI can be interpreted as
the average actual observed heterozygosity of all the genes/loci in an individual, or as the probability of heterozygosity of an one gene/locus
Hs represents the
level of heterozygosity that would be found in a subpopulation if the subpopulation were undergoing random mating, therefore Hs always equals 2piqi for a subpopulation with allele frequency pi
Hs always equals
2piqi for a subpopulation with allele frequency pi
HT represents what the
heterozygosity would be, were all sub populations pooled together and mated randomly, if the average allele frequency among subpopulations is p0 then HT equals 2p0q0
if the average allele frequency among subpopulations is p0 then
HT equals 2p0q0
what is the inbreeding coefficient
FIS
the inbreeding coefficient FIS measures
the reduction in heterozygosity of an individual due to random mating within its subpopulation
what is the equation for measuring heterozygosity of an individual due to nonrandom mating within its subpopulation
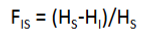
what is the equation for the reduction in heterozygosity of a subpopulation due to genetic drift
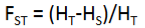
what is the equation for the reduction in heterozygosity of an individual relative to the total population
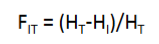
FIS is concerned with
inbreeding within individuals
FST is concerned with
inbreeding in subpopulations
FIT is concerned with
inbreeding in individuals relative to the total population
what is the most inclusive measure of inbreeding that takes into account both effects of nonrandom mating and effects of population subdivision
FIT
what is the mathematical relationship between FIS/FST/FIT
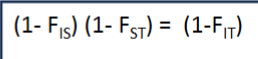
FIS in natural populations is
typically close to 0
when FIS=0 then
FST=FIT
the fixation index FST is a good estimate of
total inbreeding coefficient
FST is used as a measure of
genetic difference among populations
FST is influenced by
random genetic drift
migration
natural selection
mutation
FST minimum = 0 then
no genetic divergence
FST maximum = 1.0 then
fixation for alternative alleles in subpopulations
FST = 0-0.05 then
little genetic differentiation
FST = 0.05-0.15 then
moderate genetic differentiation
FST = 0.15-0.25 then
great genetic differentiation
FST = >0.25 then
very great genetic differentiation
as random drift occurs from generation to generation
the value of F will change (increase)