AP Biology - UNIT 6 ASSESSMENT
1/72
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
73 Terms
(DNA REPLICATION) The Molecular Basis of Inheritance - STRUCTURE OF DNA (revisited)
What is DNA?
Deoxyribonucleic acid is a coiled double helix carrying hereditary information of the cell (contains the instructions for making polypeptides from 20 different amino acids)
(DNA REPLICATION) The Molecular Basis of Inheritance - STRUCTURE OF DNA (revisited)
What is DNA wrapped around?
DNA is wrapped around histone proteins (nucleosomes) and super-coiled in order to be condensed down enough to fit into the nucleus. Appears as chromatin when cell not dividing.
(DNA REPLICATION) The Molecular Basis of Inheritance - STRUCTURE OF DNA (revisited)
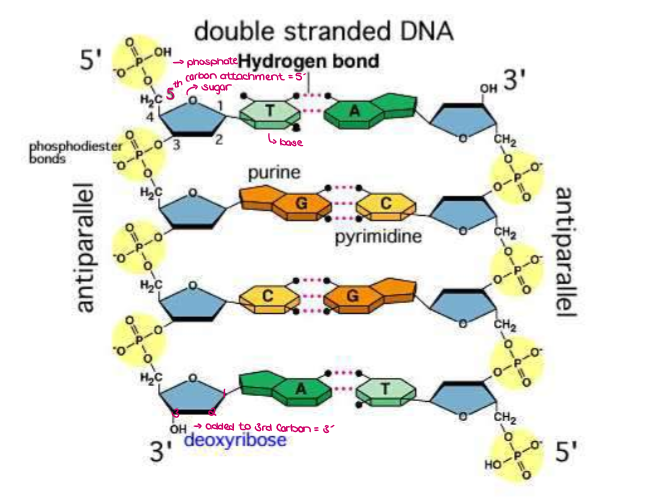
What are the “sides” of the double helix made of?
The “sides” of the double helix are made of pentose (5-sided) sugars (deoxyribose) attached to phosphate groups by phosphodiester bonds.
(DNA REPLICATION) The Molecular Basis of Inheritance - STRUCTURE OF DNA (revisited)
What are the “steps” or “rungs” of DNA made of?
The “steps” or “rungs” of DNA made of 4 nitrogen-containing bases held together by weak hydrogen bonds
(DNA REPLICATION) The Molecular Basis of Inheritance - STRUCTURE OF DNA (revisited)
What are PURINES?
Purines (double carbon-nitrogen rings) include adenine (A) and guanine (G)
(DNA REPLICATION) The Molecular Basis of Inheritance - STRUCTURE OF DNA (revisited)
What are PYRIMIDINES?
Pyrimidine (single carbon-nitrogen rings) include thymine (T) and cytosine (C)
(DNA REPLICATION) The Molecular Basis of Inheritance - STRUCTURE OF DNA (revisited)
What is BASE PAIRING?
Base pairing means a purine bonds to a pyrimidine (Example: A — T and C — G)
Chargoff’s Rule: purine always bonds with pyrimidine
(DNA REPLICATION) The Molecular Basis of Inheritance - STRUCTURE OF DNA (revisited)
Subunits/monomers of DNA are called ___________. Nucleotides contain a _________, a ___________ _____, and ___ ________ ____ (_, _, _, or _)
nucleotides
phosphate
deoxyribose sugar
one nitrogen base
A
T
C
G
(DNA REPLICATION) The Molecular Basis of Inheritance - STRUCTURE OF DNA (revisited)
What are ANTIPARALLEL STRANDS?
Antiparallel strands refers to: 2 strands run in opposite directions…
5’ → 3’ direction of one strand and the 3’ → 5’ direction of the other.
(DNA REPLICATION) The Molecular Basis of Inheritance - STRUCTURE OF DNA (revisited)
Which side are nucleotides added to?
All nucleotides can only be added to the 3’ end.
you can never add to the 5’ end
(DNA REPLICATION) Some important experiments involving DNA: The Hershey-Chase experiment was designed to answer one key question: What is the genetic material—DNA or protein?
What was the PURPOSE?
The experiment aimed to determine which molecule carries genetic information into cells during infection…nucleic acids or proteins?
(DNA REPLICATION) Some important experiments involving DNA: The Hershey-Chase experiment was designed to answer one key question: What is the genetic material—DNA or protein?
What did they test?
Used bacteriophages (viruses (protein nugget w/o cellular membrane w/ DNA) that infect bacteria → they inject their DNA into the cell, the rest of the capsule falls away, the DNA can then do its work)
phages = bacterial
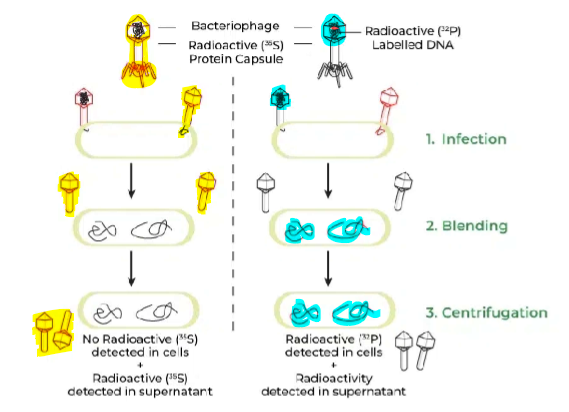
Labeled:
DNA with radioactive phosphorus (32P) → since P is only found in nucleic acids (not in proteins)
Protein with radioactive sulfur (35S) → since S is only found in proteins (not in DNA)
(DNA REPLICATION) Some important experiments involving DNA: The Hershey-Chase experiment was designed to answer one key question: What is the genetic material—DNA or protein?
What is the KEY IDEA?
If DNA is the genetic material → it should enter the bacteria
If protein is the genetic material → protein should enter the bacteria
(DNA REPLICATION) Some important experiments involving DNA: The Hershey-Chase experiment was designed to answer one key question: What is the genetic material—DNA or protein?
What is the CONCLUSION?
DNA entered the bacterial cells, while protein did not!
Therefore…
DNA (not protein) is the molecule that stores and transmits genetic information.
(DNA REPLICATION) Some important experiments involving DNA: The Meselson-Stahl experiment was designed to answer another fundamental question about DNA: How does DNA replicate?
What was the PURPOSE?
To determine whether DNA replication is:
Conservative → original DNA stays intact
Semiconservative → each new DNA has one old strand + one new strand
Dispersive → old and new DNA mixed throughout
(DNA REPLICATION) Some important experiments involving DNA: The Meselson-Stahl experiment was designed to answer another fundamental question about DNA: How does DNA replicate?
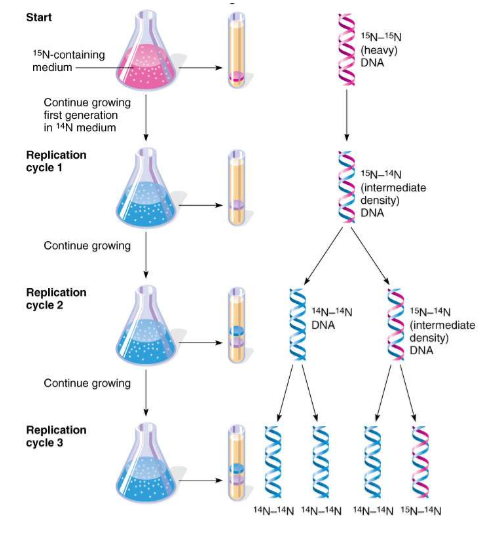
What did they do?
Grew bacteria in heavy nitrogen (15N) → DNA became “heavy”
Then moved bacteria to light nitrogen (14N)
Tracked DNA over generations using density (centrifugation)
(DNA REPLICATION) Some important experiments involving DNA: The Meselson-Stahl experiment was designed to answer another fundamental question about DNA: How does DNA replicate?
Draw the processes for CONSERVATIVE, SEMI-CONSERVATIVE, & DISPERSIVE.
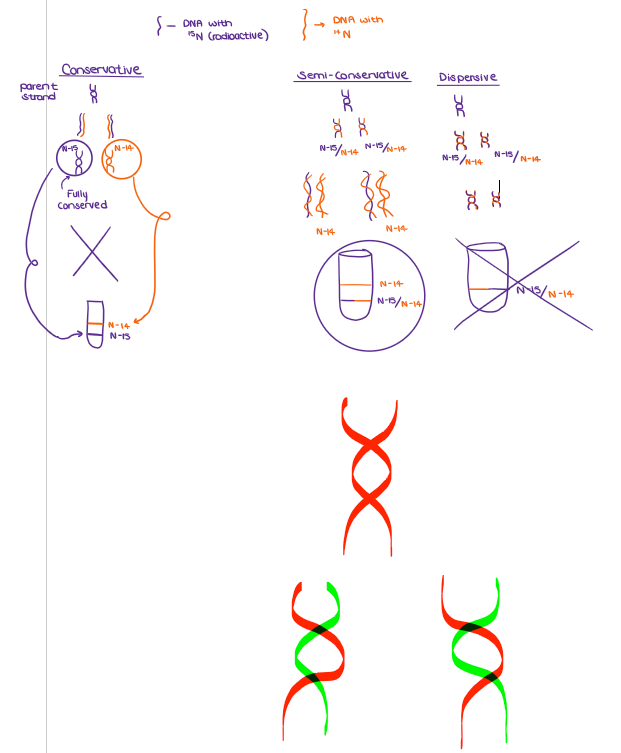
(DNA REPLICATION) Some important experiments involving DNA: The Meselson-Stahl experiment was designed to answer another fundamental question about DNA: How does DNA replicate?
What are the KEY RESULTS?
After 1 replication: DNA was intermediate (hybrid)
After 2 replications: DNA showed both hybrid and light DNA
(DNA REPLICATION) Some important experiments involving DNA: The Meselson-Stahl experiment was designed to answer another fundamental question about DNA: How does DNA replicate?
What was the CONCLUSION?
DNA replicates semi-conservatively
Each new DNA molecule contains:
One original (parental) strand
One newly synthesized strand
= ½ old/parent strand + ½ newly synthesized
(DNA REPLICATION) DNA REPLICATION
What is DNA REPLICATION?
Process by which DNA makes a copy of itself
Occurs during S phase of interphase before cell division
(There are ~6 billion nucleotide pairs in humans!)
(DNA REPLICATION) Semiconservative Model
What is the SEMICONSERVATIVE MODEL?
Each parent strand serves as the template for each new strand; ½ old - ½ new
(DNA REPLICATION) Semiconservative Model
Explain the REPLICATION BUBBLE.
Replication begins at particular sites along the DNA called ORIGINS OF REPLICATION (there are hundreds to thousands in eukaryotic DNA)
The DNA opens at these point, creating a REPLICATION BUBBLE (the more bubbles, the faster replication can occur)
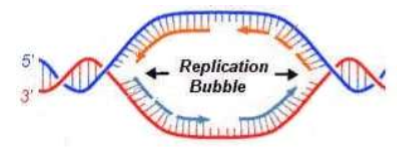
Replication proceeds in both directions until the molecule is copied
At each end of a replication bubble is a REPLICATION FORK, a Y-shaped region where the parental strands are being unwound
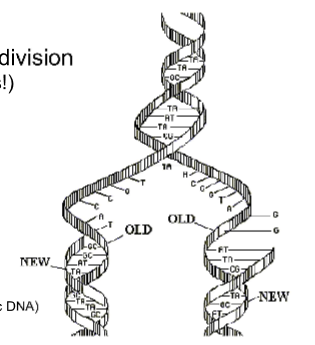
(DNA REPLICATION) Getting Started
Explain the process of DNA REPLICATION.
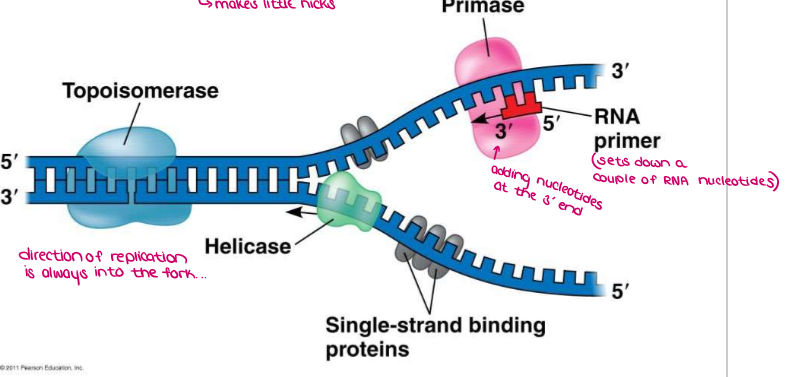
HELICASE unwinds and separates the two parental strands (unzips the H-bonds)
SINGLE STRAND BINDING PROTEINS (SSBP) stabilize the unpaired DNA. (Topoisomerase prevents the strain of re-twisting → makes little nicks)
Primase makes a short RNA PRIMER to start the replication process (DNA can’t initiate the synthesis, nucleotides can only be added to the end of an already existing chain)
→ FLATTENED, OPENED UP, RNA PRIMER ADDED
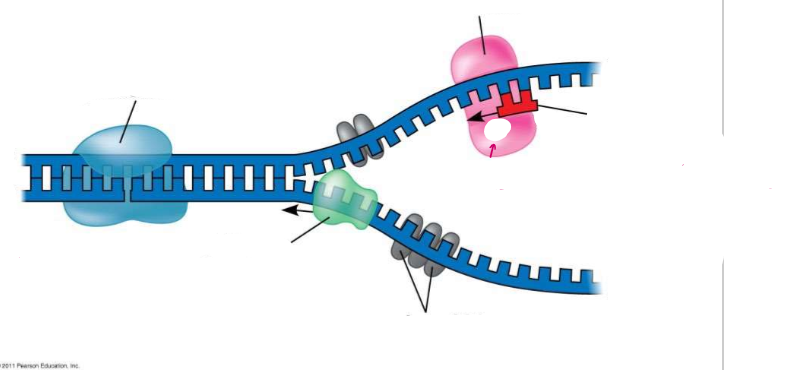
(DNA REPLICATION) Getting Started
Label the diagram of DNA replication.
(DNA REPLICATION) ELONGATION
What is the role of DNA POLYMERASE III?
DNA Polymerase III adds complementary nucleotides to the growing chain using the parental DNA strand as a template. *Nucleotides can only be added to the free 3’ end of the growing DNA strand! New DNA strands are always synthesized 5’ → 3’.
(DNA REPLICATION) ELONGATION
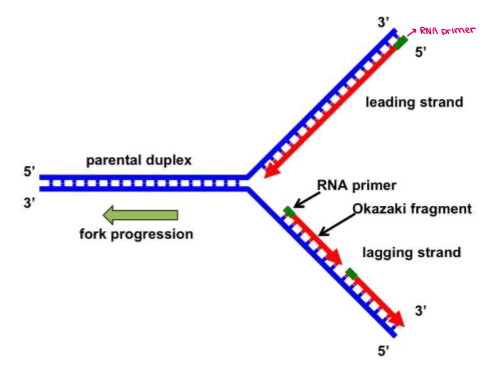
What is the LEADING STRAND?
The DNA strand that can be made continuously in the 5’ → 3’ direction (from the 3’ → 5’ template)
(DNA REPLICATION) ELONGATION
What is the LAGGING STRAND?
The DNA strand that is synthesized discontinuously in a series of OKAZAKI FRAGMENTS
(DNA REPLICATION) THE LEADING STRAND
Explain its role.
Helicase unwinds & separates parental double helix.
SSBP stabilize unwound strands.
Leading strand is synthesized continuously in the 5’ → 3’ direction by DNA pol III.
(nucleotides can only be added at the 3’ end!)
(DNA REPLICATION) THE LAGGING STRAND
Explain its role.
Primase makes several short RNA primers.
DNA pol III adds DNA nucleotides to the primer, forming Okazaki fragment (5’ — 3’)
After reaching the next RNA primer to the right, DNA pol III detaches and moves to the next primer.
Okazaki fragment 2 is synthesized until it reaches the first fragment.
DNA POLYMERAGE I cuts out the primers and replaces the RNA with DNA.
DNA LIGASE forms bonds the two fragments together. [splicing → cuts + gluing pieces together]
The lagging strand is complete in this region, and continues until entire strand is complete.
![<ol><li><p>Primase makes several short RNA primers.</p></li><li><p>DNA pol III adds DNA nucleotides to the primer, forming Okazaki fragment (5’ — 3’)</p></li><li><p>After reaching the next RNA primer to the right, DNA pol III detaches and moves to the next primer.</p></li><li><p>Okazaki fragment 2 is synthesized until it reaches the first fragment.</p></li><li><p><strong><u>DNA POLYMERAGE I</u></strong> <strong>cuts out the primers and replaces the RNA with DNA.</strong></p></li><li><p><strong><u>DNA LIGASE</u> forms bonds the two fragments together. </strong>[splicing → cuts + gluing pieces together]</p></li><li><p>The lagging strand is complete in this region, and continues until entire strand is complete.</p></li></ol><p></p>](https://assets.knowt.com/user-attachments/32647c2b-07ed-40e5-8e8d-c88d4fd7b1f5.png)
(DNA REPLICATION) DNA Replication Summary
Summarize the process!
Extreme rapid and accurate (only 1 in a billion are incorrectly paired)
Requires many enzyme & ATP (energy)
DNA polymerase III adds new nucleotides to the exposed bases in the 5’ to 3’ direction (Leading strand made continuously, Lagging strand made in Okazaki fragments and put together with DNA ligase)
DNA polymerase I replaces RNA primers with DNA nucleotides
DNA polymerase proofreads the new DNA checking for errors & repairing them; called excision repair
Helicase & SSBP come off and the two strands recoil, leaving 2 identical DNA molecules (each with an “old” strand and a “new” strand)
(DNA REPLICATION) DNA Replication Summary
How does DNA polymerase use the structure of DNA to catch errors?
DNA polymerase moves along a single strand of DNA, building the complementary strand as it goes. The two stranded molecule passes through the DNA polymerase molecules after synthesis is compelte. If the wrong base is inserted, then the bond is unstable. Because the double strand is passing through the DNA polymerase the missing base can be detected and replaced.
(DNA REPLICATION) DNA Replication Summary
What are NUCLEASES?
NUCLEASES are like genetic scissors that cut out the fault DNA.
(DNA REPLICATION) DNA Replication Summary
What is a THYMINE DIMER?
A thymine dimer is common mutation — it is a pair of abnormally chemically bonded adjacent thymine bases in DNA (T-T), resulting from damage by ultra-violet irradiation
(DNA REPLICATION) Telomerase/TELOMERE Function
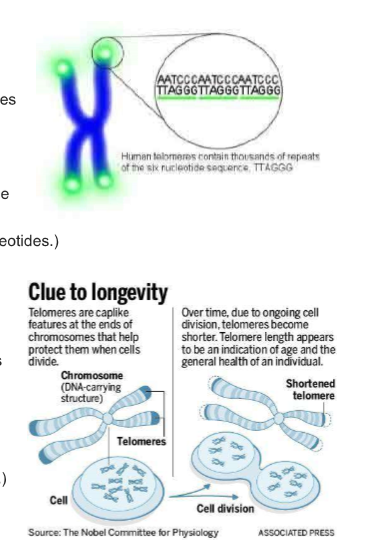
What are TELOMERES?
Telomeres are found at the end of chromosomes. They are made of repeating sequences of TTAGGG on one strand of DNA (bound to AATCCC on the other strand).
Cell division is needed so we can grow new skin, blood, bone and other cells when needed. Each time a cell divides, an average person loses 30 to 200 base pairs from the ends of that cell’s telomeres. (Since DNA pol III can’t add DNA bases from 3’ → 5’, when the RNA primer comes off of the DNA at 5’ → 3’ end, there is no way to “fill in” those nucleotides.)
The 3’ end of linear chromosomes are left with an “overhang.” Telomerase adds new DNA to the longer strand of the telomere overhang.
In human blood cells, for example, the length of telomeres ranges from 8,000 base pairs at birth to 3,000 base pairs as people age and as low as 1,500 in elderly people. (An entire chromosome has about 150 million base pairs.) Cells normally can divide only about 50 to 70 times, with telomeres getting progressively shorter until the cells become senescent, die or sustain genetic damage that can cause cancer. (Telomeres do not shorten with age in tissues such as heart muscle in which cells do not continually divide.)
Without telomeres, the main part of the chromosome - the part containing genes essential for life - would get shorter each time a cell divides. So telomeres allow cells to divide without losing genes. Without telomeres, chromosome ends would degrade the cell’s genetic blueprint, making the cell malfunction, become cancerous or die.
(GENE EXPRESSION) “ZOOMING IN” ON THE GENETIC MATERIAL
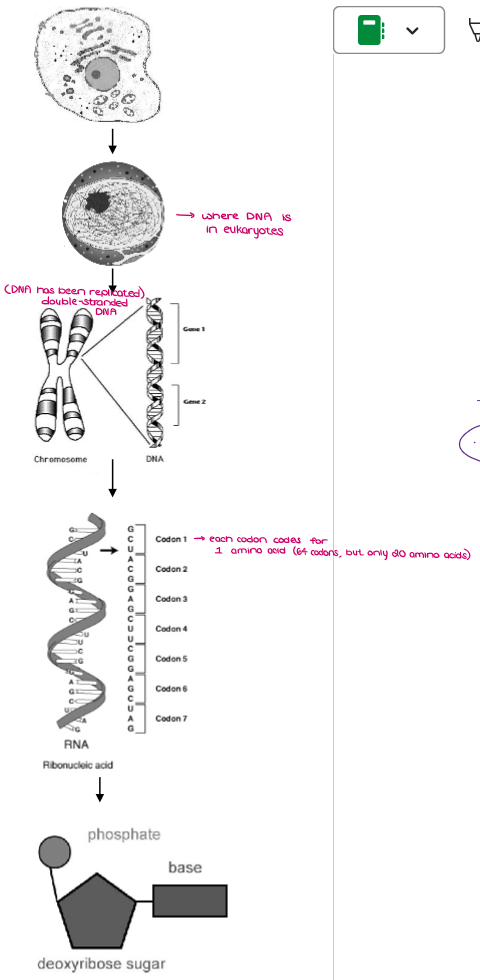
Zoom in on a CELL to a NUCLEOTIDE.
CELL
↓
NUCLEUS
↓
CHROMOSOMES (made of DNA)
↓
GENE (100s-1000s of genes on each chromosome; Specific sequence of nucleotides that codes for a polypeptide)
↓
TRIPLET (DNA)…CODON (mRNA) (Every 3 nucleotides code for 1 amino acid)
↓
NUCLEOTIDE (Consists of a phosphate group, deoxyribose sugar, nitrogenous base)
(GENE EXPRESSION) “ZOOMING IN” ON THE GENETIC MATERIAL
Draw the diagram of a PROKARYOTE.
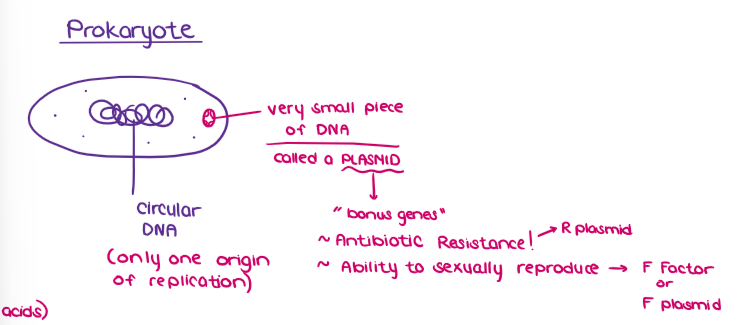
(GENE EXPRESSION) HOW DNA GETS “ORGANIZED” INTO CHROMATIN
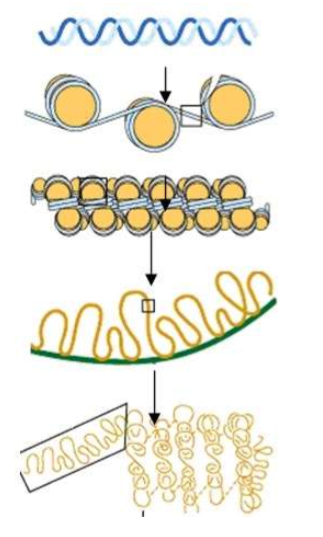
Explain this process.
DNA is a Double Helix → The DNA gets wrapped around HISTONE proteins forming NUCLEOSOMES. → The DNA continues to get coiled down into CHROMATIN FORM. (Euchromatin (active) vs. Heterochromatin (super coiled down))
(GENE EXPRESSION) One Form of Controlling Gene Expression
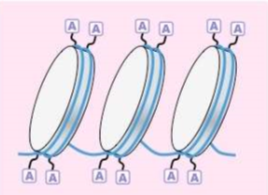
What is EUCHROMATIN?
is active DNA that can be transcribed (and translated)
Gene expressing
DNA is acetylated (acetyl groups are CH2-CH3 added to the histones, which opens up regions on DNA for Enzymes to attach)
Less condensed
(GENE EXPRESSION) One Form of Controlling Gene Expression
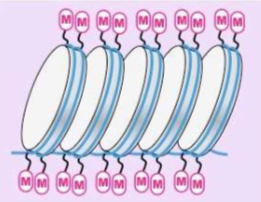
What is HETEROCHROMATIN?
is not active DNA and not transcribed
DNA is “methylated” (methyl groups -CH3 are added to histones making it impossible for transcription to occur)
Silenced genes (methylated)
More condensed
(GENE EXPRESSION) One Form of Controlling Gene Expression
What is CHROMATIN?
loosely coiled DNA (not dividing)
(GENE EXPRESSION) One Form of Controlling Gene Expression
What is ACETYLATION?
Regions with high transcriptional actively are loosely packed
(GENE EXPRESSION) One Form of Controlling Gene Expression
What is METHYLATION?
Regions with low or no transcriptional activity are densely packed
(GENE EXPRESSION) Template Strand vs. Coding Strand Notes
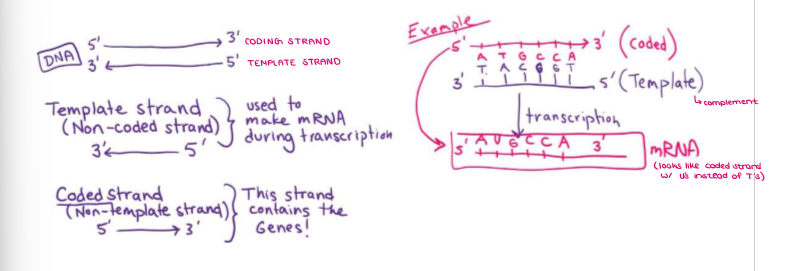
Remember that DNA strands are ANTIPARALLEL. What is the CODING STRAND?
One strand is the CODING strand (5’ → 3’), which contains the genes in the correct order.
(GENE EXPRESSION) Template Strand vs. Coding Strand Notes
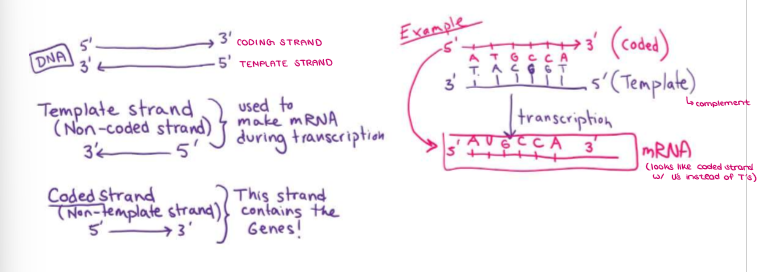
Remember that DNA strands are ANTIPARALLEL. What is the TEMPLATE STRAND?
The other is the TEMPLATE strand (3’ → 5’), which is used to make mRNA in transcription.
(GENE EXPRESSION) “GENE EXPRESSION”
What is GENE EXPRESSION?
“GENE EXPRESSION” refers to the processes: TRANSCRIPTION & TRANSLATION
These processes take our stored genetic code and make our genes usable!
Though these two processes, numerous proteins are created, and those proteins carry out the functions and give us our phenotypic traits.
(GENE EXPRESSION) One Gene - One Polypeptide Hypothesis
What is the ONE GENE - ONE POLYPEPTIDE HYPOTHESIS?
This theory proposes that each gene is responsible for the synthesis of a single polypeptide. It was originally stated as the “One gene-one, one-enzyme Hypothesis” by the US geneticist Georg Beadle in 1945 but later modified when it was realized that genes also encoded ALL polypeptides for ALL proteins (not just enzymes).

(GENE EXPRESSION) One Gene - One Polypeptide Hypothesis
Label the diagram of GENE EXPRESSION.

(GENE EXPRESSION) “GENE EXPRESSION”
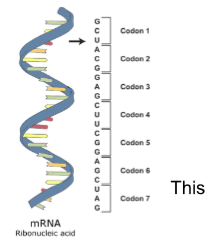
What is TRANSCRIPTION?
TRANSCRIPTION is the process of creating messenger RNA (mRNA) from the segment of DNA of our chromosomes (usually one gene).
(GENE EXPRESSION) “GENE EXPRESSION”
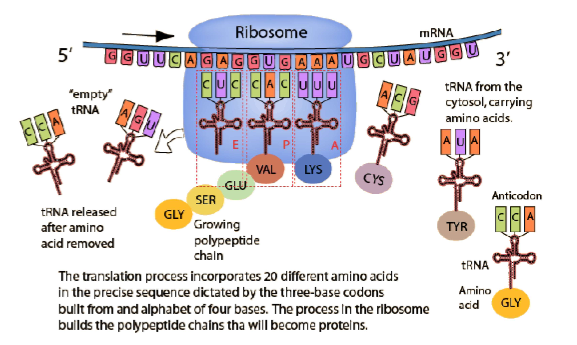
What is TRANSLATION?
TRANSLATION is the process of “reading” the mRNA and putting together the correct sequence of amino acids to make a polypeptide. This process occurs in the cytoplasm, specifically, on a ribosome.
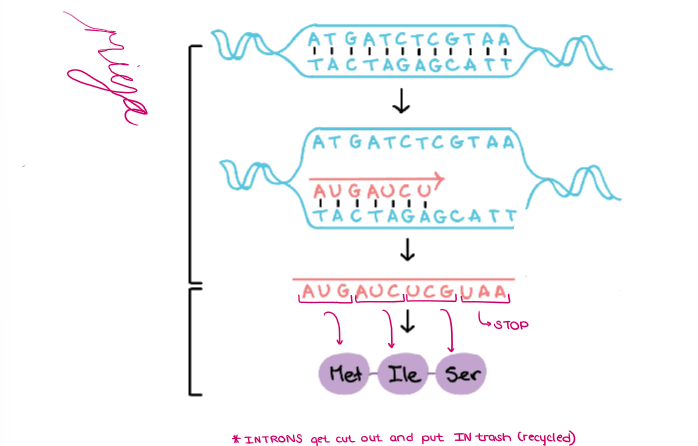
(GENE EXPRESSION) “GENE EXPRESSION”
Label the diagram of TRANSCRIPTION and TRANSLATION.
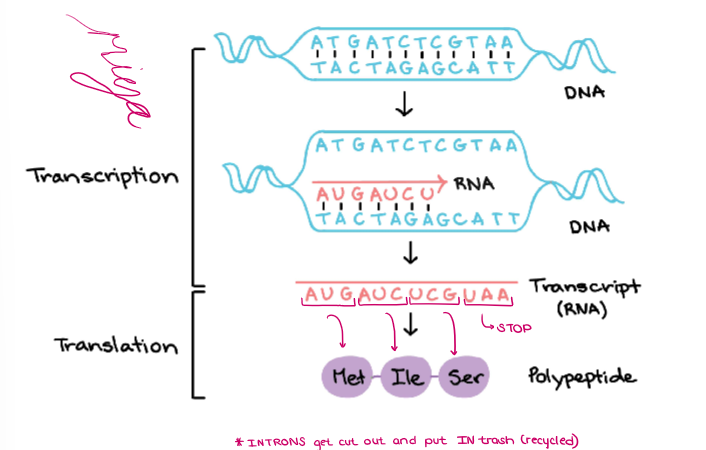
(GENE EXPRESSION) “GENE EXPRESSION”
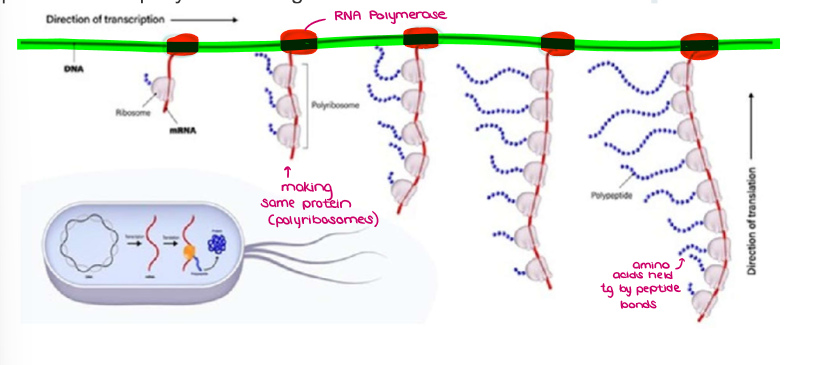
Explain gene expression in PROKARYOTES.
In prokaryotes, both processes occur simultaneously in the cytoplasm (there is no separation since they don’t have nuclei)
In prokaryotes, multiple RNA polymerases can transcribe a single bacterial gene while numerous ribosomes concurrently translate the mRNA transcripts into polypeptides (POLYRIBOSOMES). In this way, a specific protein can rapidly reach a high concentration in the bacterial cell.
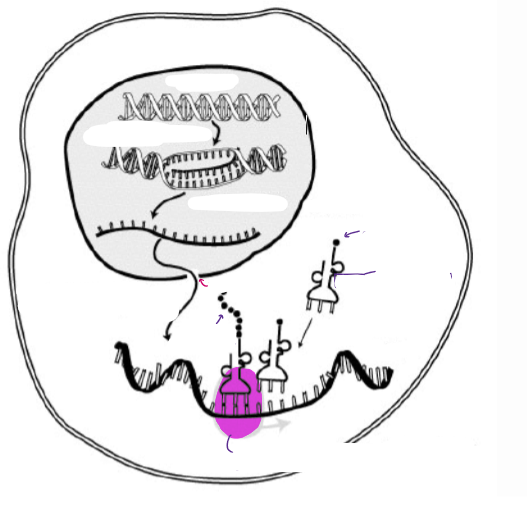
(GENE EXPRESSION) “GENE EXPRESSION”
IN EUKARYOTES, THE TWO PROCESSES ARE SEPARATED. Label the diagram.
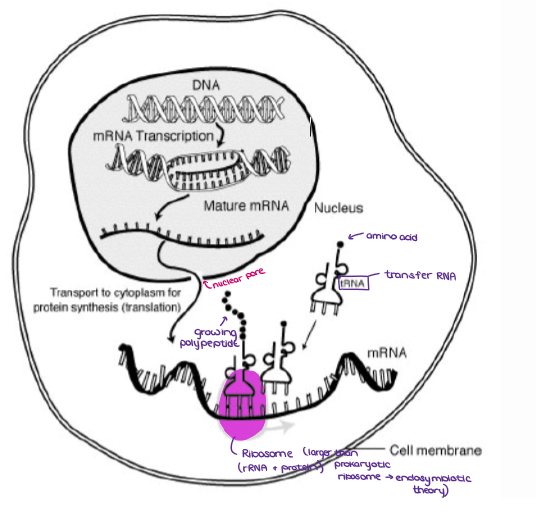
(GENE EXPRESSION) AMINO ACIDS AND THE GENETIC CODE
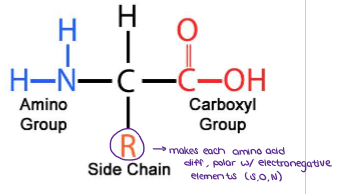
What are AMINO ACIDS?
20 different amino acids exist
Protein synthesis occurs in the cytoplasm to make polypeptides by forming PEPTIDE BONDS between amino acids to build the chain
(GENE EXPRESSION) AMINO ACIDS AND THE GENETIC CODE
Explain the role of DNA and TRIPLETS.
DNA contains the instructions for making proteins but is too large to leave the nucleus
Three consecutive bases on DNA called a triplet (e.g. TCG, ATG, ATT)
Triplets are transcribed into mRNA codons
(GENE EXPRESSION) AMINO ACIDS AND THE GENETIC CODE
What are the START and STOP CODONS?
Methionine (AUG) on mRNA is called the start codon because it triggers the linking of amino acids
UAA, UGA, & UAG are “stop” codons on mRNA signal ribosomes to stop linking amino acids together
(GENE EXPRESSION) THE GENETIC CODE
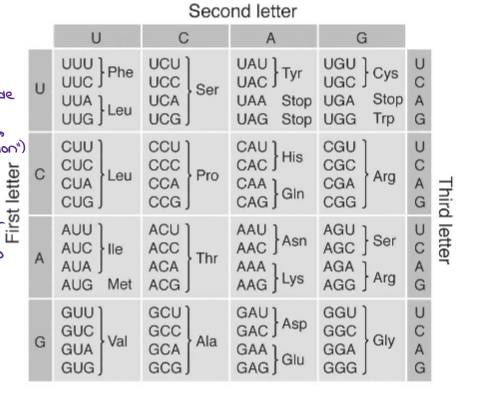
What are CODONS?
mRNA codons code for amino acids
43 = 64 codons (4 bases, arranged by 3)
There are 64 codons that code for 20 amino acids. (This reduces the # of possible mistakes in the amino acid sequence)
The code is REDUNDANT → several codons code for the same amino acids (3rd position → “Wobble Position”) → if first 2 are the same, 3rd position doesn’t matter (will still code for same amino acid)
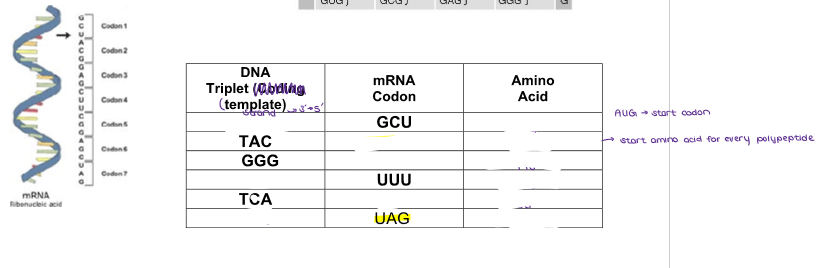
(GENE EXPRESSION) THE GENETIC CODE
Fill in this chart of DNA TRIPLETS, mRNA CODONS, and AMINO ACIDS.
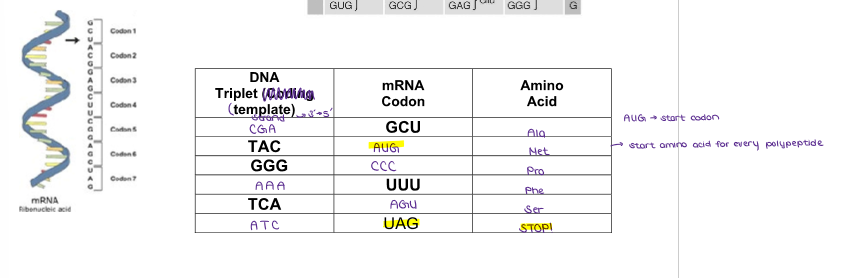
(GENE EXPRESSION) Steps in Transcription
Where does TRANSCRIPTION occur?
IN THE NUCLEUS
(GENE EXPRESSION) Steps in Transcription
What are the STEPS IN TRANSCRIPTION?
**TRANSCRIPTION FACTORS ATTACH TO THE DNA TO ACTIVATE WHICH GENE WILL BE TRANSCRIBED**
DNA helicase (enzyme) unzips the DNA molecule by breaking the hydrogen bonds between the two strands of DNA
The 2 DNA strands separate, but only one will serve as the template & be copied
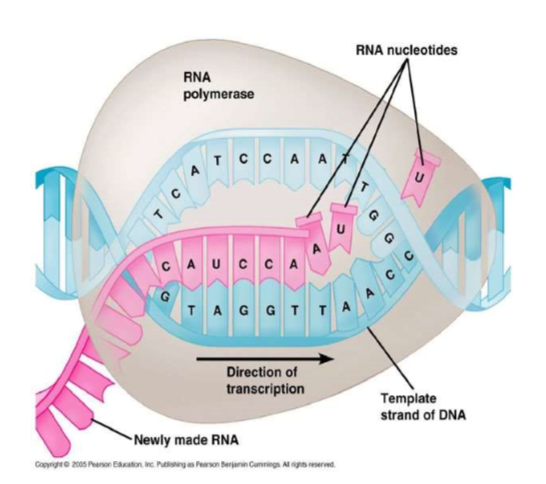
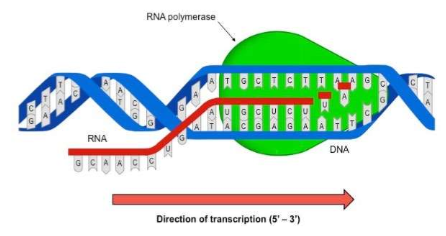
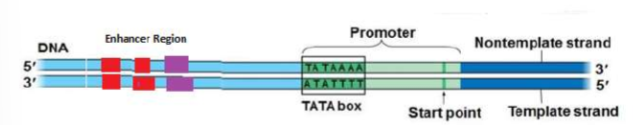
RNA polymerase (enzyme) binds to a region of DNA called the PROMOTER REGION (The “start” of a gene), which is recognized by a “TATA box” section on the DNA (helps RNA recognize that this is where I need to attach)
Free RNA nucleotide (floating in the nucleus) are joined to the template by RNA polymerase in the 5’ to 3’ direction to form the mRNA strand. The RNA nucleotides will base pair with the DNA strand…. But in place of “thymine,” URACIL will pair with Adenine.
The mRNA molecule continues to be built until the RNA polymerase reaches an area on DNA called the termination signal (→ stop codon).
RNA polymerase breaks loose from DNA and the newly made mRNA is released.
ONCE AN mRNA TRANSCRIPT IS COMPLETE, IT MUST BE EDITED BEFORE the final mRNA transcript MOVES TO THE CYTOPLASM FOR TRANSLATION.
(GENE EXPRESSION) Structure of an RNA Polymerase Promoter
What is a PROMOTER?
Region on DNA that allows the attachment of RNA Polymerase
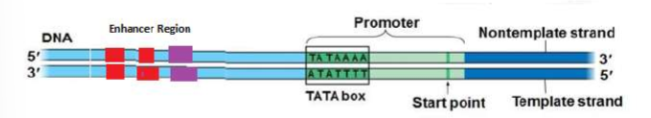
(GENE EXPRESSION) Structure of an RNA Polymerase Promoter
Explain the role of the TATA box.
Eukaryotic promoters are much larger and more complex than prokaryotic promoters, but both have a TATA box. For this gene, the exact TATA box sequence is TATAAAA, as read in the 5’ to 3’ direction. The thermostability of A—T bonds is low and this helps the DNA template to locally unwind in preparation for transcription.
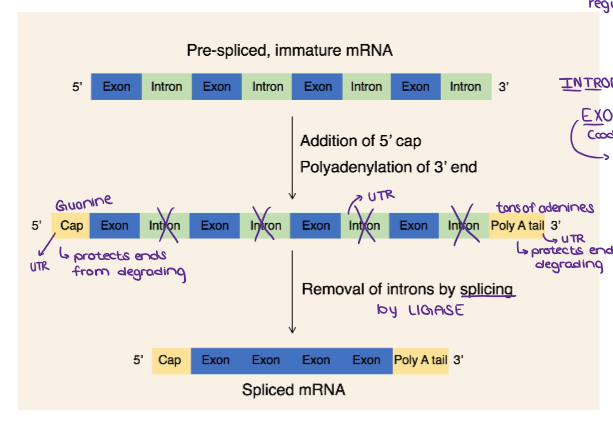
(GENE EXPRESSION) mRNA EDITING & PROCESSING (Post-Transcriptional Modification)
Explain this process.
In eukaryotes, the mRNA needs to be EDITED before it leaves the nucleus to be translated.
Add a 5’ Guanine Cap (5’ cap)
A modified guanine nucleotide is added to the front (5’ end0.
Purpose:
Protects mRNA from degradation
Helps the ribosome recognize and bind tot he mRNA for translation
Add a Poly-A Tail
A long chain of adenine nucleotides (A’s) is added to the 3’ end.
Purpose:
Increases stability (prevents breakdown)
Helps mRNA exit the nucleus
Splicing: Remove Introns
Introns (noncoding sections) are cut out and discarded
Join Exons Together
Exons (coding sequences) are spliced together to form the final edited transcript; Exit the nucleus and get expressed
(GENE EXPRESSION) mRNA EDITING & PROCESSING (Post-Transcriptional Modification)
What are UTRs?
Regions on DNA that do not code for amino acids (never become proteins):
UTRs (untranslated regions)
some of them are “regulatory” (serve another purpose) → it helps with other processes
(GENE EXPRESSION) SPECIAL PROTEINS/RNA/ENZYMES USED DURING TRANSCRIPTION “EDITING” OF RNA
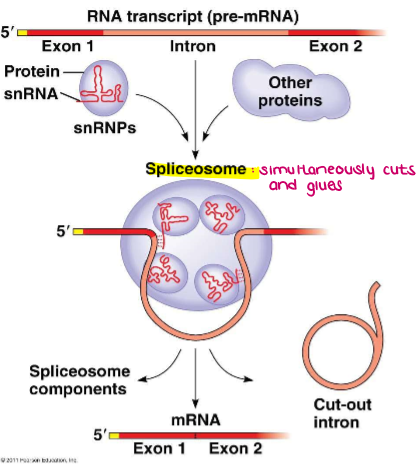
What are snRNPs (small nuclear ribonucleoproteins)?
Found inside the nucleus
They are composed of snRNA (small nuclear RNA) and protein (about 150 nucleotides long)
Recognize the “splice” or “cut” sites at the ends of introns
snRNPs join with additional proteins to form a bigger complex called a SPLICEOSOME
together they cut out the intron and put back together the exons of the mRNA!
(GENE EXPRESSION) SPECIAL PROTEINS/RNA/ENZYMES USED DURING TRANSCRIPTION “EDITING” OF RNA
What are RIBOZYMES?
RIBOZYMES are RNA molecules that function as enzymes. In some organisms, the introns are the ribozymes and catalyze its own excision from the mRNA!
evidence that RNA has high capabilities/very versatile
(GENE EXPRESSION) SPECIAL PROTEINS/RNA/ENZYMES USED DURING TRANSCRIPTION “EDITING” OF RNA
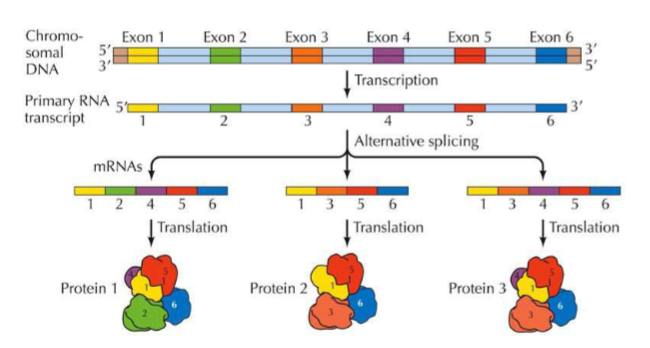
What is ALTERNATIVE SPLICING?
Alternative splicing is a regulated process during gene expression that results in a single gene coding for multiple proteins. In this process, particular exons of a gene may be included within or excluded from the final, processed mRNA produced from that gene → it increases the variety of proteins that can be made from a single mRNA transcript!
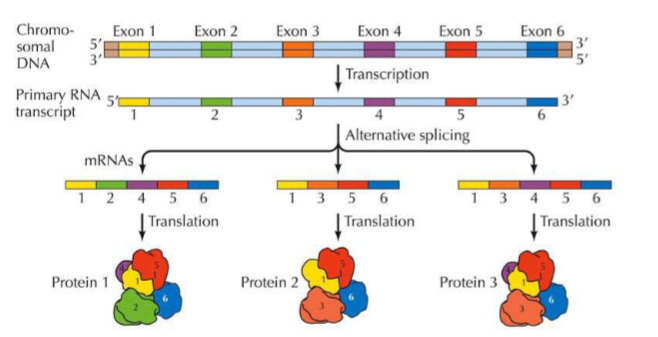
(GENE EXPRESSION) SPECIAL PROTEINS/RNA/ENZYMES USED DURING TRANSCRIPTION “EDITING” OF RNA
Using this diagram, what exon is critical for proteins on this transcript?
1, 5, 6 (present on all 3 proteins)
(GENE EXPRESSION) TRANSLATION
In order for translation to occur, the following parts are necessary…
An edited (exons only) mRNA strand — carries the copied genetic information to the cytoplasm
A ribosome - where all of the components come together to make a polypeptide
Transfer RNA (tRNA) molecules carrying amino acids.
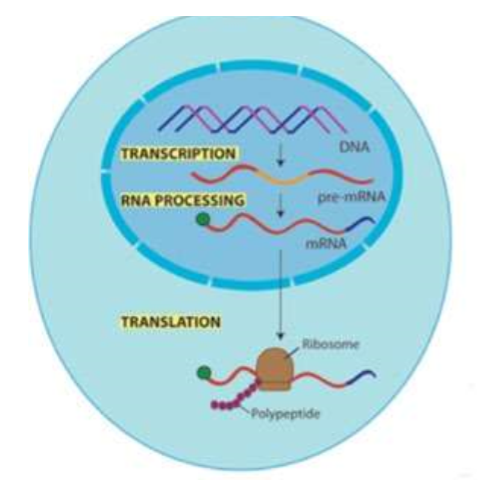
(GENE EXPRESSION) TRANSLATION
The ____ molecule is “read” _ nucleotides at a time, called a ____ — each codon codes for an _____ ____. (There are 64 codons, so each amino has more than one codon.) There is a “_____” codon and several “____” codons, signaling when to start and stop protein synthesis.
mRNA
CODON
amino acid
start
stop
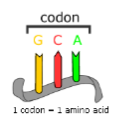
(GENE EXPRESSION) TRANSLATION
What is a tRNA MOLECULE?
A tRNA molecule is a “clover-leaf shaped molecule” with a triplet code on the end called an ANTICODON (in this case, CGG), and an amino acid attached at the other end. This anticodon will match up with an mRNA codon, GCC.
Transfer RNA (tRNA) is responsible for bringing the correct amino acid to the ribosome for attachment to the polypeptide chain.
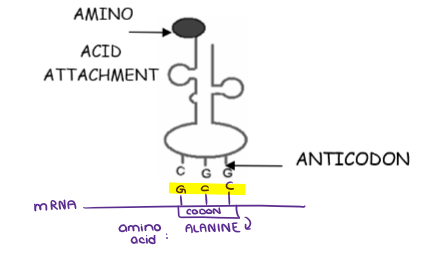
(GENE EXPRESSION) TRANSLATION
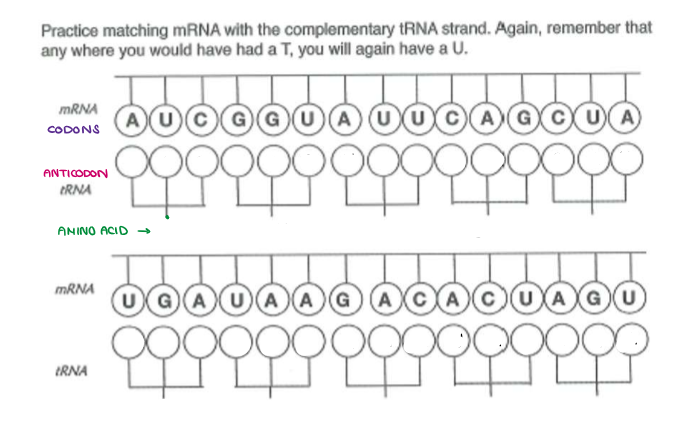
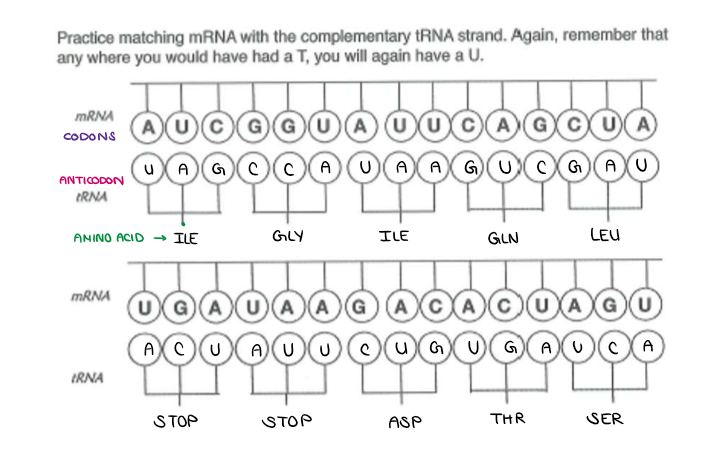
Fill in the ANTICODONS and AMINO ACIDS.
(GENE EXPRESSION) Process of Translation
Describe the process.
Initiation: The mRNA binds to the ribosome, and the start codon (AUG) is recognized.
Elongation: Transfer RNA (tRNA) molecules bring amino acids to the ribosome, where they are added one by one to the growing polypeptide chain.
Termination: The ribosome reaches a stop codon (UAA, UAG, or UGA), signaling the end of translation. The newly formed polypeptide is released.
→ The mRNA transcript can be utilized many times to make more protein product. When complete, enzymes will degrade the transcript and the nucleotides can be recycled.
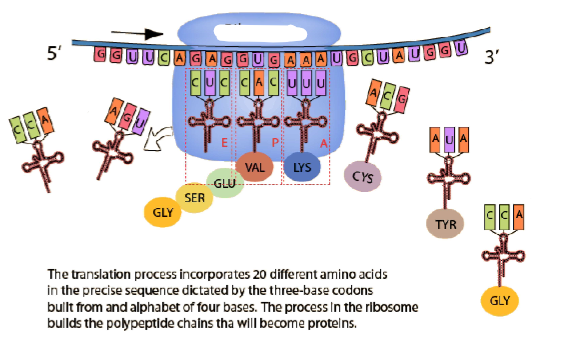
(GENE EXPRESSION) Process of Translation
Label the diagram.