Lab 5 CrRNA Design
1/14
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
15 Terms
How to visualize where protein is in vivo
One option is to make antibodies to that protein
expensive
takes considerable time
requires more animal experimentation.
Another solution is to add a marker, or tag, to our protein.
We can use a tag that encodes a peptide that can be detected with commercially available antibodies, or we can tag our gene with an auto-fluorescent protein like GFP.
This semester, we are using CrisprCas to add the DNA sequence that encodes GFP to unc-32
Review of Crispr-Cas9 components
tracrRNA*, crisprRNA* and Cas9.
In order to make controlled edits, a repair template must also be included.
tracrRNA
contains hairpin loops, giving it secondary structure that is specifically recognized by Cas9, allowing Cas9 and tracrRNA to bind to each other.
The tracrRNA also binds to the crRNA because its 5’ end is complementary to the 3’ end of the crRNA (Figure 1).
tracrRNA is required to link the crRNA to Cas9.
Cas 9 actions
bound to a guide RNA, recognizes the PAM site NGG in target DNA.
When Cas9 binds to a PAM, it unwinds the upstream DNA and checks if the 20nt at the 5’ end of the crRNA (spacer) match target DNA immediately upstream of the PAM (see also Figure 3).
If the region of the crRNA called the spacer (see Figure 1) is complementary to the target DNA, Cas9 creates a double-strand break (DSB) in the target DNA
cut occurs 3 base pairs 5’ to the PAM site.
CRISPR Cas9 diagram
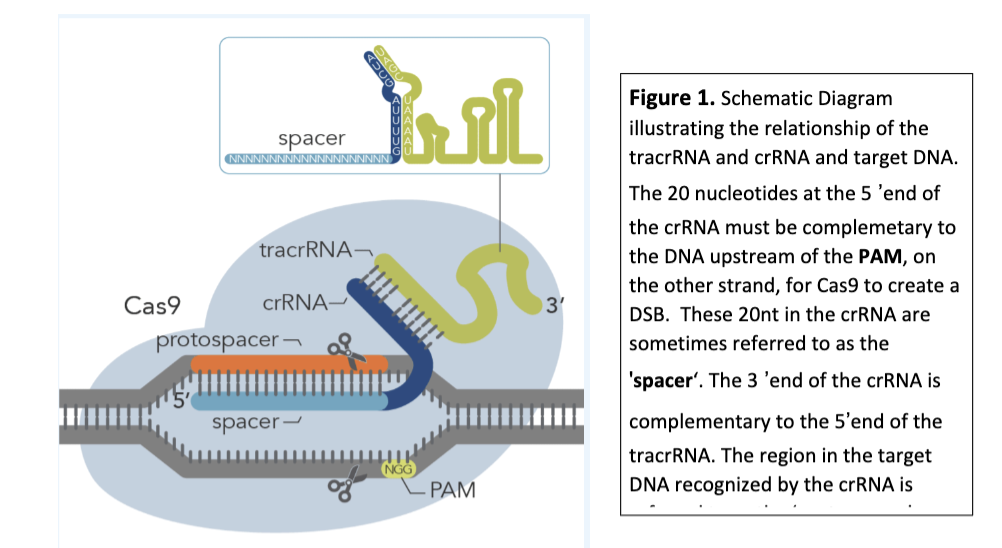
dark blue crRNA segment identical in every experiment, only spacer changes!
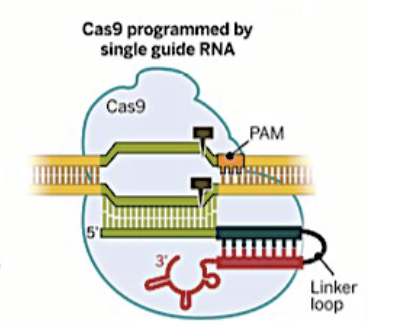
what is the difference between a guide RNA and a single-guide RNA (or sgRNA).
crRNA and trRNA are physically linked into a single molecule by a short linker sequence, called sgRNA
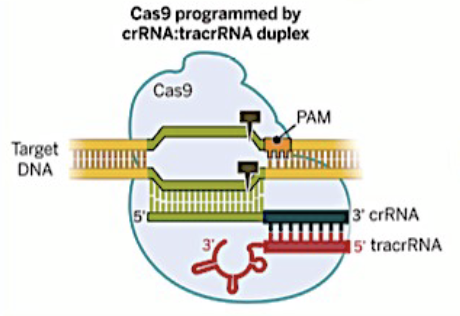
rRNA and trRNA are not physically linked; we rely on the fact that their complementary sequences will hybridize to link the two molecules, guide RNA
Cas9 target search mechanism
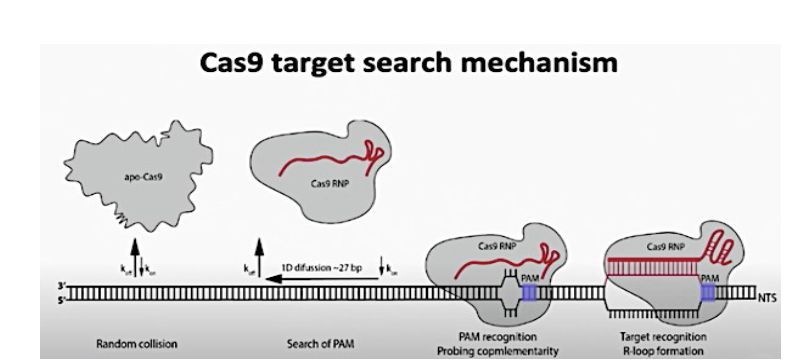
Cas-9 with its guide RNA interacting with target DNA. When Cas9 encounters a PAM site it binds to the target DNA and unwinds the upstream DNA. IF the 20 nt spacer region of the crRNA is complementary to the 20 nt in the target upstream of the PAM, Cas9 will create a DSB 3nt upstream of the PAM.
Location of the DNA encoding the Tag concerns
need the sequence encoding GFP to get translated when unc-32 gets translated, and in a way that the 2 proteins are linked.
position the tag in a way that it does not disrupt the function of UNC-32
how do we address location concerns
for gfp to be transcribed when unc-32 is transcribed, both need to be in the same open reading frame. want regulatory regions that control unc-32 expression to also drive the expression of gfp
solution: insert the DNA encoding the tag directly into the chromosomal location of our gene.
Where do we put a tag so it does not disrupt the function of a protein?
no guaruntees
If the structure of a protein is known, then one can make an educated guess about where the least disruptive locations are likely to be.
If we don’t know the structure of our protein, we usually add the tag to the N- or C-terminus, or more likely, we try both.
seems “safer” to add the tag to the ends of our protein. That way we are not interrupting the amino acid sequence of our protein with amino acids from another protein, and we hope we are not disrupting the folding or structure of our protein.
Designing our crisprRNA steps
find a PAM site near to where we are inserting gfp.
can use your understanding of how CrisprCas works to design the
crRNA.
retrieve the sequence of unc-32, before we can design crRNAs.
How was C. elegans genome sequenced
2 ways
by shotgun sequencing and by using ordered libraries of subclones of the genome.
For both methods, genome sequencing begins with chopping chromosomes up into smaller pieces, usually with restriction enzymes.
shotgun sequencing
smaller pieces are either directly sequenced or they are cloned into vectors and sequenced.
When enough fragments are generated, numerous overlapping physical pieces of DNA cover each location in the genome. (See Figure 2.)
Computer programs can be used to assemble these smaller DNA sequences by aligning overlapping regions of the fragments.
Ultimately, if enough fragments are sequenced, the assembled DNA sequence will represent a large proportion of the genome.
One drawback of this method is that sequences cannot be reliably assembled in chromosomal regions that contain extended DNA repeats.
Alternatively to shotgun sequencing
fragments can be cloned into separate vectors so they can be maintained in bacteria or yeast (Fig. 5). The “vector” you may have heard of is a plasmid. Typical plasmids hold only small pieces of DNA, usually a maximum of 10kb. Vectors called cosmids and fosmids can reliably maintain fragments of DNA around 40Kb in size. YACs, or yeast artificial chromosomes and BACs, bacterial artificial chromosomes can contain inserts as large as 1 Mb. The inserts can be placed in order along a chromosome by hybridization. The complete genome sequence of C elegans was first obtained by sequencing overlapping vector inserts.
What does the method of genome sequencing have to do with the name of genes in C. elegans?
Once the genome is sequenced, we can predict likely open reading frames (ORFs) in the genome.
These predicted ORFs are given a name based on the vector they are contained in. (an alphanumeric unique identifier)
We may know nothing about the function of these predicted genes, although if they have regions of homology to genes that have already been studied, we can form hypotheses about their function.
Another way genes are named is the way genes are classically named in genetic model organisms; a mutation was created and even well before we know anything about the gene containing that mutation, we give it a name, usually related to the mutant phenotype.
Finally, genes can also be given a name based on their homology to well-studied genes. For instance, tba-1 and tba-2 were named for their similarity to alpha tubulin, and tbb-1 and tbb-2 were named for their similarity to beta tubulin.