ap bio unit 6 Son Im Crine
1/53
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
54 Terms
pyrimidines vs purines
single ring (u, c, t) vs double ring (a, g)
prokaryotic vs eukaryotic genomes
prokaryotic:
circular
smaller
eukaryotic:
linear, multiple
larger
both contain plasmids which are circular double stranded dna molecules (cytosol in prokaryotes and nucleus in eukaryotes)
what does uracil replace in dna
it replaces thymine (so AU and GC pair up)
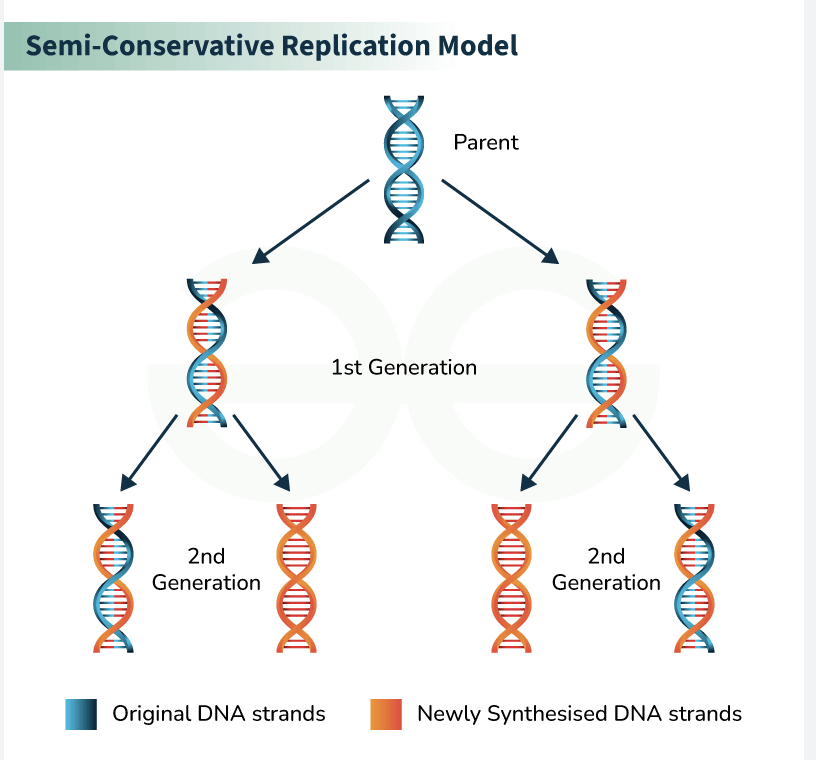
what does DNA replication being semiconservative mean
when a DNA molecule replicates, the two resulting DNA double helices each consist of one original (parental) strand and one newly synthesized strand
5 prime vs 3 prime end
5 prime: phosphate terminus (Five and PHosphate)
3 prime: hydroxyl terminus
nucleotides can only be added in the __ direction, and this means…
5’ to 3’
one strand will always be synthesized continuously (leading strand)
one strand will always be synthesized discontinuously (lagging strand) (the fragments are called okazaki)
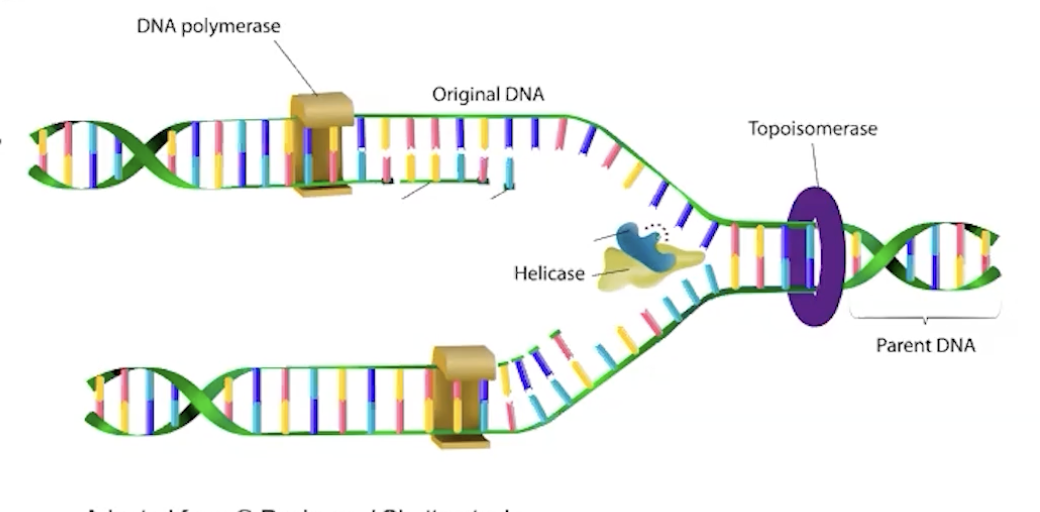
what are the enzymes involved in replication and what do they do
helicase: unwinds the dna
primase: synthesizes short RNA primers (starting from 3’ on the template strand, strands are antiparallel so at that spot the other strand is 5’) that provide starting points for DNA synthesis
topoisomerase: prevents strands from being over twisted at the replication fork (the place where the strands are separated)
dna polymerase: adds complementary DNA nucleotides to the growing strand in the 5′ → 3′ direction and proofreads for errors
ligase: joins dna fragments on lagging strand
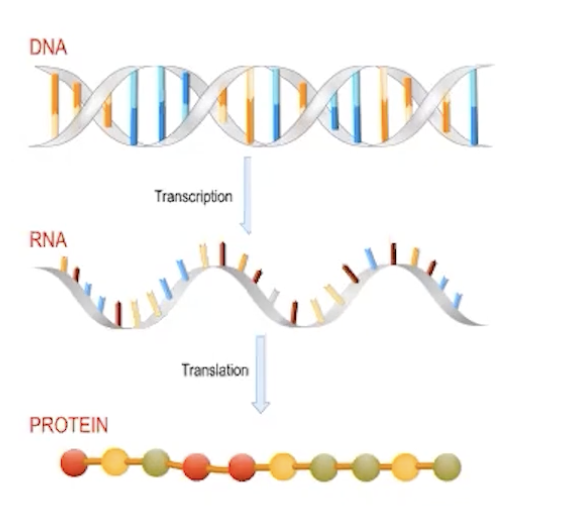
central dogma
DNA → RNA → protein
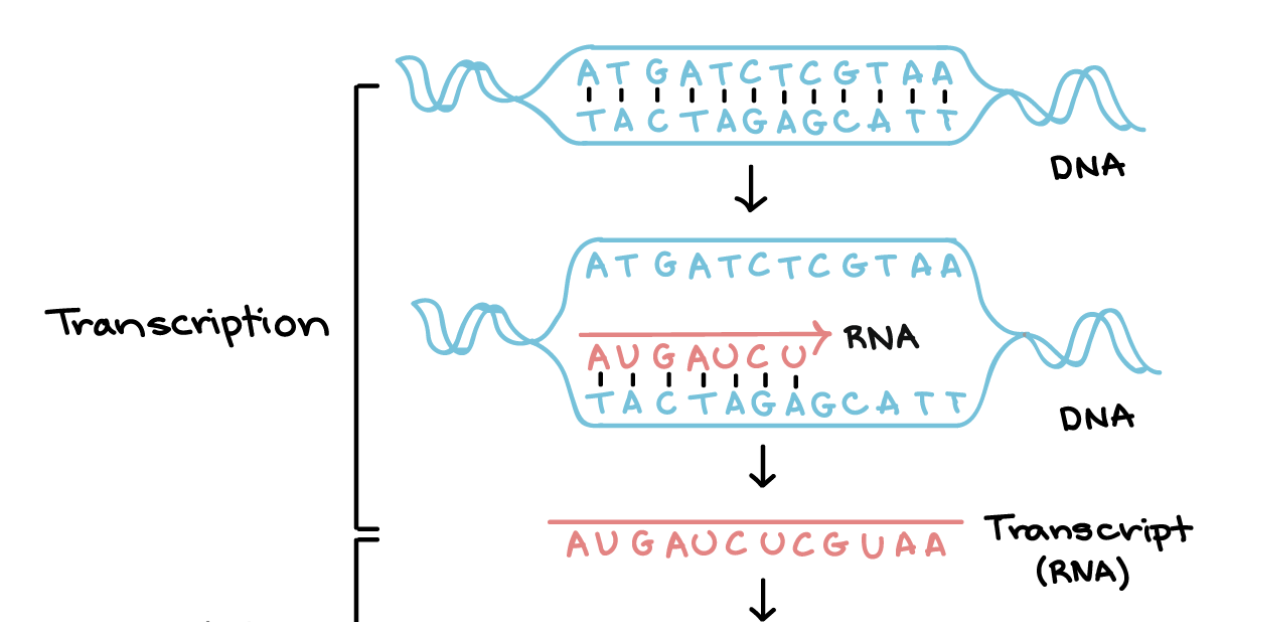
DNA transcription
initiation: RNA polymerase binds to a sequence of a DNA gene called the promoter and separates the two strands
elongation: one strand is the template (noncoding, antisense) and the other is non-template (coding). the RNA polymerase synthesizes a new RNA strand (5’ → 3’) using the template which is the same as non-template but with uracil
termination, we have a new mRNA and DNA is put back to normal
3 types of RNA
mRNA:
carries genetic info from DNA to ribosomes, 3 base sequence is codon
tRNA:
transfers specific amino acids to the ribosome; contains anticodon that pairs with mRNA codons to ensure the correct amino acid is added to the growing polypeptide chain
rRNA:
forms core and structure of ribosomes
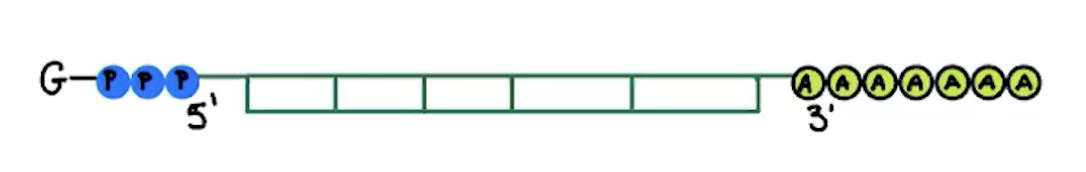
eukaryotic mRNA modifications
addition of poly-A tail
100-200 adenine nucleotides that helps stability (green circles)
addition of GTP cap
modified guanine nucleotide that helps protect it and allow ribosome to attach
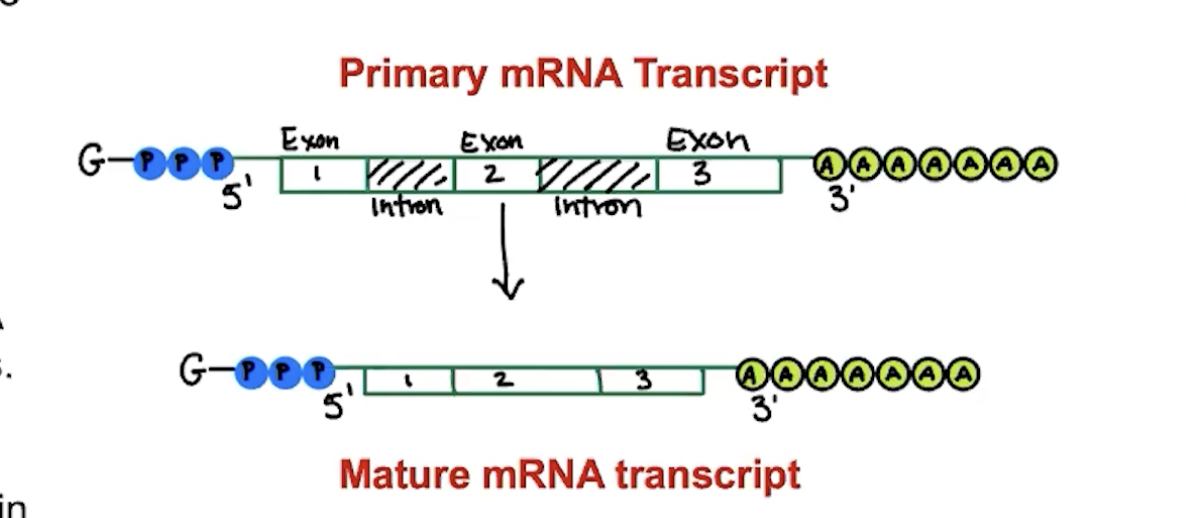
introns
sequences of mRNA that do not code for amino acids. they’re removed during RNA processing (this is called alternative splicing) and are thus not a part of the mRNA transcript
exons
sequences of mRNA that do code for amino acids and are used in processing and then connected to each other to make mature mRNA transcripts
what happens if introns are not removed
they’ll be translated along with the exons and create gibberish