AP Biology Unit 6: Gene Expression
1/53
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
54 Terms
Central Dogma of Molecular Biology
DNA makes RNA makes proteins aka DNA triplets → mRNA codons → amino acid sequence (protein).
DNA
Double stranded helical molecule made of nucleotide monomers, with each nucleotide consisting of:
deoxyribose sugar (5 carbon sugar)
phosphate group
1 of 4 nitrogenous bases (A, T, C, or G)
Sugar and phosphate form sugar phosphate backbone connected by covalent bonds
Nitrogenous bases pair the helical structure through hydrogen bonds because of the complementary base pairing (AT CG)
Strands are anti-parallel: one runs 5’ to 3’ and another 3’ to 5’ for base pairing to occur
4 Characteristics that make DNA’s life primary genetic molecule
Information Storage:
DNA’s sequence of bases (e.g., ACGT) acts as an informational code specifying RNA and protein sequences.
Replicability:
Base-pairing (A-T, G-C) allows DNA strands to serve as templates for complementary strand synthesis, ensuring accurate replication.
Stability:
The double helix protects the bases, making DNA a stable molecule.
Mutability:
DNA can mutate (spontaneously or due to environmental factors), allowing for genetic variation and evolution.
RNA
Single stranded molecule involved in protein synthesis and gene regulation
3 types of RNA:
mRNA: messenger RNA
tRNA: transfer RNA
rRNA: ribosomal RNA
Bases are AU and CG
Sugar is ribose
Prokaryotic vs. Eukaryotic Genetic Information Storage
Prokaryotes:
DNA is stored in looped circular chromosomes (no start or end).
Genomes are small (100,000 to 10 million base pairs) and naked (not wrapped around proteins).
Eukaryotes:
DNA is stored in linear chromosomes.
DNA is wrapped around proteins called histones.
Eukaryotic genomes are much larger than prokaryotic genomes.
Plasmids
Small, extra-chromosomal loops of DNA found in bacteria that facilitate:
horizontal gene transfer (transferring antibiotic resistant genes)
genetic engineering for DNA replication and expressing engineered genes in bacteria
bacterial conjugation: transfer of plasmids between cells
Origin of Replication
Specific sequence where replication begins
Helicase
Unzips DNA via breaking hydrogen bonds between strands creating a replication fork
DNA Polymerase
Synthesizes new strands by adding nucleotides to the 3’ end of a growing (extends from existing) strand
Single-Strand Binding Proteins
Prevent re-winding of DNA strand
Primase
Lays down RNA primers to provide a starting point for DNA polymerase
Ligase
Seals gaps between fragments
Topoisomerase
Relieves supercoiling during DNA replication via cutting & rejoining DNA strands
Leading Strand
Synthesized continuously in same direction as replication fork opens
Differs from lagging strand b/c of directionality of DNA polymerase
Lagging Strand
Synthesis moves in opposite direction of the opening replication fork by forming Okazaki fragments each requiring a new primer
DNA Replication
Semiconservative process with each strand of the double helix acts as a template for synthesizing a new complementary strand
Process:
Enzymes pull the double helix apart.
Each single strand acts as a template.
Free nucleotides in the nucleus pair with exposed bases following base-pairing rules:
A pairs with T, and C pairs with G.
Transcription
Process in the nucleus
Begins at the promoter region on the DNA, marking the gene’s starting point
RNA polymerase binds to the promoter and synthesizes RNA based on the DNA template strand
DNA is read in 3’ to 5’ direction and RNA is synthesized in the 5’ to 3’ direction
When RNA polymerase reaches a terminator region, transcription ends
DNA Template Strand
DNA strand that’s transcribed into RNA
Coding Strand
Complementary to the template strand that shares the same sequence as mRNA except RNA has U instead of T
Unique Features of Prokaryotic Transcription
Prokaryotes lack a nucleus so transcription and translation occur simultaneously in the cytoplasm
Transcribed RNA can be immediately translated by ribosomes
Polysomes: Multiple ribosomes can translate the same RNA strand simultaneously
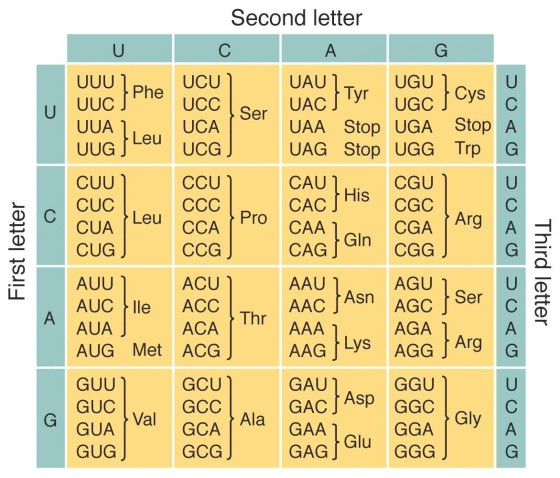
Genetic Code
The code used by living organisms to translate nucleotide sequences (RNA) into amino acid sequences (proteins).
How to read the genetic code
Use a genetic code chart (which can be in a circular or tabular format)
Match the first, second, and third bases of a codon to determine the amino acid it codes for.
Example: AUG codes for methionine (met); GAA codes for glutamic acid (Glu
Translation
Occurs in the cytoplasm
Converts an mRNA sequence into a polypeptide to produce a specific sequence of amino acids that form a polypeptide which then folds to become a functioning protein involving 3 key players:
mRNA: contains codons specifying order of amino acids
tRNA: transfers specific amino acids to the ribosome that consists of:
an anticodon that pairs w/ the mRNA codon
amino acid binding site that carries the amino acid
ribosome: synthesizes the polypeptide by forming peptide bonds between amino acids
Ribosome
Consists of a large subunit and small subunit that comes together to form a ribosome found in the cytoplasm or rough ER
Has three tRNA binding sites:
A site: accepts new tRNA with an amino acid
P site: Holds the growing polypeptide
E site: Where tRNA exits after giving up its amino acid
Steps of Translation
Inititiation:
mRNA leaves the nucleus and attatches to the small ribosomal unit, identifying its start codon AUG. Large ribosomal subunit idenfies UAC (complementary codon) and joins to the small unit, starting translation
Elongation:
tRNA brings amino acids to the ribosome that forms peptide bonds between amino acids in P and A site. Ribosomes move the tRNA to the E site to exit and repeat the process as the polypeptide chain forms and grows
Termination:
Ribosomes reach a stop codon and a release factor binds to the ribosome to release the polypeptide that folds into a functional protein
Operons
A cluster of genes in bacteria that are controlled together and transcribed into a single mRNA, allowing the cell to turn multiple genes on or off at the same time.
Trp Operon (Repressible Operon)
Makes tryptophan (Amino acid) only when the cell doesn’t have enough
How it works:
When tryptophan is LOW (absent) => ON
Repressor is inactive (can’t bind to operator)
RNA polymerase makes tryptophan
Tryptophan production eventually represses its own synthesis (negative feedback).
When tryptophan is HIGH (present) => OFF
Tryptophan acts as a repressor by binding to the operator, blocking transcription
Repressible Operon
Default condition is “on”
Presence of tryptophan turns system off
Lac Operon
Breaks down lactose when it’s available
How it works:
When lactose is ABSENT → OFF
Repressor is active
Binds to operator → blocks transcription
When lactose is PRESENT → ON
Lactose binds to the repressor that become inactive (can’t bind operator)
RNA polymerase creates enzymes to break lactose
Inducible Operon
Default condition is “off”
Lactose turns the system on as the ‘inducer’
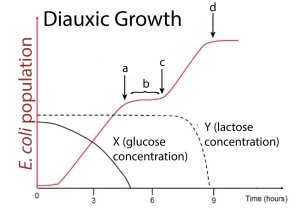
Glucose vs. Lactose Metabolism (AKA: “Diauxic” or “Two-Phase” Growth)
When provided with glucose and lactose:
E. coli prefers glucose as it’s easier to metabolize (monosaccharide vs. disaccharide)
When glucose runs out, there’s a lag while the lac operon activates and produces enzymes to digest lactose.
Euchromatin
Loosely packed (often acetylated) DNA that is transcribed into RNA and translated into protein.
Heterochromatin
tightly packed (often methylated) DNA that is NOT transcribed
RNA Processing in Eukaryotes *Why it’s more complex*
Introns and Exons
Introns: non-coding sequences removed during RNA splicing
Exons: Coding sequences spliced together to form mRNA for translation
Post-Transcriptional Modifications in mRNA
5’ GTP Cap:
Protects mRNA from degradation
Aids in nuclear export and ribosome binding
3’ Poly-A Tail:
Enhances mRNA stability and delays breakdown
microRNAS
Regulates gene expression by forming RNA silencing complexes that either pause translation or degrade mRNA so that it can no longer be translated
Mutation
A random change in DNA or an entire chromosome
Point Mutation
A change in a single nucleotide (ex: C to T)
Types of point mutations:
Silent Mutation: DNA changes, but the amino acid and protein remain the same due to redundancy in the genetic code.
Nonsense Mutation: Inserts a stop codon, halting protein synthesis.
Missense Mutation: Changes one amino acid to another.
Point Mutation Illustrative Example: Sickle Cell Disease
Cause: A missense mutation substitutes valine (non-polar) for glutamic acid (polar) in hemoglobin.
Effect:
Causes hemoglobin molecules to stick together, leading to sickled red blood cells under low oxygen conditions.
Results in tissue damage and is a recessive condition.
Heterozygote Advantage: One copy of the mutation provides resistance to malaria.
Frameshift Mutations
Caused by insertion or deletion of nucleotides, altering the reading frame of codons.
Germline Mutation
Occur in cells that produce gametes, inherited by offspring and present in every cell of the organism.
Can be passed on to future generations.
Subject to natural selection (e.g., sickle cell anemia)
Somatic Mutations
Occur in body cells, not passed to offspring.
Example: Mutations from UV exposure causing skin cancer.
Vertical Gene Transfer
Transmission of genetic material from parents to offspring, where organisms pass on all or half of their genome
For example, a bacterium copying its entire genome during reproduction or humans passing genes through gametes.
Horizontal Gene Transfer
When organisms get genes from other organisms (not their parents)
In single-celled organisms these genes are passed on when they reproduce, but in multicellular organisms they must enter reproductive cells to be inherited.
Bacterial Conjugation
Direct transfer of DNA (usually a plasmid) between two bacterial cells through a physical connection called a pilus, where the DNA is copied and passed to the recipient.
Bacterial Transformation
Process in which a bacterium takes in free DNA from its environment and incorporates it into its own genome.
Viral Transduction
Transfer of genetic material from one bacterium to another via a virus, when viral particles accidentally carry bacterial DNA and insert it into a new host cell.
Viral Recombination
Process where two different viruses infect the same cell and exchange genetic material, producing new viral combinations.
Recombinant DNA
DNA made by combining DNA from different species
Made by restriction enzymes cutting DNA creating sticky ends and matching the ends base pair via hydrogen bonding. DNA ligase seals the bond to form a DNA molecule
Recombinant Plasmids
Plasmid and foreign gene are cut with the same enzyme and if their sticky ends match ligase joins them and inserted into the bacteria
Bacteria can now produce foreign proteins
Gel Electrophoresis
Technique that separates DNA fragments by size using an electric current, where smaller fragments move farther through a gel.
Longest are near negative charge
Shortest are near positive charge
Restriction Site Mapping
Process of identifying where restriction enzymes cut DNA by analyzing fragment sizes (often using gel electrophoresis).
PCR (Polymerase Chain Reaction)
Method used to rapidly make millions of copies of a specific DNA sequence through repeated heating, cooling, and DNA replication.
DNA Sequencing
Process of determining the exact order of nucleotides (A, T, C, G) in DNA.