Activity 6 (Week 10)
1/50
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
51 Terms
RNA Pol 1
Connect fragments, removing primers (same function as in prokaryotes)
Ribosomal RNA genes
RNA Pol II
Transcribe polypeptide coding genes into mRNA (12 subunits)
mRNA and some snRNA genes
RNA Pol III
Transcribes tRNA, 5S RNA (subunit), snRNA (splicing)
Transcription Process (1)
Promoter Recognition
Transcription Process (2)
Transcription Initiation
Transcription Process (3)
Elongation
Transcription Process (4)
Termination
Transcription Start Site
DNA Sequences at Eukaryotic Promoters
+1
Usually A or G
TATA Box
DNA Sequences at Eukaryotic Promoters
-30
TATAAA
Bound by TATA Binding Protein (TBP)
Downstream Promoter Element (DPE)
DNA Sequences at Eukaryotic Promoters
+28 to +33
Bound by TAF6 and TAF9
CAAT Box
DNA Sequences at Eukaryotic Promoters
About -80
GGCCAATCT
Bound by the CAAT box transcription factor (CTF)
GC Element
DNA Sequences at Eukaryotic Promoters
Somewhere between -200 AND +1 (NO precise location)
Bound by TF SP1
General Transcription Factors
are REQUIRED to form the RNA pol II complex
Non-general Transcription Factors
are NOT REQUIRED, they just help RNA pol II find TATA box
TFIID
TAF + TBP
Binds to TATA Box
Could be accompanied by other non-general factors to other parts of the promoter
TFIIA
Stabilizes TBP binding to DNA and prevents repressor binding
TFIIB
Helps position RNA polymerase II correctly at the start site
TFIIF
Escorts RNA polymerase II to the promoter
TFIIE
Recruits the TFIIH and regulates its activity
TFIIH
Unwinds DNA (helicase) and phosphorylates RNA polymerase II CTD to start transcription
Poised State
Phosphorylation of RNA Pol II
RNA Pol II pre-initiation complex is NOT phosphorylated
Complex is formed but not transcription has occurred yet
Paused State
Phosphorylation of RNA Pol II
Recruits mRNA processing machinery
CDK-7 phosphorylates Serine 5 in C-Terminal domain of RNA pol II → now in paused state
CDK-9 phosphorylates Serine 2 → release pol II → transcription
Enhancers
(upstream ~ -500, downstream, or within the gene) work with promoters to regulate transcription = increases transcription
loop over to contact promoters and help initiate and increase transcription
Silencers
sequences that suppresses transcriptions
ex. Pierre Robin Syndrome: cleft palate, reduced lower jaw, posteriorly displaced tongue
Due to mutation in the enhancer for a gene called Sox9
Prevents binding of a TF called Twist1 to the enhancers
pre-mRNA Processing: 5’ End
Called 5’ end cap OR 5’ m7G cap → essential for translation
Importance of 3’ PolyA Tail
Poly-adenylation is part of transcriptional termination
Protecting mRNA from degradation (linked to lifespan of mRNA)
Facilitating export of mature mRNA across the nuclear membrane to the cytoplasm
Addition of the 5’ mG cap
Guanylyl transferase (GTase) adds a guanine to the 5’end of the pre-mRNA (additional enzymes then methylate the newly added guanine)
Addition of polyA tail
Addition of ~200-300 adenines to the 3’end of the pre-mRNA → polyadenylation - necessary for RNA stability and transport into the cytoplasm
RNA Splicing
Occurs for almost all protein coding genes
Specific internal regions of transcript are removed and remaining pieces joined together
Mechanism of RNA Splicing
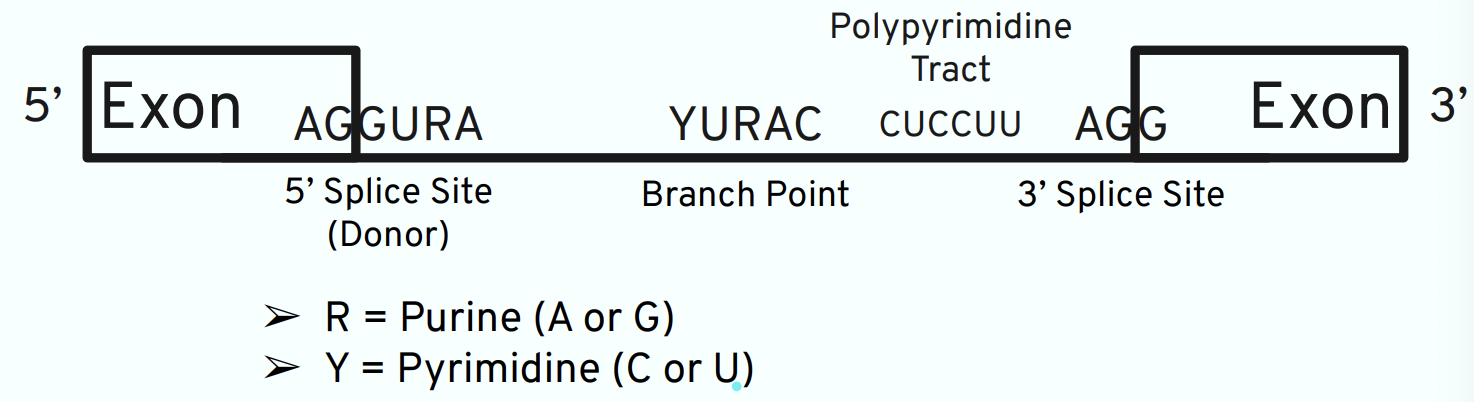
Requires specific sequences in mRNA
Cellular machinery called the spliceosome
Spliceosome - complex of proteins and snRNA
Steps in Splicing (1)
Cleavage at 5’ splice site and transesterification to join the G of 5’ splice site to branch point = forming “lariat”
Steps in Splicing (2)
Cleavage at 3’ splice site
Steps in Splicing (3)
Joining of exons and degradation of intron
Exons
become part of mature mRNA and encode segments of proteins
Introns
are intervening sequences that separate exons
during the process of splicing introns are removed and exons are spliced back together
Chromatin
Consists of DNA and nucleosomes, which are made up of proteins called Histones
Enables packaging DNA to fit in nucleus
Regulates DNA replication, transcription and mRNA processing
Half of the protein content of chromosomes are histone proteins
Histones
Nucleosome = DNA + Histone Octamer(core): 2 molecules of each of H2A, H2B, H3, and H4
High in positively charged amino acids (Lysine, Arginine, and Histidine) = can bind tightly to negatively charged DNA
Heterochromatin
Densely compacted and inhibits transcription
“closed” or “silent” chromatin
Euchromatin
loosely compacted and is accessible to transcriptional machinery
“open” chromatin
Writers
Chromatin Modifiers - Types of Enzymes
Add chemical groups
Erasers
Chromatin Modifiers - Types of Enzymes
Remove chemical groups
Readers
Chromatin Modifiers - Types of Enzymes
Proteins that recognize the modified histones
Chromatin Remodelers
enzymatic complexes that can move nucleosomes around to change compaction state of chromatin (use ATP for energy)
Nucleosome sliding
Sliding the nucleosome will expose the enhancer or promoter element = allowing TFs to bind
Chromatin modifiers
chemically modify histones by adding or removing chemical groups, called post-translation histone modifications (histone PTMs)
Acetyle, methyl and other chemical groups at the specific amino acids in the amino terminal “histone tails”
Active transcription
Lysines K9 and K27 acetylated = Neutralizes the negative charge DNA and relaxes the tight hold of nucleosome on DNA
“Silent” no transcription
Lysines K9 and K27 methylated
DNA Methylation
does not affect nucleosome and DNA interactions, but instead creates binding sites for regulatory proteins
Addition of methyl groups is accomplished by histone methyltransferases (HMTs)
H3K9 tri/di-methylation
Constitutive heterochromatin = ALWAYS “closed”
H3K27 tri/di-methylation
Faculative heterochromatin = Switches between hetero and euchromatin
H3K4 tri-methylation
Active promoters in euchromatin